Picongpu: ADIOS error
Hello,
I have an error when trying to load binary files from picongpu written through the adios plugin.
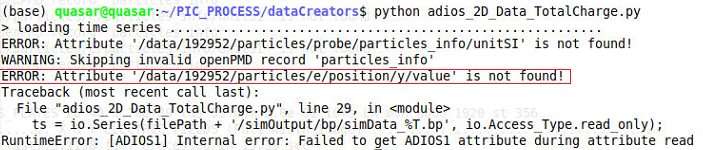
This error appears only sometimes and I cannot find a connection to the cfg file or to the model. What could it be?
The essential part of the python code is shown below:
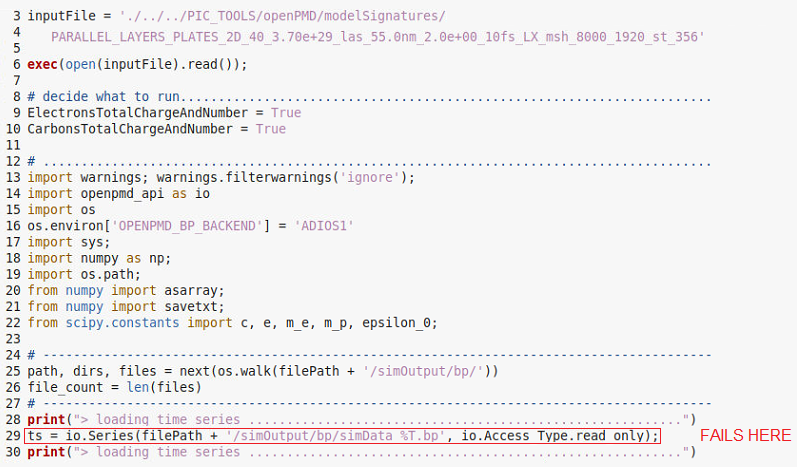
I also noticed that when loading the time series the input files are not loaded in alpahbetic order but somehow randomly. Othwerwise, I could assume that some files at the end of the simulation are corrupted and I could delete them.
Thank you.
All 20 comments
@franzpoeschel could you please help here?
Hello @cbontoiu,
can you do a dump with bpls? Something like
bpls -a simOutput/*192952*.bp
should work.
It seems that the openPMD API is looking for the attribute value of a constant dataset which happens if no actual dataset is found, so I suspect that iteration 192952 has only been partially written for some reason and the actual dataset has gone missing. Is that iteration the last iteration in your data dumps? Has your simulation finished normally or did it crash?
Maybe relevant https://github.com/openPMD/openPMD-api/issues/741
EDIT:
I also noticed that when loading the time series the input files are not loaded in alpahbetic order but somehow randomly. Othwerwise, I could assume that some files at the end of the simulation are corrupted and I could delete them.
The order is the same as the order returned by readdir on UNIX systems:
The order in which filenames are read by successive calls to readdir() depends on the filesystem implementation; it is unlikely that the names will be sorted in any fashion.
I think it is still the most likely case that the last iterations are faulty.
@franzpoeschel
Hello and thanks for your quick reply.
No that iteration is not the last one but somehwere in the middle. The last one is simData_388396.bp.dir. Last time when this occured it happened for about the 10th iteration back from the end, so I removed all those 10 files but then it loaded again and failed with the same warning about the y-value somewhere in the first half of the files, previously (assumed) correctly loaded if in alphabetic order.
The command you suggested issued in a terminal where the files are stored /simOutput/bp/ resulted in a list of attributes like the figure below but didn't solve the problem. Notice that I am using adios 1 not 2
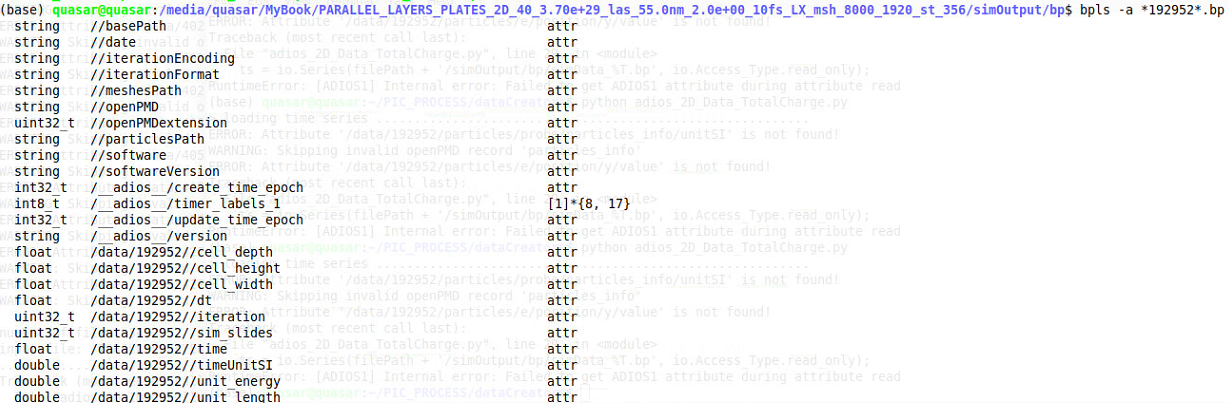
Alright, that's good to know.
