I am trying to write up procedures for splitting a lot when one item split from the lot is re-cataloged. I would like this process to facilitate reporting on collection management effort in Arctos. Here is a sketch of what I think should be done:
- Create a loan that includes subsamples of all specimen parts that are being split from a lot.
Authorized by – name of agent who authorized the lot split
Received by – collection receiving the split part
In-House Contact – agent name of "donating" collection manager
Outside Contact – agent name of person who determined an identification or created other
information that requires the specimen to be given its own catalog number
Loan Type – in house
Transaction Date – date of lot split
Due Date – date of lot split unless loan is being recorded as part of legacy data, then date of recording
Loan Status – closed
Nature of Material - Loan to split lots and separately catalog specimens with published data
All subsamples in the loan should have disposition of "recataloged"
- Create an Accession for the split parts that will be given new catalog numbers
How obtained? – transfer
Status – complete
Received date – date of lot split
Nature of Material – specimens split from lots in existing cataloged items.
Authorized by – name of agent who authorized the lot split
In-House Contact – agent name of receiving collection manager
Outside Contact – agent name(s) of person(people) who determined an identification or created other information that requires the specimen to be given its own catalog number
Received from – agent name of donating collection manager
- Catalog newly split parts
Clone the original cataloged item to create a new cataloged item for the split part
Add identifier of original cataloged item with "same lot as" relationship to the new record
It is the last step that seems like excessive work when there are a lot of splits happening at once. Splitting a single lot doesn't take too long. I recently worked on a loan and publication that resulted in over 60 split lots which can take a really long time. I'm open to ideas for making this process more streamlined that still allows everyone to understand what happened and provides attribution to everyone in the process for the work involved. I considered a download of items in the original loan to create a new bulkload for the new specimens, but stuff in the download requires A LOT of parsing and work before I can get it into bulkoad format.
All 23 comments
@DerekSikes does this often and correctly and hopefully will chime in.
Not sure I see the counting coup angle or the need for a bunch of new transactions. Lots are split when someone didn't catalog the useful thing in the first place, sometimes because the original was always intended for splitting (think 5 gallons of "tiny things that show up in light traps" or "guts for the parasitologists to dig through"). Whatever the situation, this just seems like normal curatorial work to me - you're fixing a previous choice that can't support the science you're trying to do, delaying the work of sorting/identifying/cataloging (and possibly leveraging some experts), etc.
There are at least three tools which should make this easy.
- data entry has a clone by barcode option
- individual specimen records have two clone options
- specienresults has a download for bulkloader option
need for a bunch of new transactions.
It is the only way we can associate stuff with projects? I want to be able to demonstrate that these things were identified as part of a specific project.
Yes, we do this often.
In some cases we empty the original container so for example 1 jar of 1000+
insects that has a catalog number becomes 1000+ records of pinned insects,
each with their own records. We just use the clone by barcode tool: clone
the original into the bulkloader (& edit to change 'ethanol' to 'pinned'),
use the tool to get to the interface to scan all the new barcodes in, click
the button & 1000+ new records are created each with their own barcodes.
Then delete the original since it's an empty jar now, ready to be used for
another batch later.
Not sure what problem Teresa was trying to solve with all those transaction
steps. It's pretty easy to add these newly made records to the project they
belong to...
-Derek
On Wed, Aug 28, 2019 at 12:30 PM dustymc notifications@github.com wrote:
@DerekSikes https://github.com/DerekSikes does this often and correctly
and hopefully will chime in.Not sure I see the counting coup angle or the need for a bunch of new
transactions. Lots are split when someone didn't catalog the useful thing
in the first place, sometimes because the original was always intended for
splitting (think 5 gallons of "tiny things that show up in light traps" or
"guts for the parasitologists to dig through"). Whatever the situation,
this just seems like normal curatorial work to me - you're fixing a
previous choice that can't support the science you're trying to do,
delaying the work of sorting/identifying/cataloging (and possibly
leveraging some experts), etc.There are at least three tools which should make this easy.
