Picongpu: Write out data in periods
Hello,
I have a challenge for you maybe.
I know how to use the period syntax. For example with TBG_hdf5="--hdf5.period 253 --hdf5.file simData --hdf5.source 'species_all,fields_all'" I get a data file every 253 steps and, depending on the mesh, this file can be quite heavy (6GB). I typically compute this number of steps (253) such that I sample data four times a laser period, so it makes sense to see fields increasing and decreasing on both negative and positive ranges. However, most of the time, storage space is a problem. As an example, for a FWHM laser pulse duration of 25 fs I need to run the simulation during 825995 time steps (DELTA_T_SI) if I am interested in a time range as long as double the whole pulse duration, that is 6*2.35*sigmaTime and this will end up in writing out 3270 files (!).
With such a high sampling rate it does not make a huge difference if I am looking at data from t = 10*T or t = 11*T where T is the laser period. What I need is lets say sample data at (1*N+0), (1*N+1), (1*N+2), (1*N+4) next time at (2*N+0), (2*N+1), (2*N+2), (2*N+4) where N can be 10 for example and I will end up with only 327 files.
But this means that in my .cfg file I should replace hdf5.period 253 by imbricated for loops:
for(int i = 0; i < Imax; i++)
for(int j = 0; j < Jmax; j++)
hdf5.period i*N + j
or similarly with some if statements?
Could you help me along this idea, please?
An alternative would be to write some script that parses the names of the files in a folder and deletes those which do not have the right timestamp, but this would need to be a repetitive action and would need to check the file size as well to decide when it is ready to be deleted.
All 32 comments
Hi, that's already possible with the period syntax 😃. You can have comma seperated periods.
https://picongpu.readthedocs.io/en/latest/usage/plugins.html#period-syntax
Thank you. I have difficulties to understand the example
42,30:50:10: at steps 30 40 42 50 84 126 168 …
which I think would apply in my case. Let's say I want data at the following steps:
0, 117, 234, 351
1170, 1287, 1404, 1521
....
93600, 93717, 93834, 93951
They all can be obtained with S = 117 and
K = 0; -> K, K + S, K + 2S, K + 3S
K = 10S -> K, K + S, K + 2S, K + 3S
....
K = 800S -> K, K + S, K + 2S, K + 3S
What would be the correct cfg syntax to contain all these 81 batches of four samples each?
10S , 0: 3S : S , just a try
42,30:50:10: at steps 30 40 42 50 84 126 168 means every 42nd and also every 10th from 10 to 50. Btw. I think that would also dump at 0.
Regarding your example: 0:351:117,1170:1521:117,..., 93600:93951:117. You would have to write all of them out... . So some bash script that would create that string in a loop and maybe safe as an environment variable so you can use it later in the .cfg file could be a good idea.
Thank you. I actually generate and use a script for those files which I don't need. It looks like this
#!/bin/bash
file="simData_1108.h5"
rm -f $file
file="simData_1385.h5"
rm -f $file
file="simData_1662.h5"
rm -f $file
....
The only problem is that I have to run it manually from time to time but there is a solution for this as well.
The main question here is if allowing picongpu to write out uselss data files consumes a significant time.
Regards.
@cbontoiu Yes, writing out large files that you do not need will definitely slow down your simulation. Thus following the suggestion by @pordyna would definitely speed up your simulation.
Depending on the file system used and depending on how many MPI ranks you use for your simulation, writing to a file system is usually the slowest part in PIConGPU. For highly parallel file systems (many aggrigators) and many MPI ranks, our ADIOS back-end might be faster. For details please see the book section On the Scalability of Data Reduction Techniques in Current and Upcoming HPC Systems from an Application Perspective by @ax3l in Lecture Notes in Computer Science DOI: 10.1007/978-3-319-67630-2_2 or ArXiV: arXiv:1706.00522v1
If you do not need all output data, you could also reduce the content of the hdf5 output files to only contain relevant field and/or particle data. This tremendously reduces the size of these files. For that, please adjust the fileOutput.param file.
Thank you all for your help.
