Picongpu: cannot use more cells in the transverse directions
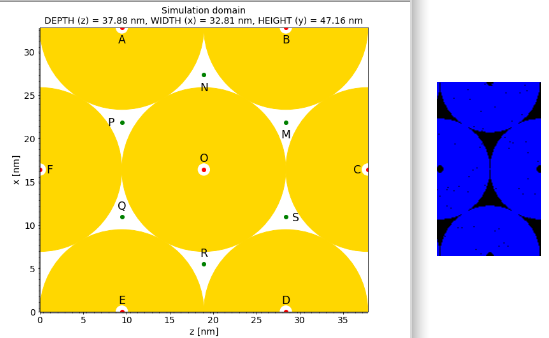
I want to simulate only half (Z) of a tubular structure. It is designed in density.param and grid.param as shown in the yellow picture below. However in the 1.cfg file I specify only half of the cells along the z-axis making sure that this number is multiple of 8 because using SuperCellSize = mCT::Int< 8, 8, 8 >;. This works for the number of X:Y:Z cells 160:400:200(100) as shown by the blue picture.
However I need more resolution in the X and Z directions so I tried 320:400:384(192) and there is no density appearing in the png figures. This configuration is attached in the .zip folder. I tried changing 8 to 16 as using SuperCellSize = mCT::Int< 16, 16, 16 >; but then the error was
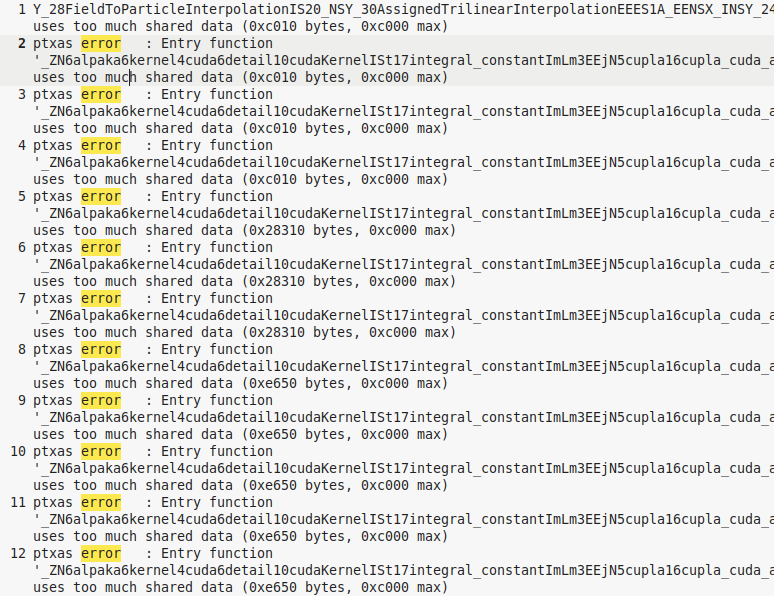
Is it something related to the number of macroparticles? Do I need to use something else instead of static constexpr uint32_t numParticlesPerCell = 6u;? What can I try else? Please have a look.
Thank you.
All 3 comments
using SuperCellSize = mCT::Int< 16, 16, 16 >; is not possible if you run on a NVIDIA GPU. X*Y*Z must be <= 1024. The supercell size is also controlling the caching within the GPU so I expect that even mCT::Int< 16, 8, 8 > is to large.
I would suggest to use not more than 8x8x8 cells for a supercell, beside this out of shared memory compiler error you showed it could have negative performance effects.
Is it something related to the number of macroparticles? Do I need to use something else instead of static constexpr uint32_t numParticlesPerCell = 6u;? What can I try else? Please have a look.
The error shown is coming from the supercell size.
However I need more resolution in the X and Z directions so I tried 320:400:384(192) and there is no density appearing in the png figures.
I have not reproduced this but could you please check your density profile and check if the density problem is coming from the hard coded sizes.
//##################################################
constexpr double WAVELENGTH = 282.192e-09;
constexpr double hTL = 0.2*WAVELENGTH; // hollowtube length
constexpr uint32_t CELLSX = 320;
constexpr uint32_t CELLSY = 400;
constexpr uint32_t CELLSZ = 384;
constexpr double OUTER = 0;
Hello, Thank you for your answers. The error was due to the MIN_WEIGHTING parameter in the particle.param file. With more mesh cells the actual weigh went below the MIN_WEIGHTING value.
Most helpful comment
using SuperCellSize = mCT::Int< 16, 16, 16 >;is not possible if you run on a NVIDIA GPU.X*Y*Zmust be <=1024. The supercell size is also controlling the caching within the GPU so I expect that evenmCT::Int< 16, 8, 8 >is to large.I would suggest to use not more than
8x8x8cells for a supercell, beside this out of shared memory compiler error you showed it could have negative performance effects.The error shown is coming from the supercell size.