Picongpu: time-resolved probing of a single location
Hi there,
For some plasma instability studies it would be really useful to have a way to probe the fields (including charge-densities) at a given location, and output it as a resolved time-series.
I imagine that one simple way to do this might be setting a single immobile probe particle and gather the fields on it. I've see that there is already a way to do the probes, so technically it should not be too complicated to implement such plug-in for a case of a single particle and arbitrary fields.. what you think ?
All 26 comments
Hi! Yes, the easiest way to achieve this is via probing, immobile particles.
Here is a workflow on it, also including which plugin (HDF5 or ADIOS) can create the output for you: https://picongpu.readthedocs.io/en/0.4.2/usage/workflows/probeParticles.html
In order to probe densities, one can use the same plugins and just needs to modify the attributes stored on the probe particles (speciesAttributes.param, speciesAttributes.unitless -> then put into speciesDefinition.param) and also the "pusher" that stores the fields on the particle attributes: include/picongpu/particles/pusher/particlePusherProbe.hpp
Alternatively, we also have a "slice field" plugin which also records locally fixed values over time. It desperately deserves some updates such as support for more fields and writing to HDF5.
OK, I'm trying the probe particle's method, but I don't get how to limit creation of these probes.
This is supposed to be controlled via EveryNthCellImpl, but whatever I put there (4 4 4 or 1000 1000 1000), it creates particles in all cells.
What could I be missing/should I check ? I followed more or less all the steps from the workflow, but don't see effect of EveryNthCellImpl
@hightower8083 the issue you describe reminds of a rather recent issue #2763 , which has been fixed by #2768; this fix is part of 0.4.1 release. In case you are using a version earlier than 0.4.1 this might be it. However, in case you are using the latest release (or latest dev), this has to be smth else.
Hi @sbastrakov. Thank you for the prompt response!
I think this is actually the case for my problem -- i'm running tests on my desktop machine, where I've 0.4.0 installed. I'll check it on 0.4.2 asap.
By the way, do you have recommendations on a best way to update the spack installation?
Yes, for 0.4.0 this has to be that bug. Sorry, I have no experience with spack.
cc @berceanu just in case you know and have time.
@hightower8083 Can you confirm that #2768 fixed your issue? I am surprised that issue #2763 occurred for your, since you used an equal number of particles per cell thus distributing the macro-particles should have worked. (The bug only occurred for non-equal spacing.)
@hightower8083 go to ~/src/spack-repo and update it's sources. (git fetch --all; git pull --ff-only)
If you now spack install picongpu it will install the latest version alongside the 0.4.0 you have installed.
When loading, make sure to specify the PIConGPU version spack load [email protected] so it knows which version you want. (You can also spack uninstall [email protected] if it is no longer needed).
@ax3l thanx -- upgrade works with no problems
@PrometheusPi -- I've just redone the test and you are right, the issue is still there for 0.4.2.
To clarify
- I clone LWFA example
- do all the steps from
https://picongpu.readthedocs.io/en/0.4.2/usage/workflows/probeParticles.html - in
etc/picongpu/1.cfg, I only change
TBG_steps="100"
[...]
TBG_hdf5="--hdf5.period 1000 \
--hdf5.file simData \
--hdf5.source 'species_all'"
[...]
TBG_plugins="!TBG_hdf5"
- check the output (i'm using opmd_viewer)
from opmd_viewer import OpenPMDTimeSeries
ts = OpenPMDTimeSeries('./test_lwfa_01/simOutput/h5/',)
x, = ts.get_particle(species='probe', var_list=['x',])
print(x.shape)
x, = ts.get_particle(species='e', var_list=['x',])
print(x.shape)
and get 2359296 and 4497408. Factor ~2 difference here is from 1ppc for probes and 2ppc for eons -- should be factor 2x4x4x4=128.
Am I missing something or is a real issue?
@hightower8083 can you please post your speciesInitialization.param?