For reference, here is the solution which works for me following your advices
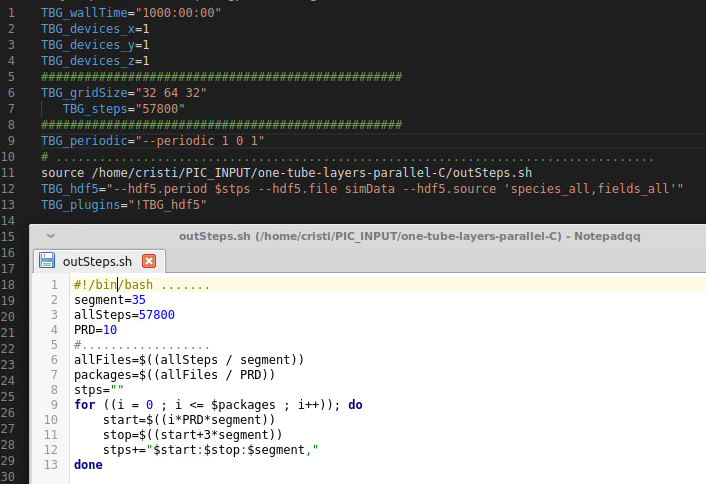
I would like to use the adios plugin instead of the hdf5 plugin but as I rememeber from a few months ago, the openPMD viewer couldn't handle files written with adios.
I do all my postprocessing using a Python script or sometimes a JupyterLab notebook with:
from openpmd_viewer import OpenPMDTimeSeries;
ts = OpenPMDTimeSeries(filePath + '/simOutput/h5/', check_all_files=False)
What would be my options for postprocessing if I switch to adios?
Regards,
Cristian
Using openPMD-enabled tools, your Python / Jupyter should literally be the same, just a different path to the time series.
Edit: ah, sorry, my comment referred to the openPMD-api itself, but the viewer may indeed have limitations.
@sbastrakov is right - currently the openPMD viewer is still not ADIOS capable (as reference @cbontoiu issue).
@cbontoiu Sorry from your image, I did not understand your solution now. Did you reduce your file output via fileOutput.param?
@PrometheusPi
There were two questions adressed in parallel. One of them was if the time needed to write out a file as big as a few GB is significant and on my computer it takes let's say 20 seconds. Actually the question was if while writing out, the code really halts solving the equations or continues at lower speed; this depends how it is threaded internaly. The other question was how to avoid writing all files if only some of them are needed. The picture shows how to source a .sh file which generates the variable $stps which is in fact a very long string based on the period syntax rules. Indeed I redued the size of the .h5 files disabling in fileOutput.param some quantities, such as the momentum. Overall the size dropped from 7GB to 3 GB for my simulations. The discovery of the possibility to script the time steps in an external file and call it from the .cfg file, really helped me, because now I don't fill up the disk with unecessry files, neither do I need to "manually" delete them from time to time. This is important since I am preparing to run on a cluster, where space is limited. Thank you.
Currently, the file output writing effectively halts the computational part of the code.
I am not sure if I understand the other question correctly, sorry if not. Hdf5 and Adios are multi-plugins (e.g. please see the note for hdf5 on the linked page), you can configure output of different quantities with different periods.
@sbastrakov
The problem is that when switching to ADIOS I don't know how would I replace the 1,2,3 commands in the image below such that I can do my postprocessing in Pyhton.

If you want to interact with the raw data instead of using the plotting functionality of openPMD-viewer, maybe just use the h5 and bp (ADIOS) agnostic API in openPMD-api for now: https://openpmd-api.readthedocs.io/en/0.11.1-alpha/usage/firstread.html
You can read your data into numpy arrays with it and do the same analysis and matplotlib plotting as you already do.
@ax3l
I have my files in the "simOutput/bp/" folder:
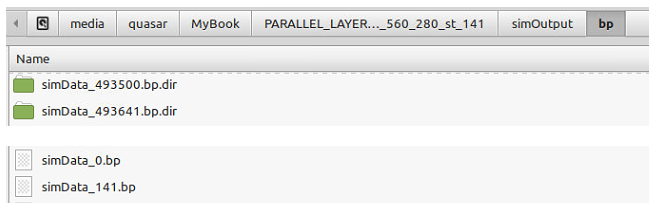
and I try to load them as a series with:

but I get the folliwing error:
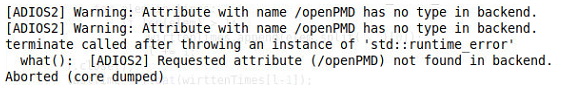
Could you guide me on what is wrong here?
