Picongpu: Data storing with ADIOS
I am looking for methods to compress my output data. So far I worked with TBG_hdf5="--hdf5.period 2 --hdf5.file simData" and I am aware about the possibility to write only fields and even further only the electric fields and possibly use it as a multi-plugin. I just discovered that it is possible to output particles to one disk and fields to another disk. That is not all hdf5 in a single simOutput folder but in two independent folders. I mean using something like this
TBG_hdf5_A="--hdf5.period 10 --hdf5.file /home/quasar/testA/simData --hdf5.source 'fields_all'"
TBG_hdf5_B="--hdf5.period 20 --hdf5.file /home/quasar/testB/simData --hdf5.source 'particles_all'"
Further, I also managed to use the ADIOS plugin for the first time and it is impressive that using compression a snapshot is reduced to ~20MB instead of a ~1GB as it HDF5. I disabled the the meta information as --adios.disable-meta 1 and successfully run bpmeta simData_xxx.bp.
1) What can I do with these bp.0 files? Are they useful only for continuation of the simulations, using the checkpoint plugin or I can retrieve my data from them using the OpenPMD viewer or something else?
2) Is there a script that I can run in order to apply the bpmeta operation to all files at once? Perhaps I can create one. Do I have to apply the bpmeta operation to all files in order to get some kind of consistency check file at the end?
3) What other route do you suggest in order to encapsulate the data but still be able to read it?
Thank you
Cristian
All 13 comments
I also think there is an error at the bottom of this page
https://picongpu.readthedocs.io/en/0.4.3/usage/plugins/hdf5.html
Namely to swap the description of the two plugins.
.......................................................................
--hdf5.period 128 --hdf5.file simData1
--hdf5.period 1000 --hdf5.file simData2 --hdf5.source 'species_all'
creates two plugins:
dump all species data each 128th time step.
dump all fields and species data (this is the default) data each 1000th time step.
I also think there is an error at the bottom of this page
https://picongpu.readthedocs.io/en/0.4.3/usage/plugins/hdf5.html
Namely to swap the description of the two plugins.
.......................................................................
--hdf5.period 128 --hdf5.file simData1
--hdf5.period 1000 --hdf5.file simData2 --hdf5.source 'species_all'
creates two plugins:
dump all species data each 128th time step.
dump all fields and species data (this is the default) data each 1000th time step.
The description in our documentation is correct. The behavior is very confusing because the command line parser framework we are currently using not support hierarchical options.
Therefore the command line in the example is equal to --hdf5.period 128 --hdf5.file simData1 --hdf5.source 'species_all --hdf5.period 1000 --hdf5.file simData2
The option --hdf5.source is only used once and is therefore part of the first instance of the multi plugin.
@psychocoderHPC but then I think the description is confusing at least and we should move --hdf5.source 'species_all' to the first line of the sample?
@cbontoiu both HDF5 and ADIOS output should be openPMD-compatible, so tools like openPMD-viewer should work. This concerns both checkpoints and regular output.
@cbontoiu regarding your question 1.:
You still need the bp.0 file as your actual data is in there. The .bp files are just meta files that are required to actually access you data.
Also, we usually use --adios.disable-meta 0 (and not 1) in order to generate the meta files directly while the simulation runs. We only turn this off when running on the largest systems and writing hundreds of Terabytes of data because then the generation of the .bp meta files can take minutes which can be a significant amount of time from our precious compute time on the system. Considering that you run locally or on smaller clusters where your output is considerably smaller, your time loss is negligible and generating the *.bp files automatically while the simulation runs is a lot more convenient.
If you need a script, you may create one along the lines of the following, which was first created by @ax3l (as far as i remember)
#!/bin/bash
# 1. Go to a directory where you have PIConGPU ADIOS output, e.g. "simOutput/bp/"
# 2. Create a list of filenames matching your pattern, e.g. "simData"
f=$(\ls | grep simData)
# 3. Run bpmeta on all of them
for fdir in $f; do
# The naming pattern is for me "simData_fields_all_<step>.bp.dir"
# We cut at every "." and only select the field 1 (till <step>) and 2 (bp) so that we do
# bpmeta "simData_field_all_<step>.bp"
file=$(echo $fdir | cut -d . -f1,2)
bpmeta -z $file
done
Please be aware, that the -z option to bpmeta made some problems in the past with empty datasets and restarting from checkpoints, see (https://github.com/ComputationalRadiationPhysics/picongpu/issues/2953#issuecomment-489585221). (Which is another reason for being lazy and let PIConGPU directly generate the *.bp)
@psychocoderHPC but then I think the description is confusing at least and we should move
--hdf5.source 'species_all'to the first line of the sample?
The example is fine, it should show exactly this strange behavior. Therefore we have the comment In the case where a optional parameter with a default value is explicitly defined the parameter will be always passed to the instance of the multi plugin where the parameter is not set. e.g. before.
I am not sure how we can describe it better, any suggestion is welcome!
@cbontoiu both HDF5 and ADIOS output should be openPMD-compatible, so tools like openPMD-viewer should work. This concerns both checkpoints and regular output.
It actually does not work for me and I don't know why. Attached are snapshots of the Jupyter import statements and of my files produced with the ADIOS plugin Could anyone help me to load these files in the openPMD_Viewer. This setup used to work fine with HDF5 files as written by TBG_hdf5="--hdf5.period 100 --hdf5.file simData" but now I used TBG_adios="--adios.period 100 --adios.file simData".