For the first time step, this can be true since probes are everywhere and regular e- for the gas are just starting (vacuum then upramp in the first simulation steps until moving window starts.) Maybe you can plot each one's "density" for a quick check as well.
here's the speciesInitialization.param file. I'll add the data from simulation shortly
#pragma once
#include "picongpu/particles/InitFunctors.hpp"
namespace picongpu
{
namespace particles
{
/** InitPipeline define in which order species are initialized
*
* the functors are called in order (from first to last functor)
*/
using InitPipeline = bmpl::vector<
#if( PARAM_IONIZATION == 0 )
CreateDensity<
densityProfiles::Gaussian,
startPosition::Random2ppc,
PIC_Electrons
>
# if( PARAM_IONS == 1 )
,
Derive<
PIC_Electrons,
PIC_Ions
>
# endif
#else
CreateDensity<
densityProfiles::Gaussian,
startPosition::Random2ppc,
PIC_Ions
>,
Manipulate<
manipulators::SetBoundElectrons,
PIC_Ions
>
#endif
,
CreateDensity<
densityProfiles::ProbeEveryFourthCell,
startPosition::OnePosition,
Probes
>
>;
} // namespace particles
} // namespace picongpu
Well, looking at the particle's positions in the cells (z-integrated, probes - orange, eons blue, dashed lines are grid), I'd say that ProbeEveryFourthCell does not do what I though it should do -- it creates a particle at each cell
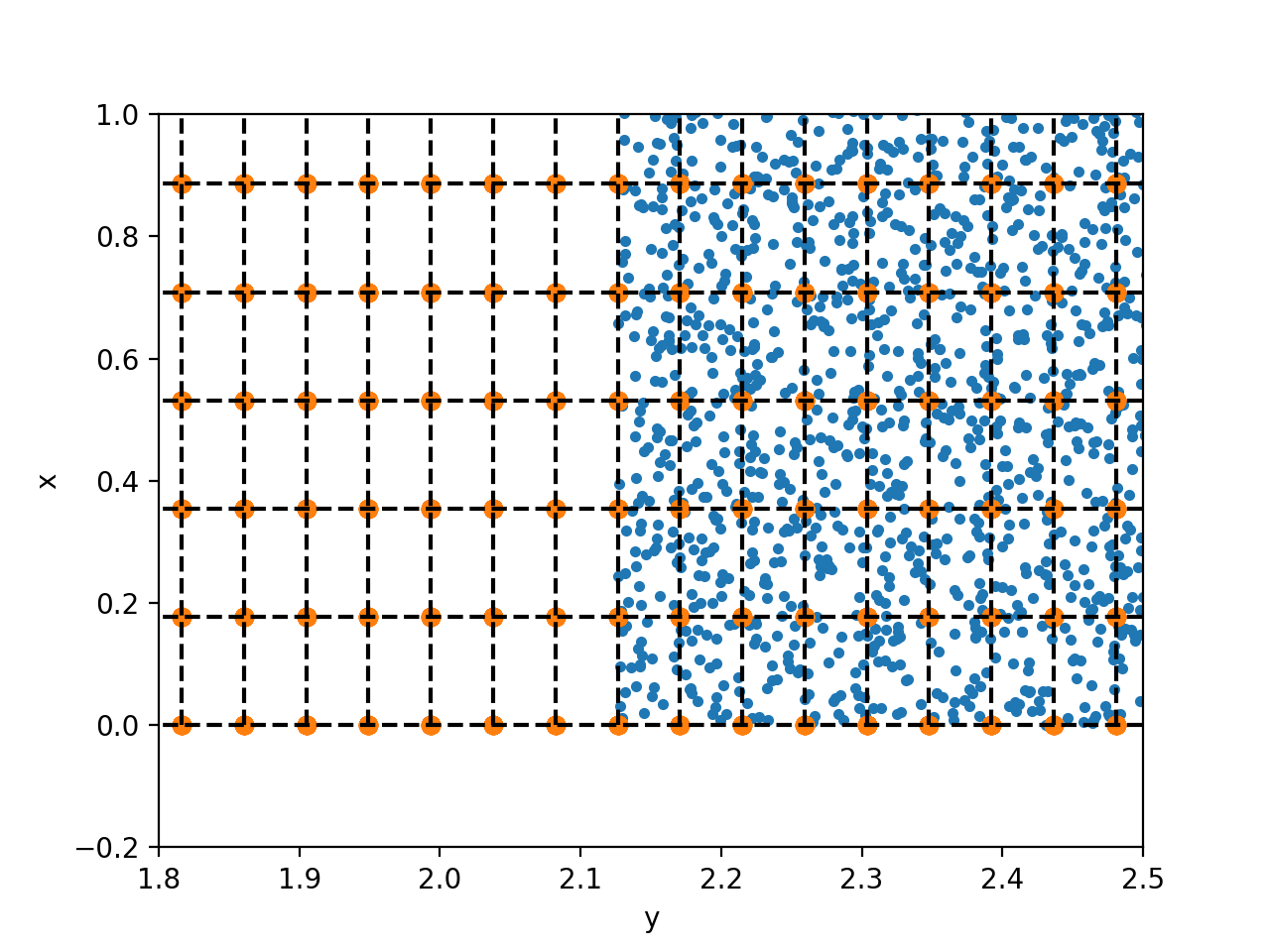
For me ideally would be a plugin which operates with individual locations, but with such numbers of probes, any frequent (e.g. stepwise) output would be enormous
Thanks for checking this!