I found some instructions about a json file and I wrote one like this:
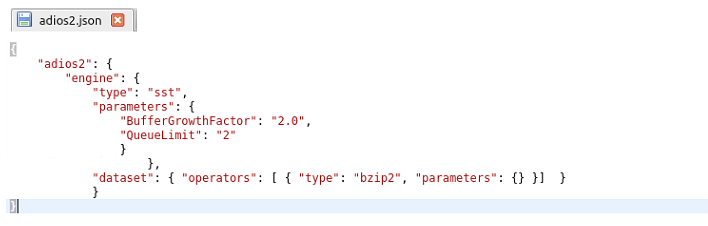
but I have no idea where should I place it and if it needs to be loaded somehow.
Thank you.
@cbontoiu I am not that experienced with ADIOS errors, but did you create the meta-data files?
(Either you do not disable meta-data journaled output despite it being slower or you create the meta data afterwards)
Yes, he created the meta files (.bp files visible in screenshot).
Oh no, that's an ADIOS2 reader error...
You have to export OPENPMD_BP_BACKEND="ADIOS1" (in bash) or in your python code just do:
import os
os.environ['OPENPMD_BP_BACKEND'] = 'ADIOS1'
to explicit open ADIOS1 files with ADIOS1. Set this prior to io.Series(...) that opens the files.
I am sorry about that, I will open an issue in ADIOS2 about this...
@cbontoiu Just as follow-up, we reported this with ADIOS2 and the next release of ADIOS2, probably v2.6.0 and later, will contain a fix to read BP3 files from ADIOS1 more robustly: https://github.com/ornladios/ADIOS2/pull/2153
Until then, please use the work-around posted in the above comment.
@ax3l Thank you,
About the meta files I am not sure because although there is a .bp file for every .bp.dir I used

in my cfg file and maybe this is not right.
I implemented your suggestion as
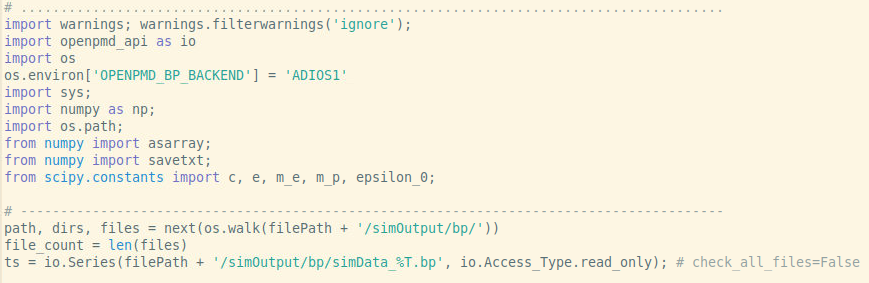
and the previous error is gone but there is and error and a warning for each data time step as

which indicates that some information is missing and files cannot be used;
In addition the typical syntax for the h5 files with openPMD does not work
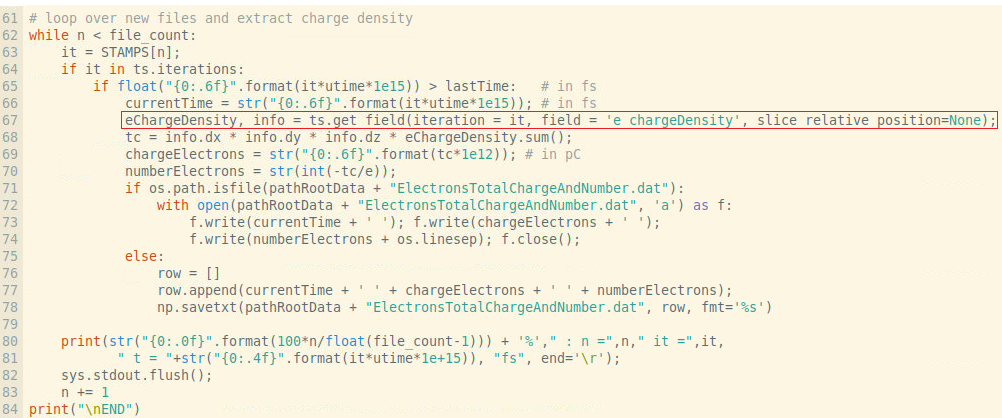
and here is the error

Any suggestion or link to the adios syntax for importing data fields and cutplanes through 3D data would be very appreciated.