The command you suggested issued in a terminal where the files are stored /simOutput/bp/ resulted in a list of attributes like the figure below but didn't solve the problem. Notice that I am using adios 1 not 2
Yes, the command was not supposed to fix the dataset, I would just like to see its complete output to verify my suspicion that the dataset has not been written. If it has been correctly written, that output should include not only attributes, but also some n-dimensional datasets. For comparison, a correctly written field might look similar to this (some minor ADIOS1 vs. ADIOS2 differences aside):
string /data/2000/fields/B/axisLabels attr
string /data/2000/fields/B/dataOrder attr
string /data/2000/fields/B/fieldSmoothing attr
string /data/2000/fields/B/geometry attr
double /data/2000/fields/B/gridGlobalOffset attr
float /data/2000/fields/B/gridSpacing attr
double /data/2000/fields/B/gridUnitSI attr
float /data/2000/fields/B/timeOffset attr
double /data/2000/fields/B/unitDimension attr
float /data/2000/fields/B/x {128, 2048, 128}
float /data/2000/fields/B/x/position attr
double /data/2000/fields/B/x/unitSI attr
float /data/2000/fields/B/y {128, 2048, 128}
float /data/2000/fields/B/y/position attr
double /data/2000/fields/B/y/unitSI attr
float /data/2000/fields/B/z {128, 2048, 128}
float /data/2000/fields/B/z/position attr
double /data/2000/fields/B/z/unitSI attr
Can you paste the complete output of the command, please?
Also, did you produce any logs while running PIConGPU?
Here is the output of the bpls command
bpls_command_result.txt
the only log of picongpu is the output text file but contains little information
output.txt
Maybe I can obtain more meaningful log if modifyng my cfg file? At the moment it looks like this
cfg_file.txt
Note that the cfg file calls another executable file called outSteps.sh which defines the variable stps as a long string with all output times. I have no doubt about that file because it always worked with both adios and hdf5
#!/bin/bash .......
segment=356
allSteps=388484
PRD=10
#..................
allFiles=$((allSteps / segment))
packages=$((allFiles / PRD))
stps=""
for ((i = 0 ; i <= $packages ; i++)); do
start=$((i*PRD*segment))
stop=$((start+4*segment))
stps+="$start:$stop:$segment,"
done
Thank you
You can enable more verbose logging at compile time, but it is probably not going to be necessary. I think I know what the error might be
int8_t /data/192952/particles/e/position/x scalar
It seems there are zero electron particles present in that iteration, is that possible? The ADIOS plugin was written before the openPMD API was able to handle zero-extent datasets, so I'm not sure it is able to handle that case. (FYI we are currently in the process of replacing the ADIOS plugin with an openPMD plugin which handles that case correctly)
@psychocoderHPC Do you have any experience with the ADIOS plugin and zero-extent datasets?
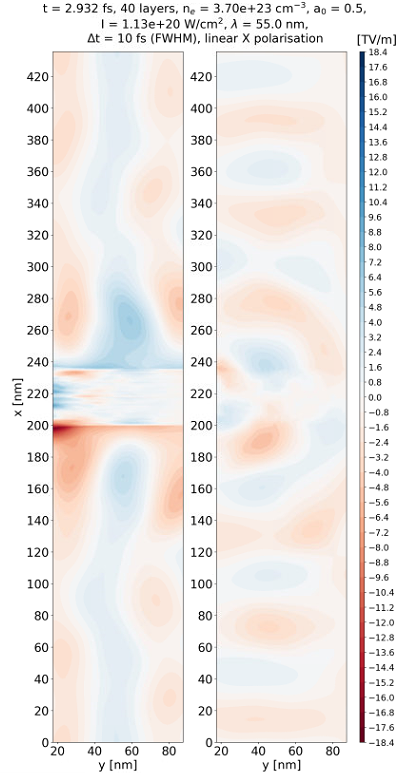
Yes it is possible and most likely the case. I used a powerful laser and due to limited size of the simulation domains the electrons are pushed out. It happened before as you can see in the figure below.
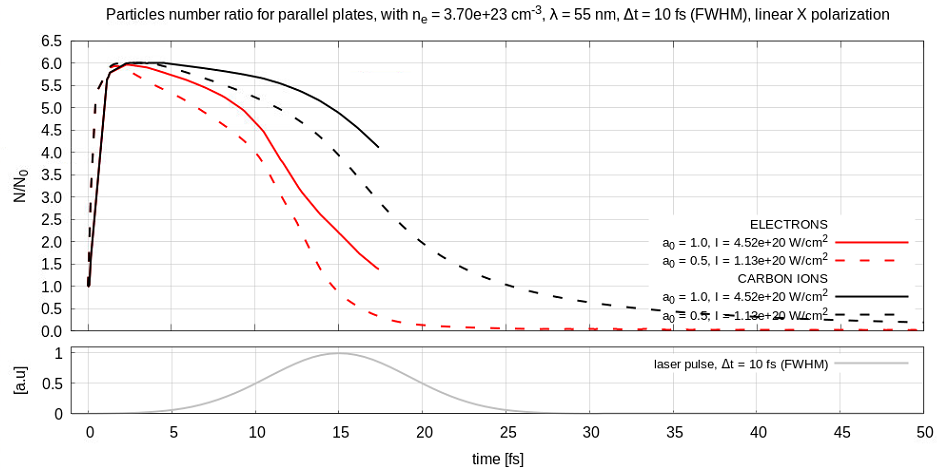
I now understand the problem but I don't know how to overcome it. Ideally I would like both large laser intensity and also be able to keep the electrons inside the simulation domain. The problem is that I have really tiny structures (0.1 nm carbon nano tube layers) so I need to resolve them with a fine mesh. Accordingly I cannot increase the domain too much. The situation (transverse and longitudinal electric fields) is like in the figure below
I don't have experience with ADIOS and zero data sets but I am happy to study the problem since it will be recurring in the future. Thank you!
