Picongpu: model without target
Hello,
I wanted to compile a model without any target at all just to see how the laser behaves when compared with the case when the target is present. So I thought the simplest way is to set the density function to 0 as float_64 dens = 0.0;, but I also set the constexpr float_64 BASE_DENSITY_SI = 1.00e+0; so very small. One of these settings or the combinations of two resulted in the error below. Setting back constexpr float_64 BASE_DENSITY_SI = 1.00e+28; enabled the model to compile. Is this error normal or you may want to get rid of it? What would be the safest way to run a model without target material and without altering the density functor in use?
Thank you
Cristian
/home/quasar/src/spack/opt/spack/linux-linuxmint19-skylake/gcc-7.4.0/picongpu-0.5.0-jwzpukc7grapgx2ixr263anam2xiquxw/include/pmacc/../picongpu/particles/bremsstrahlung/ScaledSpectrum.tpp: In member function ‘picongpu::float_64 picongpu::particles::bremsstrahlung::ScaledSpectrum::dcs(picongpu::float_64, picongpu::float_64, picongpu::float_64) const’:
/home/quasar/src/spack/opt/spack/linux-linuxmint19-skylake/gcc-7.4.0/picongpu-0.5.0-jwzpukc7grapgx2ixr263anam2xiquxw/include/pmacc/../picongpu/particles/bremsstrahlung/ScaledSpectrum.tpp:98:69: error: ‘(-5.88119869e+26 * -5.88119869e+26)’ is not a constant expression
constexpr float_64 classicalElRadius = float_64(ELECTRON_CHARGE*ELECTRON_CHARGE) / (pi * 4.0 * EPS0 * ELECTRON_MASS * SPEED_OF_LIGHT*SPEED_OF_LIGHT);
All 13 comments
Hello @cbontoiu ,
Thanks for reporting, not sure if that's intended. We will have a look.
The simplest way to have no macroparticles in a simulation is to not generate any initially. This is controlled by file speciesInitialization.param. Normally with no particles you want no initialization, for that you can simply replace the definition of InitPipeline with using InitPipeline = bmpl::vector<>; (but please note that you can't remove this definition alltogether - it won't compile then)
Also I have to add that my method rendered the ADIOS files unable to be read. There is some error with my approach. I would try the method that @sbastrakov suggested. Thank you.
@cbontoiu Thanks for reporting this issue.
As @sbastrakov already mentioned, the common way to initialize no target is to set using InitPipeline = bmpl::vector<>; in speciesInitialization.param.
My guess for the error is that setting constexpr float_64 BASE_DENSITY_SI = 1.00e+0; caused our internal unit system to create numbers that are not representable by single precision floats thus causing the issue when defining the classical electron radius in the Bremsstrahlung plugin. Most likely this also causes under-/overflows in the data output of single precision float values in ADIOS. If it is no issue or you to keep the BASE_DENSITY_SI, I would recommand to use the default value.
I also think it is the case @PrometheusPi . Just the error message is not very clear, but I definitely saw this type of error before regarding results of constexpr expressions being out-of-range. If this is the case, it is not that easy to catch everything and report properly, as that would require manual copy-pasting of checks and I guess not worth it, except few select cases. Perhaps (but I don't know, just a speculation) with a new unit system being worked on by @psychocoderHPC we may be able to check and report it better, but that needs to be checked.
Hello,
Today I had a chance to run a simulation without a target as indicated above but there is an error when I try to load the output of the ADIOS plugin.
RuntimeError: [ADIOS1] Internal error: Failed to get ADIOS1 attribute during attribute read
I remember a similar situation when the target was completely ionized and the electrons were pushed away from the simulation domain. There was a similar error from the adios files because there were no particles.
https://github.com/ComputationalRadiationPhysics/picongpu/issues/3256
Is there a workaround to simulate only the laser? I guess one will be to output h5 files.
thank you
It might be that the error originates not from the missing particles but from a error we have currently in PIConGPU that misplaces the particle attributes. Some openPMD-api readers try to read these attributes and thus fail.
A fix is on the way - see #3389.
How did you load the ADIOS data?
@PrometheusPi Hello and Happy New Year. I wish you all the best!