Hi @cbontoiu,
if there is a .bp file for every .bp.dir than you do have ADIOS1 meta files. All good.
The "error particles_info and warning" thing you see is a inconsistency when I implemented openPMD in PIConGPU's ADIOS1 plugin. This is just ignored (but warned on) while reading in openPMD-api, you can ignore these messages, sorry that one reads ERROR instead of WARNING.
openPMD-api is not openPMD-viewer, the latter has a get_field method. Please see the Python examples of openPMD-api for read examples for fields (meshes):
https://openpmd-api.readthedocs.io/en/0.11.1-alpha/usage/examples.html#python
https://github.com/openPMD/openPMD-api/blob/dev/examples/2_read_serial.py
You can quickly get the same functionality as ts.get_field in your highlighted line by implementing a function:
import openpmd_api as io
import numpy as np
# ...
def get_field(series, iteration, field, coord=io.Mesh_Record_Component.SCALAR):
""" this is a partial implementation of openPMD-viewer's get_field with openPMD-api """
it = series.iterations[iteration]
mesh = it.meshes[field]
component = mesh[coord]
data = component[()] * component.unit_SI
grid_spacing = mesh.grid_spacing
return data, grid_spacing
now replace the two lines starting at ts.get_field line with:
eChargeDensity, grid_spacing = get_field(ts, iteration=it, field='e_chargeDensity')
tc = np.prod(grid_spacing) * eChargeDensity.sum()
General python note: no ; is needed at the end of python lines.
If you want to create cut-planes ("slices" in python), just use the eChargeDensity numpy array and use the slice syntax as discussed recently. The order of index labels is stored in mesh.axis_labels.
Does that help you? :)
We are planning to get openPMD-api below the openPMD-viewer read you already know. If you or anyone wants to contribute to this effort, these are the few files that need an update: https://github.com/openPMD/openPMD-viewer/tree/dev/openpmd_viewer/openpmd_timeseries/data_reader
Back to .bp meta-files: ADIOS2 (2.5.0+) does not require these anymore, you can just open the directory directly "as if it were a .bp-ending file". As soon as the new implementation lands in PIConGPU, you can take advantage of that #2966 . Also, the warnings mentioned above will go away.
@ax3l
Thank you so much for these advices. I am closer now to the solution but I still have troubles. Here is my code
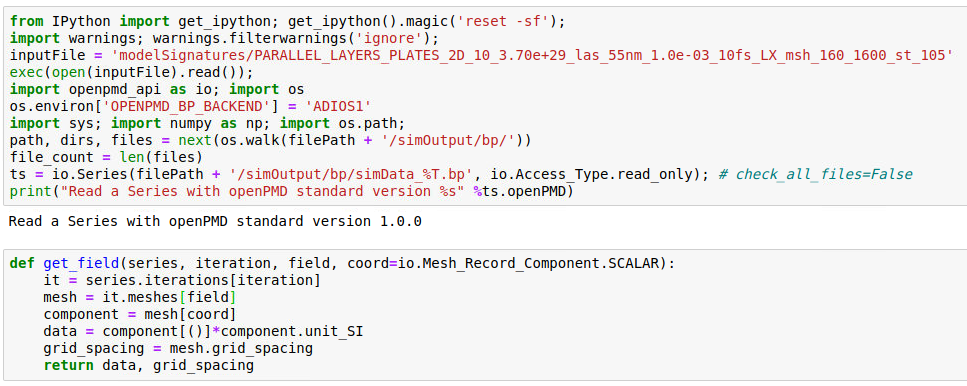

but when printing data I get sometimes:
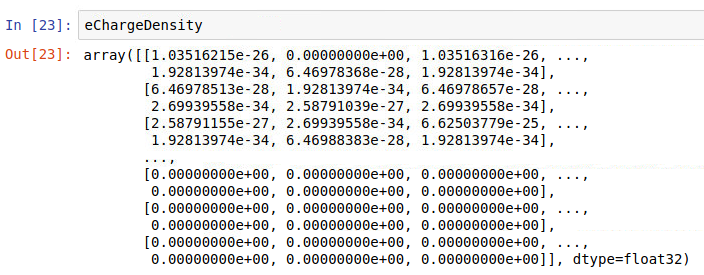
and at other times, for the same iteration (it = 0):
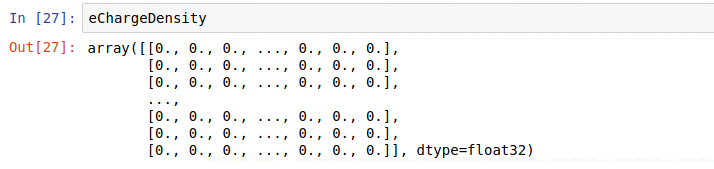
for the same iteration and this explains why sometimes I get a large number for the sum and at other times I get a zero (first column is time in fs)
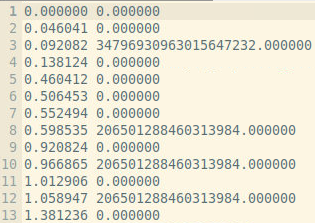
I also notice that the mesh units are not what I expected. I know that dy = 0.189 nm and dx = 0.182 nm but the code gives me
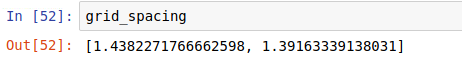
Perhaps I fail understanding how to tread my 2D data packages or maybe the units are messed up somehow. My code is here attached
chargeScript.txt
and here is the data only for the first time step (it = 0) with its meta file archived together.
