Picongpu: particle tracking with openPMD
Hello,
I am learning to use openPMD viewer using picongpu h5 data. More exactly those produced in the LWFA example. I load data as
ts_3d = OpenPMDTimeSeries('./../../PIC_RUN/myLaserWakefield_02/simOutput/h5/', check_all_files=False)
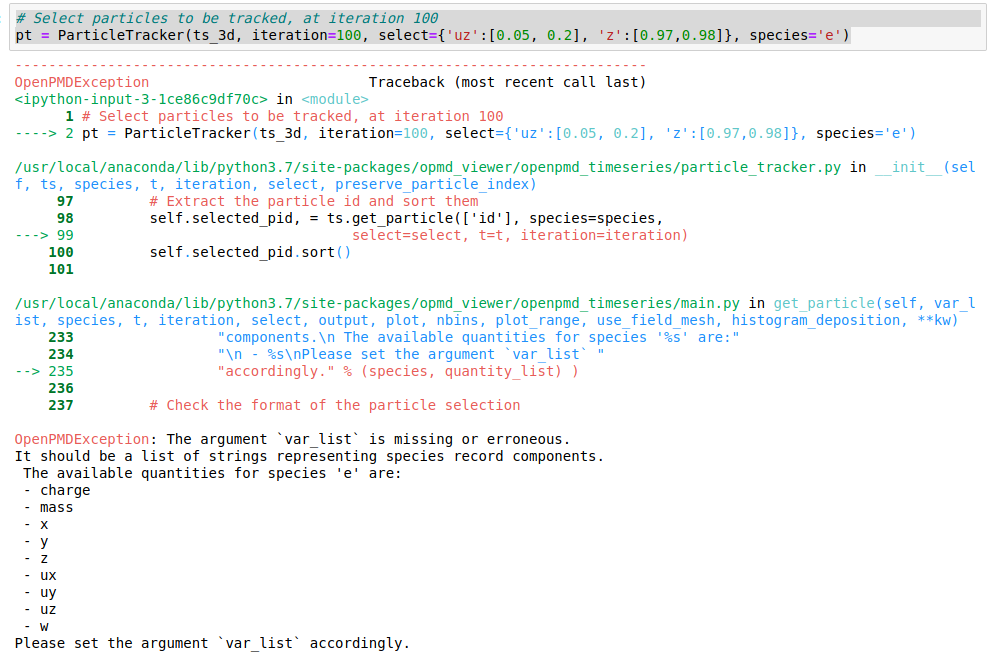
and everything goes smoothly, getting various field and phase space plots until I reach the ParticleTracker part. The error is attached in the figure. I tried various changes to the syntax and to the order of the argument without any luck.
Could anyone advice me on the tracker, please?
Kind Regards
All 20 comments
Hi,
probably the right place to ask questions about openPMD-viewer is its own issue tracker here: https://github.com/openPMD/openPMD-viewer/
@RemiLehe can probably help with it, looks like you are not passing a valid argument to ParticleTracker. Maybe try ParticleTracker? in a new Jupyter cell to see its API doc string?
Oh, it could be the case that the electrons species e that you are trying to track has not particle-id argument yet.
In order to add an id attribute to particles, go to speciesDefinition.param and add another attribute for ids to the list:
/** describe attributes of a particle*/
using DefaultParticleAttributes = MakeSeq_t<
position< position_pic >,
momentum,
weighting,
particleId
>;
Yes, @ax3l is absolutely right. The ParticleTracker requires the id of the particles to be written to the openPMD file.
This is mentioned in the docstring of ParticleTracker, but I agree that it is not easy to find, and that it should be better documented:
https://github.com/openPMD/openPMD-viewer/blob/7029c7c19ec02c108d00e5b8ac04c13a6f276c95/opmd_viewer/openpmd_timeseries/particle_tracker.py#L45
Thanks for your help.
I rerun the model with using DefaultParticleAttributes = MakeSeq_t< position< position_pic >, momentum, weighting, particleId >; as suggested and I can see the field particleId through the openPMD GUI. It is there but I don't know how to set up a correct call in my notebook.