To continue on this problem, I always load my adios-generated data as
os.environ['OPENPMD_BP_BACKEND'] = 'ADIOS1'
path, dirs, files = next(os.walk(OUTPUT_NAME + '/simOutput/bp/'))
ts = io.Series(OUTPUT_NAME + '/simOutput/bp/simData_%T.bp', io.Access_Type.read_only);
and in this case when running without a target i obtained the error shown in this image.
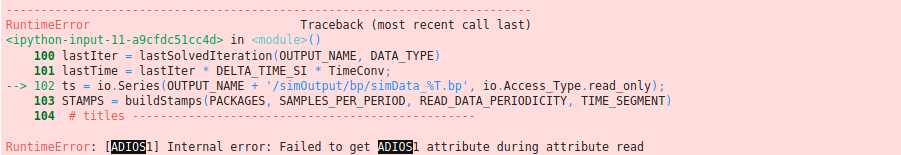
I could attach a file from the results for investigation. it is very small since there are no particles.
@cbontoiu Happy New Year to you as well!
This is an interesting issue. This might be caused by the fact, that out adios1 implementation is not fully openPMD compatible.
Which plugin are your currently (and with target) using to generate the output? (PIConGPU's Adios plugin or openPMD plugin)
Which version of python-adios are you using?
Thanks for offering to share the file - this would be very helpful indeed.
@PrometheusPi Thank you for your reply. To write out PIConGPU data I use in all cases
the adios plugin
TBG_adios="--adios.period !TBG_plugin_period --adios.file simData --adios.source 'species_all,fields_all' --adios.disable-meta 0 --adios.compression blosc:threshold=2048,shuffle=bit,lvl=1,threads=8,compressor=zstd"
and as I remember data written out like this can only be read with the adios module. Thus by doing
os.environ['OPENPMD_BP_BACKEND'] = 'ADIOS1'
I think I use the first version of adios. Listing modules installed for Python I get
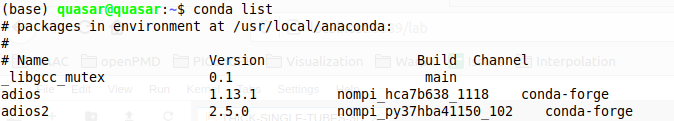
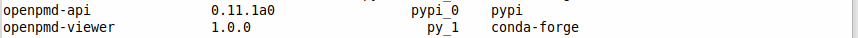
I attach the whole output folder so, if necessary, you could also inspect the input folder apart from the data files.
I am now using PIConGPU: 0.5.0 Build-Type: Release Do I need to upgrade to @develop version in order to use the openPMD plugin? And, if using it, what type of data shall I obtain? How would I read that data then?
Many thanks for your kind help.
Cristian
@cbontoiu Please excuse the late response. I now had the chance to look into your data. The issue is not the one I suspected with our slightly wrong implementation of the adios/openpmd output. (This would create valid adios files with missing openPMD attributes.) However, the adios files provided are not valid adios files. Thus the error occurs at the adios level. Even
bpls -a simData_0.bp
creates a segfault.
If you did not change anything with your libraries, could it be that the file system was corrupted?
What do you get when running bpls on your output data?
Your file is defintly corruped. I used bprecover and it returned:
File size in bytes: 36148
========================================================
Look for a Process Group (PG) at offset 0
PG reported size: 2
=== PG reported size is too small. This is not a (good) PG.
========================================================
Found 0 PGs to be processable
Index metadata size is 96 bytes
To actually recover the file by rewriting the index, use -f option.
Final file will be truncted to 46912571903712 bytes
Following https://github.com/ornladios/ADIOS/issues/124 (for a much older version), the issue could be caused by MPI. Did you change the MPI library between this erroneous simulation and the last time, the adios output was created correctly?
Most helpful comment
I also think it is the case @PrometheusPi . Just the error message is not very clear, but I definitely saw this type of error before regarding results of constexpr expressions being out-of-range. If this is the case, it is not that easy to catch everything and report properly, as that would require manual copy-pasting of checks and I guess not worth it, except few select cases. Perhaps (but I don't know, just a speculation) with a new unit system being worked on by @psychocoderHPC we may be able to check and report it better, but that needs to be checked.