data.zip
Any help is appreciated.
Thank you,
Cristian
@cbontoiu the grid_spacing value you see is given in the PIConGPU unit system. You need to multiply it with its unitSI attribute (or alternatively with the unit length attribute).
However, the ratio 189nm/182nm and 1.438227/1.3916333 are still of by 0.4%. That's a bit confusing.
If the code you gave for it = 0 results in non-zero charge values (as seen in the first array you provided) the initialization is not charge neutral and thus the field solver has a wrong starting condition. What are the differences between those setups where at ts=0 the charge is not zero everywhere and those where it is?
Furthermore from the values you showed in the first array, the individual contributions are << 1e-20, but the sum over all array values is >1e18. This more than 38 orders of magnitude difference can only occur if your 2D simulations has 1e19 cells in both dimension (which would never fit into memory 😉) or if you have much larger charge values in your array. Perhaps identifying these extreme charge values might help identifying the origin of this strange behavior of the total charge.
@PrometheusPi Thank you. The number of cells was x = 160 and y = 1600, along the tubes and dimensions are here (longitudinally the plates extend about 300 nm):
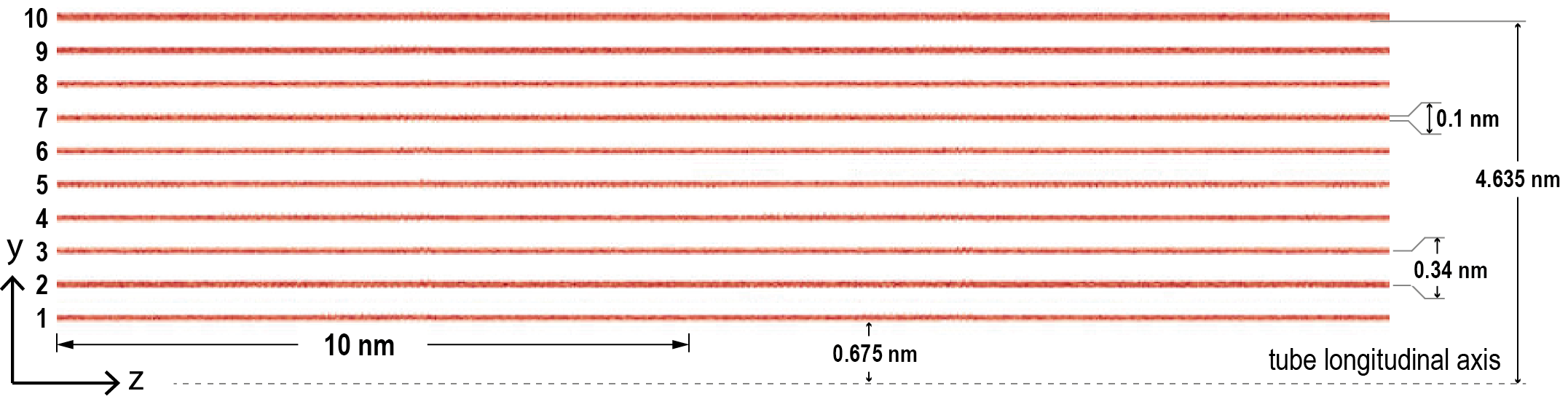