- data entry has a clone by barcode option
- individual specimen records have two clone options
- specienresults has a download for bulkloader option
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/2239?email_source=notifications&email_token=ACFNUM4RTG3GRFSBMHIXVODQG3N6DA5CNFSM4IRR54B2YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD5MMAJI#issuecomment-525910053,
or mute the thread
https://github.com/notifications/unsubscribe-auth/ACFNUM4AC3LHZE5TFQAYYLLQG3N6DANCNFSM4IRR54BQ
.
--
+++++++++++++++++++++++++++++++++++
Derek S. Sikes, Curator of Insects
Professor of Entomology
University of Alaska Museum
1962 Yukon Drive
Fairbanks, AK 99775-6960
phone: 907-474-6278
FAX: 907-474-5469
University of Alaska Museum - search 400,276 digitized arthropod records
http://arctos.database.museum/uam_ento_all
http://www.uaf.edu/museum/collections/ento/
+++++++++++++++++++++++++++++++++++
Interested in Alaskan Entomology? Join the Alaska Entomological
Society and / or sign up for the email listserv "Alaska Entomological
Network" at
http://www.akentsoc.org/contact_us http://www.akentsoc.org/contact.php
You can't add specimens to a project, only transactions. Am I wrong there?
Also, Derek - do you have any of this written down in any more detail than that? I want to create a how to. Thanks!
I'll let Dusty weigh in on the adding to projects bit. I suppose you can
just make a data loan for the specimens, add them & then add that loan to
the project?
& No, I don't have that written down.
-Derek
On Wed, Aug 28, 2019 at 3:32 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
Also, Derek - do you have any of this written down in any more detail than
that? I want to create a how to. Thanks!—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/2239?email_source=notifications&email_token=ACFNUMYFGNYSI4YGNSS5VWLQG4DKFA5CNFSM4IRR54B2YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD5MYVCA#issuecomment-525961864,
or mute the thread
https://github.com/notifications/unsubscribe-auth/ACFNUMY25Q73KGVC2GWQTKDQG4DKFANCNFSM4IRR54BQ
.
--
+++++++++++++++++++++++++++++++++++
Derek S. Sikes, Curator of Insects
Professor of Entomology
University of Alaska Museum
1962 Yukon Drive
Fairbanks, AK 99775-6960
phone: 907-474-6278
FAX: 907-474-5469
University of Alaska Museum - search 400,276 digitized arthropod records
http://arctos.database.museum/uam_ento_all
http://www.uaf.edu/museum/collections/ento/
+++++++++++++++++++++++++++++++++++
Interested in Alaskan Entomology? Join the Alaska Entomological
Society and / or sign up for the email listserv "Alaska Entomological
Network" at
http://www.akentsoc.org/contact_us http://www.akentsoc.org/contact.php
Yes projects are transaction-based.
Whatever is causing the split is likely loan-worthy anyway; it's ultimately the same situation as it would have been if you'd have had the resources/expertise/patience/whatever to sort to individual when originally cataloging.
If there are barcodes involved, adding them to a loan should be trivial.
the split is likely loan-worthy anyway;
That's why I set things up the way I did.
There are at least three tools which should make this easy.
data entry has a clone by barcode option
individual specimen records have two clone options
specimen results has a download for bulkloader option
Where are instructions for the "clone options" for individual specimen records? A search hasn't turned anything up in the handbook and I don't see "clone" in any of our specimen records. Thanks.