Can you post the snippets from density.param and particles.param where densityProfiles::ProbeEveryFourthCell and startPosition::OnePosition are defined?
Can you post the head output of your simOutput/output just to verify it's really v0.4.2 you are using?
From looking at https://github.com/ComputationalRadiationPhysics/picongpu/pull/2768, I can't see what might be off right away.
For me ideally would be a plugin which operates with individual locations
Oh you can do this. E.g. by using a finely tailored "free density" profile (in density.param) instead of the pre-defined densityProfiles::ProbeEveryFourthCell and startPosition::OnePosition.
Sure, here they are (its really only a copy-paste):
simOutput/output
PIConGPU: 0.4.2
Build-Type: Release
Third party:
OS: Linux-3.10.0-862.14.4.el7.x86_64
arch: x86_64
CXX: GNU (7.3.0)
CMake: 3.12.3
CUDA: 9.2.88
mallocMC: 2.3.0
Boost: 1.65.1
MPI:
standard: 3.1
flavor: OpenMPI (3.1.2)
PNGwriter: 0.7.0
libSplash: 1.7.0 (Format 4.0)
ADIOS: NOTFOUND
density.param
#pragma once
#include "picongpu/particles/densityProfiles/profiles.def"
/* preprocessor struct generator */
#include <pmacc/preprocessor/struct.hpp>
namespace picongpu
{
namespace SI
{
/** Base density in particles per m^3 in the density profiles.
*
* This is often taken as reference maximum density in normalized profiles.
* Individual particle species can define a `densityRatio` flag relative
* to this value.
*
* unit: ELEMENTS/m^3
*/
#ifndef PARAM_BASE_DENSITY_SI
# define PARAM_BASE_DENSITY_SI 1.e25
#endif
constexpr float_64 BASE_DENSITY_SI = PARAM_BASE_DENSITY_SI;
}
namespace densityProfiles
{
PMACC_STRUCT(GaussianParameter,
/** Profile Formula:
* constexpr float_X exponent = abs((y - gasCenter_SI) / gasSigma_SI);
* constexpr float_X density = exp(gasFactor * pow(exponent, gasPower));
*
* takes `gasCenterLeft_SI for y < gasCenterLeft_SI`,
* `gasCenterRight_SI for y > gasCenterRight_SI`,
* and exponent = 0.0 for gasCenterLeft_SI < y < gasCenterRight_SI
*/
(PMACC_C_VALUE(float_X, gasFactor, -1.0))
(PMACC_C_VALUE(float_X, gasPower, 4.0))
/** height of vacuum area on top border
*
* this vacuum is important because of the laser initialization,
* which is done in the first cells of the simulation and
* assumes a charge-free volume
* unit: cells
*/
(PMACC_C_VALUE(uint32_t, vacuumCellsY, 50))
/** The central position of the gas distribution
* unit: meter
*/
(PMACC_C_VALUE(float_64, gasCenterLeft_SI, 8.0e-5))
(PMACC_C_VALUE(float_64, gasCenterRight_SI, 10.0e-5))
/** the distance from gasCenter_SI until the gas density decreases to its 1/e-th part
* unit: meter
*/
(PMACC_C_VALUE(float_64, gasSigmaLeft_SI, 8.0e-5))
(PMACC_C_VALUE(float_64, gasSigmaRight_SI, 8.0e-5))
); /* struct GaussianParam */
/* definition of density with Gaussian profile */
using Gaussian = GaussianImpl< GaussianParameter >;
using ProbeEveryFourthCell = EveryNthCellImpl<
mCT::UInt32<
4,
4,
4
>
>;
}
}
particles.param
#pragma once
#include "picongpu/particles/startPosition/functors.def"
#include "picongpu/particles/manipulators/manipulators.def"
#include <pmacc/nvidia/functors/Assign.hpp>
namespace picongpu
{
namespace particles
{
/** a particle with a weighting below MIN_WEIGHTING will not
* be created / will be deleted
* unit: none
*/
constexpr float_X MIN_WEIGHTING = 10.0;
namespace startPosition
{
struct RandomParameter2ppc
{
/** Count of particles per cell at initial state
* unit: none
*/
static constexpr uint32_t numParticlesPerCell = 2u;
};
using Random2ppc = RandomImpl< RandomParameter2ppc >;
CONST_VECTOR(
float_X,
3,
InCellOffset,
/* each x, y, z in-cell position component
* in range [0.0, 1.0) */
0.0,
0.0,
0.0
);
struct OnePositionParameter
{
static constexpr uint32_t numParticlesPerCell = 1u;
const InCellOffset_t inCellOffset;
};
using OnePosition = OnePositionImpl< OnePositionParameter >;
} // namespace startPosition
/** During unit normalization, we assume this is a typical
* number of particles per cell for normalization of weighted
* particle attributes.