A very useful feature in picongpu will be the possibility to degrade the mesh quality radially from the centre towards the boundaries of the simulation domain.
In that case, we have very likely found the problem.
Unfortunately, I think the situation requires a bugfix from our side, or we should alternatively merge the openPMD plugin soon.
Bugfix: The ADIOS plugin should use constant record components to write zero-extent datasets. (This is the way it is implemented in the openPMD API since that is the only way to do so that does not cause compatibility issues across different backends)
I don't have experience with ADIOS and zero data sets but I am happy to study the problem since it will be recurring in the future.
That question was to René, sorry for the misunderstanding. I would like to hear his feedback on the issue before proceeding, since I haven't really used the ADIOS plugin extensively.
Good! I am glad we understood the issue. I think a solution to prevent loosing all the electrons would be to use a moving window. I haven't done it so far as this requires a cluster with at least two GPUs, but I am going to try.
One little hack that might also work for you in the meantime:
- Identify an iteration that you want to inspect with the openPMD API in your Python script
- Move that iteration to a folder of its own. One iteration consists of a file ending with
.bpand a directory ending with.bp.dir - Open that iteration in your Python script in the usual way, from the new folder
This will obviously only work with iterations that do not have the issue above. You can use bpls in the described way to checked whether there is a scalar dataset.
@franzpoeschel If I understanood you correctly, It is no option to catch the "attribute not found" error in the python code of io.Series (openPMD_api) and handle it by returning an empty data set instead due to keeping the compatibility with other backends than ADIOS. Is this correct?
Do you mean catching the error right in the openPMD API or in the script that uses it?
If you are talking about catching it right in the openPMD API, I think the issue that I linked above might be of interest. Catching the error in the script that imports openpmd_api will not work, since the error occurs during construction of the series object. So you can definitely catch it, but then there's no series :/
@franzpoeschel Sorry, I overlooked the link you provided earlier - yes this is exactly what I meant :+1:
Yes, sadly catching the error in line 29 of @cbontoiu script is not an option.
I just noticed that you were maybe referring to this?
This is the way it is implemented in the openPMD API since that is the only way to do so that does not cause compatibility issues across different backends
So what you are suggesting is that the openPMD API should recognize empty datasets of this form? The reason why we are currently not doing this is because ADIOS loses information on dimensionality in that case ({0, 5} is not {5, 0} is not {0} – ADIOS only reports these datasets as scalar). But maybe that's an issue we could work around. Should I open an issue in the openPMD API to discuss this?
@franzpoeschel Yes, I war referring to that statement - if you think the issue you already linked did not represent the issue with valid but empty datasets, feel free to open an issue in the openPMDF-api repo.
@PrometheusPi It is slightly different I think, because the openPMD API should maybe actually consider this dataset as well-formed. If the issue is accepted in the openPMD API, a bugfix in PIConGPU will not be necessary.
I can remember that we had problems with zero size data in the past and we handled it somehow in PIConGPU. If I remember correctly openPMD used another way than PIConGPU to create zero size datasets.
We patched zero size writes in the past: https://github.com/ComputationalRadiationPhysics/picongpu/pull/2561
I am not sure if this is related here.
It would also be useful if you could find a way to hide the warnings/errors when loading the series.
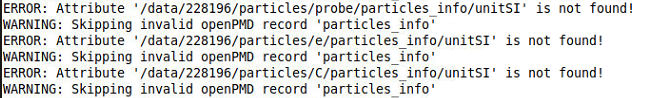
There are hundreds of lines in the terminal filled with warnings/errors like this, but the data is still loaded corectly and can be processed. Still the error on unitSI might be the explanation for the difficulties I describe in my last post of issue #3223.
Still the error on
unitSImight be the explanation for the difficulties I describe in my last post of issue #3223.
Is there an issue with unitSI? @cbontoiu could you please open a new issue and short describe what is the problem with unitSI. The issue you pointed to is already closed because it was something else and from looking to the linked issue I can not see the problem.
Most helpful comment
One little hack that might also work for you in the meantime:
.bpand a directory ending with.bp.dirThis will obviously only work with iterations that do not have the issue above. You can use
bplsin the described way to checked whether there is ascalardataset.