I attach snapshots of a few cases that I tried. The online example on mybinder would correspond to my case 6 and my syntax for importin data is data = OpenPMDTimeSeries('./../../PIC_RUN/myLaserWakefield_01/simOutput/h5/', check_all_files=False)
Please advice me how to correctly call the method.
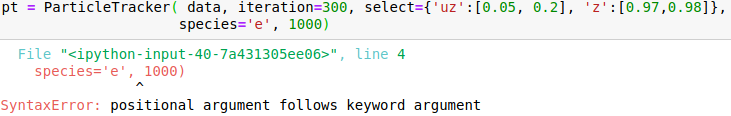
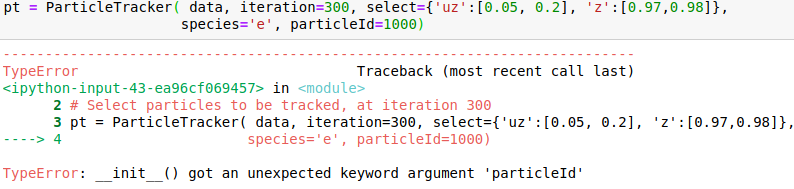
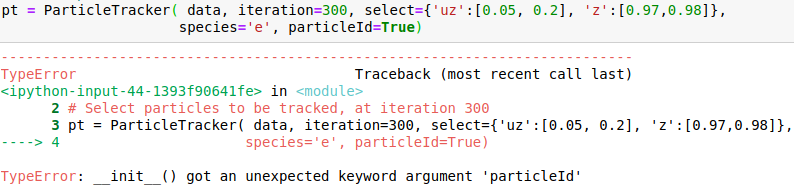
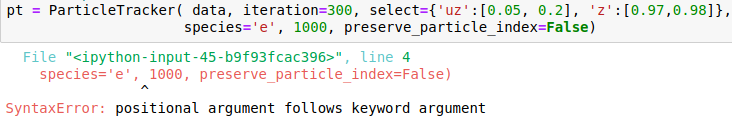
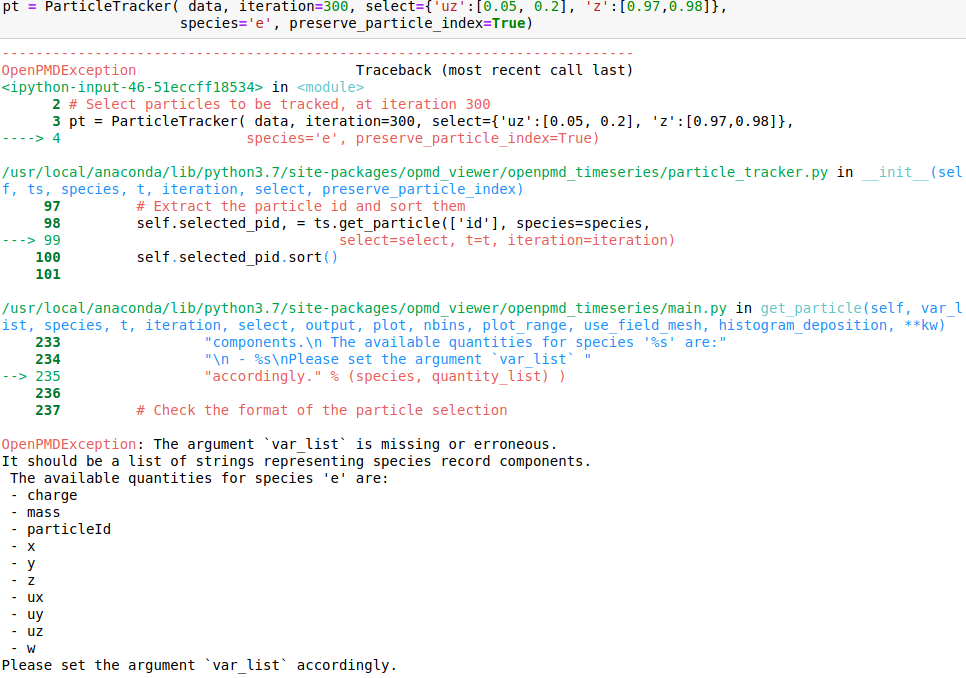
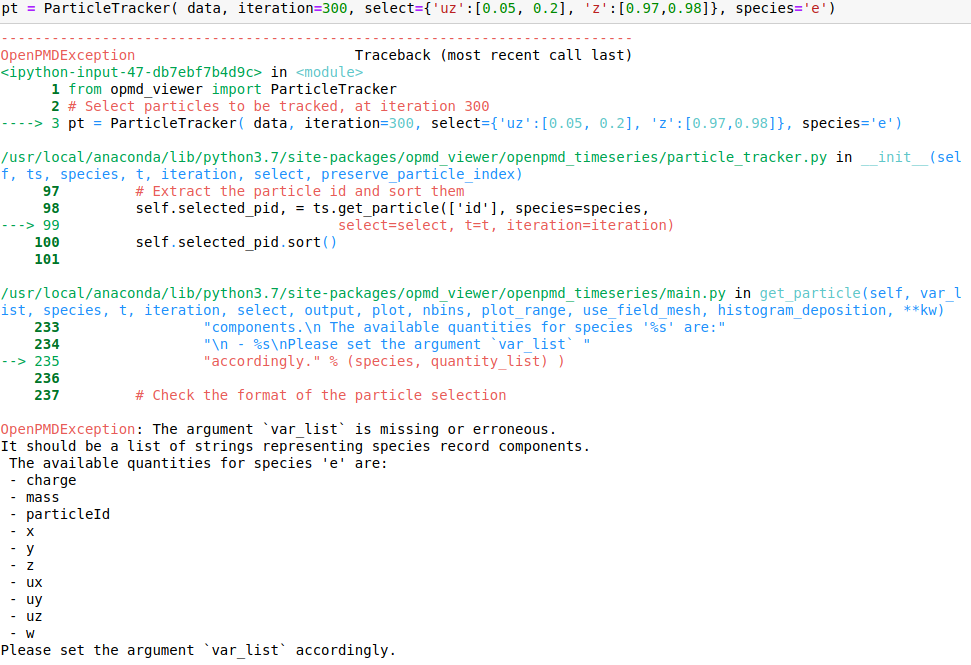
Hello @cbontoiu . I am not an expert in this, but it seems in your earlier calls some issues are caused by using positional (unnamed) parameters after the named ones, which is not allowed. E.g., after you have species='e', all the following parameters also have to be named, so given in the same form with explicitly giving the name. The names have to match the interface, so e.g. particleId is not a valid parameter name for this function. The last call is syntactically correct, and I can't see why it causes a crash.
The comment about particle id earlier in this discussion referred to the fact these ids must be present in the file you are reading (and for this perhaps configured in the .param file as @ax3l wrote). From the output it seems like particleIds are already there.
@sbastrakov @ax3l Actually it seems to me that @cbontoiu's last command line is valid and should have worked
(pt = ParticleTracker(data, iteration=300, select={'uz':[0.05, 0.2], 'z':[0.97, 0.98]}, species='e'))
I am wondering if, in fact, the problem is that PIConGPU maybe does not fully comply with the openPMD standard. More specifically, PIConGPU stores the particle ID as particleId ; but the openPMD standard expects the particle ID to be stored as id:
https://github.com/openPMD/openPMD-standard/blob/latest/STANDARD.md#records-for-each-particle-species
and openPMD-viewer does rely on this convention.
Does this seem correct?
Yes, that last also looks good to me.
I think you are right about particleId in PIConGPU. This can actually be "overloaded" with a trait specification.
There is no trait for this yet, but you can generalize the implementation of the static getName() method in include/pmacc/identifier/value_identifier.hpp to support something like this, e.g. a pmacc::getName() free standing function with ADL lookup or similar.
I see that the header of speciesDefinition.param already includes pmacc/identifier/value_identifier.hpp but what can I do exactly to get the particle ids?
Thank you.
How can I change the color style in density plots? I tried adding colorbar='rainbow' but it is not accepted.
I am not sure where exactly did you try this parameter. It might be that in the viewer it is called cmap, not colorbar. At least this name is used in this example.