*/
constexpr uint32_t TYPICAL_PARTICLES_PER_CELL =
startPosition::RandomParameter2ppc::numParticlesPerCell;
namespace manipulators
{
struct SetIonToNeutral
{
template< typename T_Particle >
DINLINE void operator()( T_Particle & particle )
{
using Particle = T_Particle;
// number of bound electrons at initialization state of the neutral atom
float_X const protonNumber = traits::GetAtomicNumbers< T_Particle >::type::numberOfProtons;
particle[ boundElectrons_ ] = protonNumber;
}
};
using SetBoundElectrons = generic::Free< SetIonToNeutral >;
} // namespace manipulators
} // namespace particles
} // namespace picongpu
Can you please add the line in the first block that also contains the PIConGPU version number?
cc @psychocoderHPC @sbastrakov @PrometheusPi @n01r do you have any idea what might cause this?
Can't see what's off right away.
I can confirm that I see this, too in my probes output.
At the time I was blaming it on the #2763 issue, though.
I just had an idea:
Could it actually be that returning a zero weighting doesn't do anything here?
In our particle creation we reduce the number of macroparticles per cell based on the macroparticles' weighting, right?
I didn't follow it through to the end, yet but probes do not have a weighting.
In [15]: print(probes.vars.keys())
dict_keys(['position/x', 'position/y', 'position/z', 'probeB/x', 'probeB/y', 'probeB/z', 'probeE/x', 'probeE/y', 'probeE/z', 'positionOffset/x', 'positionOffset/y', 'positionOffset/z', 'particles_info'])
So even if we give the probes a weighting of 0 will they actually be removed or not initialized?
offline with @psychocoderHPC
That's what it was:
https://github.com/ComputationalRadiationPhysics/picongpu/blob/dde9b40e30d00b0a7ae27906acc91abb13485553/include/picongpu/particles/startPosition/OnePositionImpl.hpp#L116-L125
We set the numParticlesPerCell to 1u because that's the default in OnePosition but the check that realParticlesPerCell is larger than 0 is missing and it plays a role currently only after the check if the species has a weighting attribute, thus we skip that part and always initialize 1 macroparticle.
Thanks @hightower8083 for the detailed bug description!
Cool that you've found the reason so fast ! You are very welcome for the bug descriptions -- unfortunately can't really trace or fix those myself (need you guys to provide some training =) )
I am currently writing the fix. The pull request will be open after lunch.
@hightower8083 Update, the fix takes a little bit longer. My tests show something strange what I need to understand first before I can roll out a fix.
From the quote above, the logic should probably just read
// include/picongpu/particles/startPosition/OnePositionImpl.hpp
// ...
if( hasWeighting )
return startPosition::detail::WeightMacroParticles{}(
realParticlesPerCell,
T_ParamClass::numParticlesPerCell,
m_weighting
);
else
{
float_X init_weighting = float_X( T_ParamClass::numParticlesPerCell );
uint32_t const result = startPosition::detail::WeightMacroParticles{}(
realParticlesPerCell,
T_ParamClass::numParticlesPerCell,
init_weighting
);
if( result > 0u )
return T_ParamClass::numParticlesPerCell;
else
return 0u;
}
but it could be that we need to make the "MIN_WEIGHTING" constant a cleaner trait so we can ignore it for probes that have in most cases no weighting attribute... Otherwise we might end up with lacking probes where we would expect some.
I opened a bugfix for this issue in #2831
The issue should be fixed in PIConGPU 0.4.3
Most helpful comment
I am currently writing the fix. The pull request will be open after lunch.