Lightgbm: monotonic constraints not working?
I have rerun my checks on monotonic constraints and could not reproduce them under the current lightgbm release 2.2.3 as well as under 2.1.x. The constraints are not satisfied.
Environment info
Operating System: Windows
CPU/GPU model: CPU
C++/Python/R version: R 3.5.0
Error message
None.
Reproducible examples
Steps to reproduce
library(tidyverse)
library(caret)
library(lightgbm)
library(xgboost)
# Generate function with one input x
set.seed(20)
x <- seq(-0.2, 1, by = 0.001)
y <- x^2 + rnorm(length(x), sd = 0.1)
dat <- data.frame(x, y)
base_plot <- ggplot(dat, aes(x = x, y = y)) +
geom_point()
base_plot
# train/test split
ind <- caret::createDataPartition(y, p = 0.8, list = FALSE) %>% c
train <- dat[ind, ] %>% data.matrix
test <- dat[-ind, ] %>% data.matrix
# LGB
lgb_data <- lgb.Dataset(train[, "x", drop = FALSE], label = train[, "y"])
# Some reasonable parameters (would need to be optimized)
params <- list(learning_rate = 0.03,
num_leaves = 31,
min_data_in_leaf = 10,
bagging_fraction = 0.7,
bagging_freq = 4,
nthread = 4,
max_bin = 255,
metric = "rmse",
objective = "regression")
# Use cross-validation to figure out an appropriate number of iterations
fit_cv <- lgb.cv(params = params,
data = lgb_data,
nfold = 5,
nrounds = 1000,
early_stopping_rounds = 20,
verbose = 0)
fit_cv$best_iter
fit <- lgb.train(params = params,
data = lgb_data,
nrounds = fit_cv$best_iter,
verbose = 0)
# Visualize predictions on test data
test_pred <- data.frame(test, pred = predict(fit, test))
base_plot +
geom_line(data = test_pred, aes(x = x, y = pred),
col = "blue", lwd = 2)
# Now with monotonicity constraints
fit_monotone <- lgb.train(params = c(params, monotone_constraints = '1'),
data = lgb_data,
nrounds = fit_cv$best_iter,
verbose = 1)
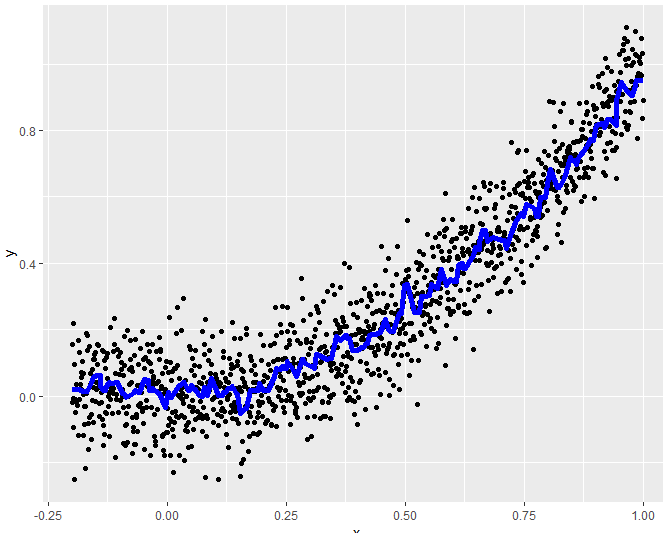
# Visualize predictions on test data
test_pred$pred_monotone <- predict(fit_monotone, test)
base_plot +
geom_line(data = test_pred, aes(x = x, y = pred_monotone),
col = "blue", lwd = 2)
All 7 comments
@mayer79
we have test for mc in python package: https://github.com/Microsoft/LightGBM/blob/af3c4f89ff3989516c60d3f1f1d1e10c7758b855/tests/python_package_test/test_engine.py#L670-L708
can you try to pass mc parameter to dataset? (or remave the cv).
@guolinke: I could not reproduce the problem in Python, so I tried out different things to identify what goes wrong in the example code above.
It seems that old instances of "params" are stuck somewhere in C++ aether (?) and are not being updated by the next call of lgb.train. Adding one row
lgb_data <- lgb.Dataset(train[, "x", drop = FALSE], label = train[, "y"])
just before calling the constrained model solves the problem. Do you think when running gridsearch-CV, we would need to construct the data set like this within the gridsearch loop?
The Dataset will be initialized only one-time, and use the parameter it got first to initialize. For example, in you case, the parameter in the cv is used to initialize the dataset.
After initialize of dataset, the parameter cannot be override.
For grid search, it depends. Some parameter don't affect the dataset object, like num_trees.
Oh - this is a very valuable information, best thanks. So, nothing is wrong with monotonic constraints and we can close this issue. I am wondering how complex it would be to provide an warning (or even error) if the current parameters are inconsistent with those used in initialization of the data set?
@guolinke is there a way to get the parameters of the Dataset and update them? Or should we use free data parameter when creating the Dataset?
I think it is not possible to updata dataset's parameters now.
Free_raw_data cannot help as well, since the constructed dataset is not freed.
I will fix this when I have time.
Most helpful comment
The Dataset will be initialized only one-time, and use the parameter it got first to initialize. For example, in you case, the parameter in the cv is used to initialize the dataset.
After initialize of dataset, the parameter cannot be override.