The confusion comes from the fact that with h5 files I never had this troubles for this model type, though I used better mesh. I tried to simply change the cfg file and use the hdf5 plugin but got the error below
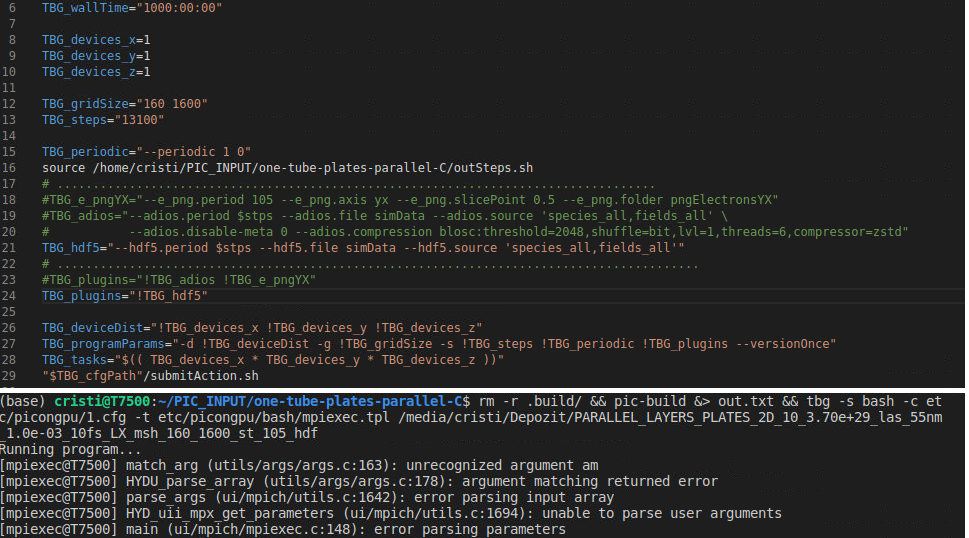
Maybe this is an indication. I will try running exactly this model on a different computer with the hdf5 plugin to check again. Maybe it is a problem of openmpi now but there has never been an issue with charge calculation before (hf5 files) and whatever the mesh is I would expect the total charge to be about 4 fC in the beginning as it can be seen in this figure
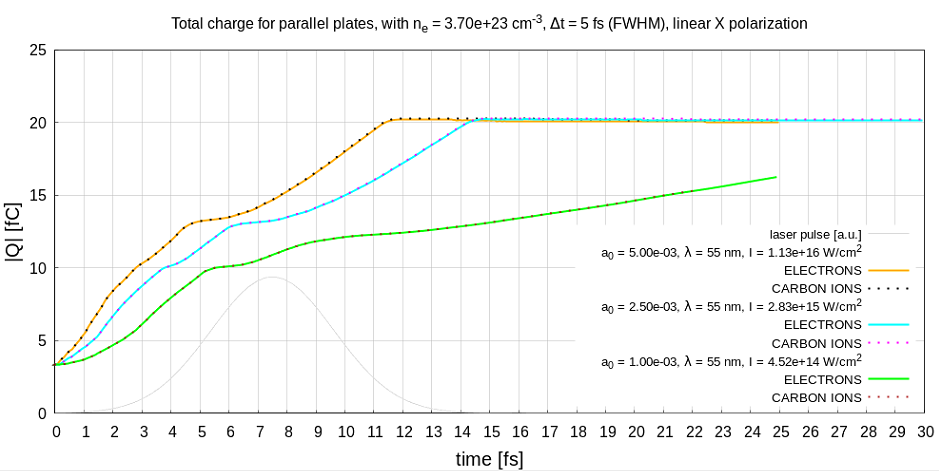
That you never encountered these charge issues with hdf5 and only now with adios seems to be an interesting indicator. I am very interested in your comparison.
This is just now only guessing what the error might by: I assume you initialize your particles charge neutral, thus positively and negatively charged particles should be places exactly on top of each other, correct? (Or do you initialize neutral atoms.) You could check the particles them self in the adios files. If they are not distributed neutrally, then there is an initialization problem. If they are placed correctly, then we might have an issue with field initialization in PIConGPU.
That you never encountered these charge issues with hdf5 and only now with adios seems to be an interesting indicator. I am very interested in your comparison.
