Picongpu: installation issues
Hello,
I have a new installation of the latest dev version of PIConGPU but compiling the default LWFA mode does not work at compilation. I end up with this error. Attached is my log file
/home/quasar/src/spack/opt/spack/linux-linuxmint19-skylake/gcc-7.4.0/picongpu-develop-3datygwkvmtjbvaoihuzksvpmjxqhbxa/include/pmacc/../picongpu/plugins/PluginController.hpp(226): error #864: PerSuperCell is not a template
/home/quasar/src/spack/opt/spack/linux-linuxmint19-skylake/gcc-7.4.0/boost-1.70.0-c2sa4tgp44w22dstw23dzepx3t7yumg7/include/boost/mpl/sequence_tag.hpp(111): error #70: incomplete type is not allowed
Any idea of how can I solve the problem is appreciated. I installed with gcc 7.4 as spack load picongpu@develop +adios %[email protected]and CUDA 10 built system-wide.
The same compilation error appears for the Bremsstrahlung example.
All 17 comments
Hello @cbontoiu , thanks for your report. It seems to be a small bug accidentally introduced with the recently merged #3570. I am looking into it.
@cbontoiu could you try a fix #3589 ? It is a single-line change, so you could as well do it manually in your local dev branch.
@sbastrakov Thank you. Do you mean i go to the file include/pmacc/../picongpu/plugins/PluginController.hpp(226)
and repalce a code block with
auto gridSpacing = // SuperCellSize::toRT() * cellSize;
[]() {
auto superCellSize = SuperCellSize::toRT();
using float_type = decltype(superCellSize[0] * cellSize.x());
return std::vector<float_type>{
superCellSize[2] * cellSize.z(),
superCellSize[1] * cellSize.y(),
superCellSize[0] * cellSize.x()};
}();
mesh.setGridSpacing(gridSpacing);
?
@sbastrakov it compiles now. thank you so much
Thanks for reporting, it slipped through as our CI didn't have this combination as we are moving away from native HDF5 support (but will still do hdf5 output via openPMD API). I will close the issue once my PR is merged.
now the problem appears at the cuda test. I used tbg -s bash -c etc/picongpu/1.cfg -t etc/picongpu/bash/mpiexec.tpl /media/quasar/RawDataDisk/PICONGPU/TESTS/myLaserWakefield_01 so there should be one GPU device recognized but this line fails: cudaGetDeviceCount(&num_gpus);CUERR;
[04/15/2021 14:16:05][quasar][0]:ERROR: CUDA error: unknown error, line 279, file /home/quasar/src/spack/opt/spack/linux-linuxmint19-skylake/gcc-7.4.0/picongpu-develop-3datygwkvmtjbvaoihuzksvpmjxqhbxa/thirdParty/cuda_memtest/cuda_memtest.cu
cuda_memtest crash: see file /media/quasar/RawDataDisk/PICONGPU/TESTS/myLaserWakefield_01/simOutput/cuda_memtest_quasar_0.err
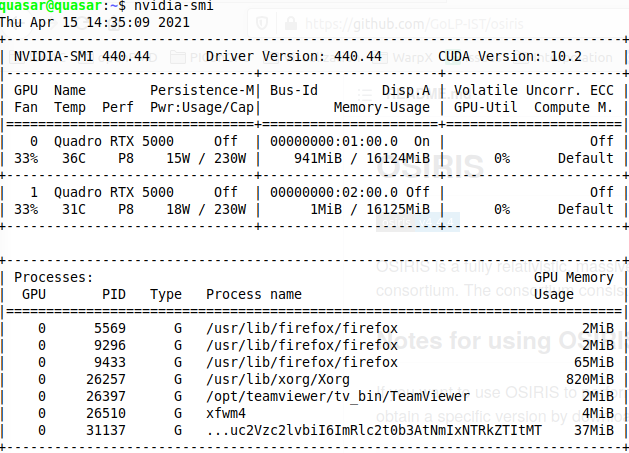
Yes, so it seems cuda_memtest did not see any GPUs. In my opinion, a way to investigate is to get on that system or node interactively and run nvidia-smi there.
i am on a desktop with two GPUs. They are fine and I used them this morning with the previous PIConGPU installation, before deciding to upgrade to the latest dev version.
Interesting. If you suspect the issue is not in the system, but something inside cuda_memtest, you could skip running it. It is a separate app ran before PIConGPU. To skip it, simply comment out its call inside your version of the .tpl file. In our reposotiry that would be e..g here, but you need to do it in your local copy of it. In your example above that would be etc/picongpu/bash/mpiexec.tpl inside your setup directory.
Great. I disabled the cuda memory test and the reading of the cfg file starts. i encounter though the error
unrecognised option '--e_phaseSpace.period'
I guess it has to do with the openPMD package for Spack?
Yes, now almost every plugin requires openPMD as it's the main way of PIConGPU output (and a couple remaining non-openPMD plugins will be converted soon). So these plugins are only enabled, and so can be used from .cfg file, when the code is compiled with openPMD API. So in my opinion it's worth always compiling PIConGPU with openPMD API, this can be done via spack.
ok, so instead of
source $HOME/src/spack/share/spack/setup-env.sh && spack load picongpu && spack load adios && export PIC_BACKEND="cuda:75" && export OMPI_MCA_io=^ompio
I should load picongpu with openPMD rather than with adios
So I did compile with openPMD loaded and the previous error disappeared but maybe at the cfg file, the GPUs failed to be recognized with this new error Unhandled exception of type 'St13runtime_error' with message 'no CUDA capable devices detected', terminating
I'm not an expert, we have a description here. I think you need to spack-install openPMD API with appropriate backends first, and then just use the variant of PIConGPU spack package with openPMD.
Okay, so somehow both cuda_memtest and PIConGPU do not see your GPUs. Maybe the issue is in CUDA version or drivers. You could try compiling some starndard CUDA examples with the same environment and see if it works.
ok, so i will modify the packages.py script to allow picongpu to install its one version of cuda rather then using the system cuda 10. I just hope the new version will be compatible with the cpp compiler.