Generally, in python one might call help([function name or object])
I have 3 additional questions here.
1) I define three types of particles and use them in speciesDefinition.param which is attached
I then import data in openPMD viewer as
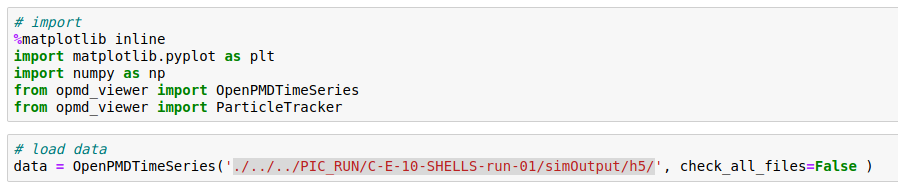
But it appears that only the probes are stored. Is this right?
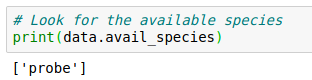
How can I have a figure/data of all electrons or Carbon ions at a given time snapshot? Can this be achieved through the Probes or they are only for mapping the fields?
2) Where can I find the full API which allows me to get the most of the graphics options. For example I don't find how should I increase the size of the images generated in the Jupyter notebook. How can I add labels, annotations, gridlines etc?
3) What would be the syntax (or maybe the easiest way out of the openPMD viewer) to export field and particle data along a line or across a slice? In this way I could use date with Gnuplot for example. My data source is in hdf5 format as produced by
TBG_hdf5="--hdf5.period 500 --hdf5.file simData --hdf5.source 'species_all,fields_all'"
but maybe there are other ways to store/extract data.
Thank you
- In terms of output, all 3 of your species are independent, so it's not possible to restore electrons or carbon from the probes. It might be that you only enabled output of the probes. From the
TBG_hdf5="--hdf5.period 500 --hdf5.file simData --hdf5.source 'species_all,fields_all'"line it looks like you request to output all species. Could you check thefileOutput.paramfiles and see the "value" ofFileOutputParticles? To output all species, it has to beusing FileOutputParticles = VectorAllSpecies;for all species or an explicit list of species you want, likeusing FileOutputParticles = MakeSeq_t< Electrons, Carbon, Probes >;(or you pick a subset of those)
- For getting the API docs, please use
help([name of object, function or basically whatever])or[name of object, function or basically whatever]?. The latter is demonstrated in this example byts_2d.get_mean_gamma?. Generally, for openPMD-side API questions, perhaps their repository is a more suitable place. Btw, they support running the Jupyper examples interactively (and so you can modify and play around as well) in a browser.
- I am not sure about the best way here. Probably the openPMD-viewer
selectargument is of help here (not sure)? In case you know which directions are of interest before doing the simulation, this filtering can also be done using PIConGPU plugins, would also save disk space and output time. However, in case these are not known in advance, one has to output everything first and then indeed apply some filtering.
1. FileOutputParticles = VectorAllSpecies;
Indeed this was the problem. Thank you. In this case what is the role for the Probes?
The role is to track physical quantities (like field values) at certain points of the simulation domain, hence the name. There is more info in the docs. Ofc you can name such species differently, or have multiple species with such properties, we just use Probes as name in the examples.
Actually much can be learnt from the matplotlib documentation which I just found
I am re-opening this since someone has to implement the renaming in PIConGPU from particleId to id during I/O.
Update: I guess that someone fell to me, implemented in #3165 :gift:
solved in #3165
Most helpful comment
@sbastrakov @ax3l Actually it seems to me that @cbontoiu's last command line is valid and should have worked
(
pt = ParticleTracker(data, iteration=300, select={'uz':[0.05, 0.2], 'z':[0.97, 0.98]}, species='e'))I am wondering if, in fact, the problem is that PIConGPU maybe does not fully comply with the openPMD standard. More specifically, PIConGPU stores the particle ID as
particleId; but the openPMD standard expects the particle ID to be stored asid:https://github.com/openPMD/openPMD-standard/blob/latest/STANDARD.md#records-for-each-particle-species
and
openPMD-viewerdoes rely on this convention.Does this seem correct?