This is just now only guessing what the error might by: I assume you initialize your particles charge neutral, thus positively and negatively charged particles should be places exactly on top of each other, correct? (Or do you initialize neutral atoms.) You could check the particles them self in the adios files. If they are not distributed neutrally, then there is an initialization problem. If they are placed correctly, then we might have an issue with field initialization in PIConGPU.
no, I initialize an electron for each C+ ion
@ax3l @PrometheusPi
OK, so there is difference for the same model between extracting data from h5 files and bp files, as I get total charge data like this:
time [fs] totalElectronsCharge[fC] electrons
0.000000 -2.600602 16231
0.046041 -2.600602 16231
0.092082 -2.600602 16231
0.138124 -2.600602 16231
0.460412 -2.600602 16231
0.506453 -2.600602 16231
..............
Notice that due to the much lower mesh size (that is 10-fold) I get half 2.60 fC instead of 3.38 fC charge before, but still there are no zero values and neither very large. More exactly the mesh was before uXspace = 1.829375e-11m, uYspace = 1.890625e-11 m and now it is uXspace = 1.829375e-10 m, uYspace = 1.890625e-10 m
I attach the full picongpu model:
one-tube-plates-parallel-C.zip
and here is the script which extracts the charge and the number of particles
the other script (using adios) is uploaded above under the name chargeScript.txt
With that script I checked again the total charge this time using a different computer and obtained confusing, random (?) results:
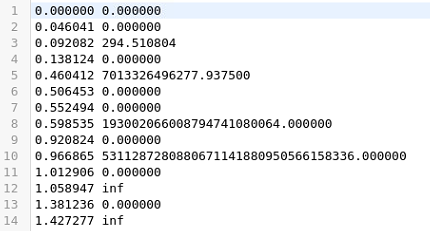
and
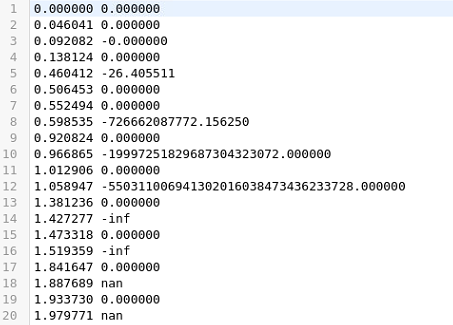
There must be something wrong in my script. Please let me know if you can advice me.
Regards,
Cristian
Hi @cbontoiu , I think that you are not flushing your series in your script. In openpmd-api you have to flush the series to actually completely copy the chunks of data to the initialized numpy arrays. You could be just reading random data that just happened to be in the buffer at the moment. Try adding ts.flush() ad the end of your get_field function.
Please ignore this if you are already doing it somewhere else and I just didn't find it :)
ts.flush()
Great! That is the problem! Thank you. I manage to get the data below
0.000000 -54686.503291
0.046041 -54686.503291
0.092082 -54686.503291
0.138124 -54686.503291
0.460412 -54686.499382
0.506453 -54686.503291
0.552494 -54686.503291
Now my problem is related to units or to the concepts of sum() and prod() I think because instead of -2.6 [fC] I get -54686.5 [C/m ?] which by multiplication with 0.1e-09 [m] would give me 54.6 [fC] .
The key part of the script is here

I just checked what comes from the product of the grid units and what comes for the sum of charge density
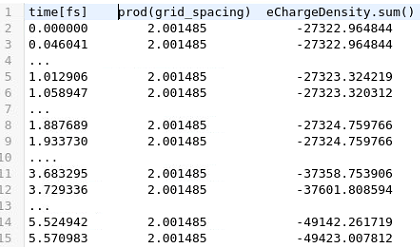
The equivalent with hdf5 is
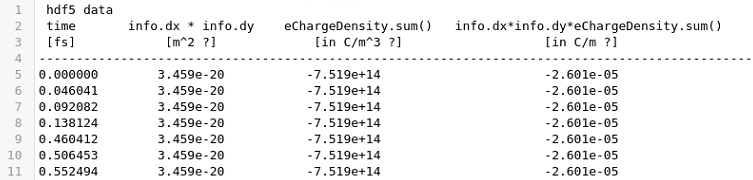
I normally multiply the last column by 0.1 nm the (3rd dimension) to account for one atom thickness to get the the charge in Coulombs
@cbontoiu Thanks for posting this update - the ratio change is indeed strange.