An additional question is if compressed ADIOS files generated with TBG_adios="--adios.period 100 --adios.file simData --adios.compression zlib" can also be imported in openPMD_Viewer.
Thank you
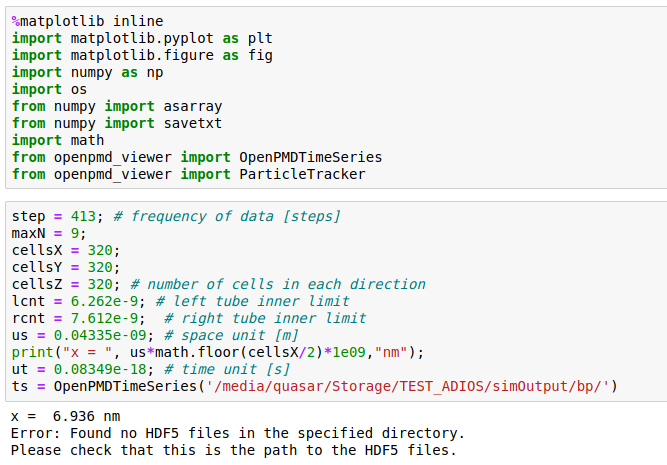
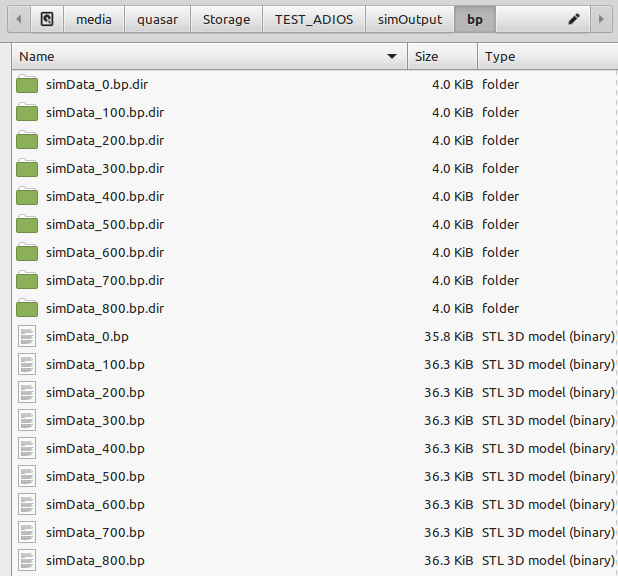
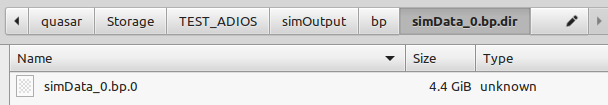
@cbontoiu could it be that your openPMD-viewer (or openPMD-api) was built without ADIOS? Not sure if there is an easy way to check, but perhaps their documentation offers it.
openPMD-viewer is not yet using openPMD-api and therefore the viewer supports HDF5 only. We plan to change that soonish, please use HDF5 in the meantime.
bpmeta's bugs from the past are fixed in latest adios 1.13.1.
Script: add a check if the file exists before calling it for all files again to safe time.
if [ ! -f $file ]; then bpmeta .... fi
openPMD-viewer is not yet using openPMD-api and therefore the viewer supports HDF5 only. We plan to change that soonish, please use HDF5 in the meantime. bpmeta's bugs from the past are fixed in latest adios 1.13.1. Script: add a check if the file exists before calling it for all files again to safe time. if [ ! -f $file ]; then bpmeta .... fi
Thank you. In conclusion, without ADIOS there is not better way to compress ouput files.
but then I think the description is confusing at least and we should move --hdf5.source 'species_all' to the first line of the sample?
Actually, that documentation example tries to document this exact confusing behaviour by posting that example. Maybe someone can re-phrase the "Note" text and add a first, simple correct example before the confusion-causing example?
Thank you. In conclusion, without ADIOS there is not better way to compress ouput files.
Our goals are to provide the same tooling for ADIOS files since this methods scales and compresses well at scale as well as in smaller sims. We'll hopefully be there soon with the viewer too - so you have a seamless experience with both outputs (openPMD-api already works, as well as native ADIOS python bindings for post-processing).
Until then, if you have HDF5 files already on disk you can re-pack them after a run by re-packing the files with transparent, in-format compression:
https://github.com/ax3l/cluster-scripts/blob/master/bash-h5compress.sh
This usually saves a factor 3-5x on disk for me. Maybe that's somewhat helpful in the meantime for you?
Actually, that documentation example tries to document this exact confusing behaviour by posting that example.
You are right. But it became clear to me only after reading this piece for the 3rd time. The first two I thought it just explained that one could run 2 different instances of the plugin with different source data. (Ofc could also be that i should have just read more carefully)
Nah, I think a "that's how it should be used" example and then "careful, don't make this mistake" with the current example could make this section easier to read.
Most helpful comment
openPMD-viewer is not yet using openPMD-api and therefore the viewer supports HDF5 only. We plan to change that soonish, please use HDF5 in the meantime.
bpmeta's bugs from the past are fixed in latest adios 1.13.1.
Script: add a check if the file exists before calling it for all files again to safe time.
if [ ! -f $file ]; then bpmeta .... fi