@cbontoiu Thanks for posting this update - the ratio change is indeed strange.
@PrometheusPi Sorry I had to delete my last message about the mesh ratio because I am not sure about the correctness. It might have happened that I misinterpreted the input and the ratio was really roughly 1 for the binary data and 10 for the hdf data. However the problems I have as shown above are still not solved. Thank you.
@ax3l @PrometheusPi
Hello,
I finally achieved convergence between hdf5 and adios results with some minor modifications
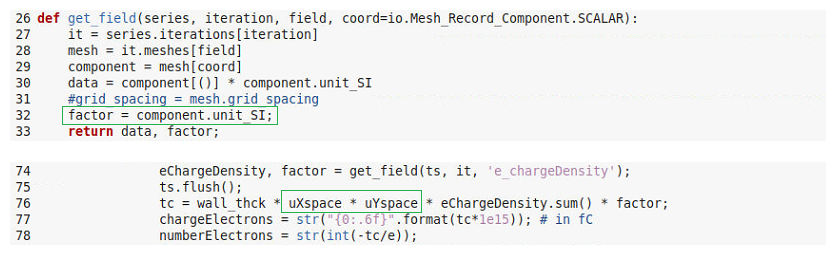
but this means I cannot access the grid_spacing anymore and I need to rely on my calculated mesh units for x and y. I just wanted to share this with you because grid_spacing coming for the get_field definition never made sense for me and maybe you want to check.
Here is the result
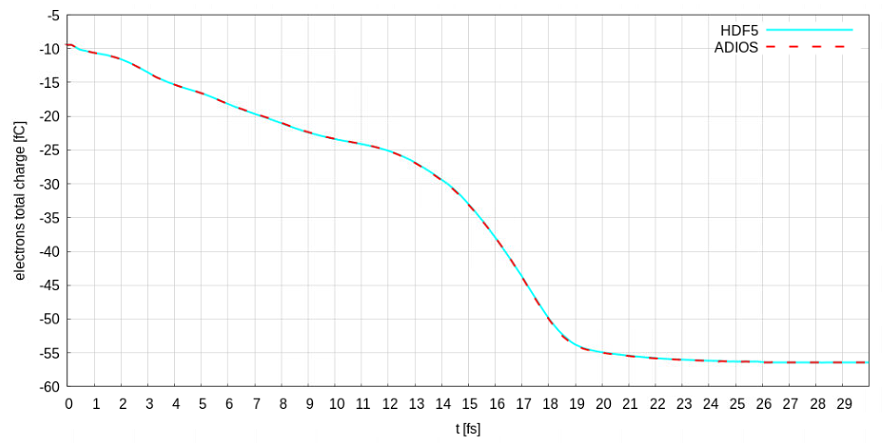
Another thing: a paper based on PIConGPU was submitted by our group and will be published in the proceedings of EAAC 2018 by Elsevier/ScienceDirect. Maybe you want to prepare a space where you can list publications making use of PIConGPU
Regards and many thanks for your help
Cristian
Hi @cbontoiu
Thanks for posting the update and great it works for you (somehow) now. I am sorry, I do not understand the code you provided - thus I can not follow your issue with grid_spacing.
Thanks for sharing your idea on a public list of papers based on PIConGPU. I think this is a great idea. @ComputationalRadiationPhysics/picongpu-developers what do you think. We could add this in the ReadTheDocs. I will open a issue for voting in this.
Most helpful comment
@cbontoiu Yes, writing out large files that you do not need will definitely slow down your simulation. Thus following the suggestion by @pordyna would definitely speed up your simulation.
Depending on the file system used and depending on how many MPI ranks you use for your simulation, writing to a file system is usually the slowest part in PIConGPU. For highly parallel file systems (many aggrigators) and many MPI ranks, our ADIOS back-end might be faster. For details please see the book section On the Scalability of Data Reduction Techniques in Current and Upcoming HPC Systems from an Application Perspective by @ax3l in Lecture Notes in Computer Science DOI:
10.1007/978-3-319-67630-2_2or ArXiV: arXiv:1706.00522v1If you do not need all output data, you could also reduce the content of the hdf5 output files to only contain relevant field and/or particle data. This tremendously reduces the size of these files. For that, please adjust the
fileOutput.paramfile.