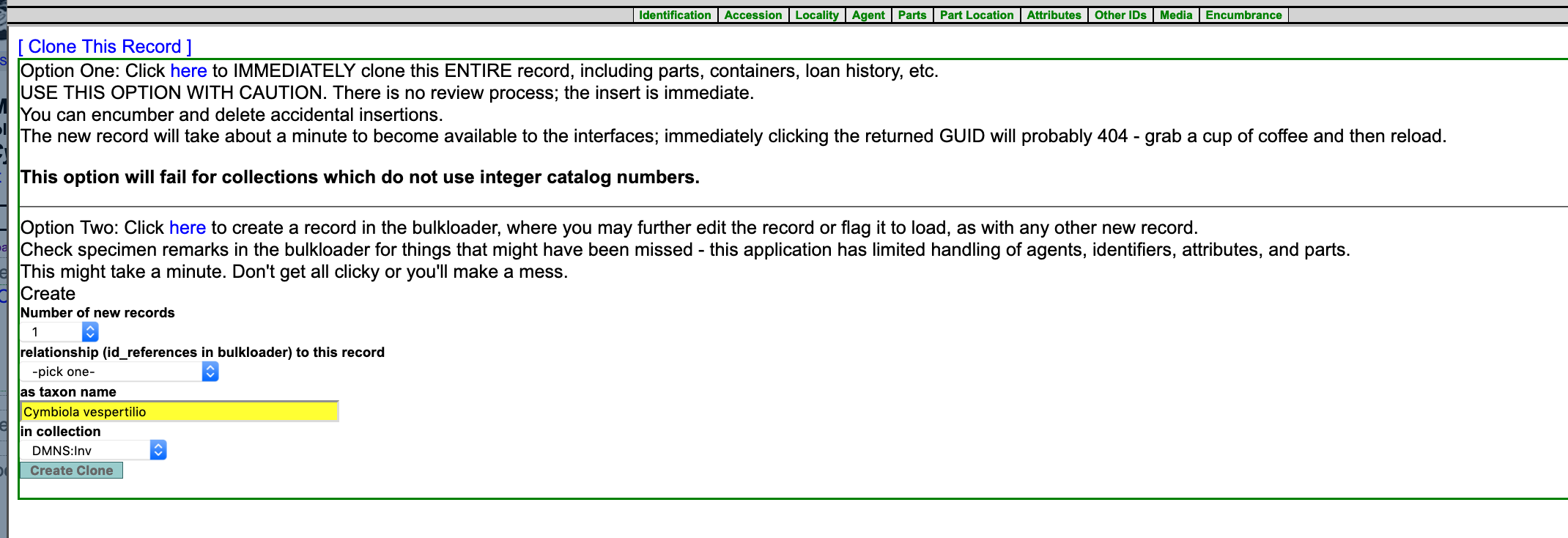
Thanks! Any reason why it's tucked in under the "Other IDs" tab?
I get it now! Relationships start out in cataloging as being entered as "Other IDs". I had to do a couple to figure this out.
Relationships are just otherIDs with pointers to different catalog records. "Normal" otherIDs have a 'relationship' of "self" - it's not usually displayed, but the model is consistent. Arctos provides actionable identifiers for about everything, so this is all of the functionality that a key-pair relationship could provide, plus it gives collections total control over their data (reciprocals are suggested, but never forced in), allows forming resolvable relationships to anything with a decent identifier, and allows forming "soft relationships" to things that do not have useful identifiers. "Parasite of ABC-123" leaves available data (cruddy as it might be!) in the usual spot, and is easily converted to something that can DO STUFF if anyone ever manages to figure out what ABC refers to.
We need to revisit this. I am dealing with splitting lots one record at a time for 57 different catnums that may or may not already have been split out and recataloged under a variety of possible identifiers and versions of the identifier . . . each has to be checked individually and searched using the ID "contains" field, which only allows one record to be searched at a time. This is for legacy parasite material which was sent out prior to the existence of a Para collection, with no loan documentation . . . It is taking me approx 20 - 30 min per record, with the risk of mistake at each step. Plus the single record data entry form and clone function is very buggy at the moment.
See also #1966
https://github.com/ArctosDB/arctos/issues/1966
We need one ID to rule them all, and an easy way to split and link them to the original.
We need documentation for this - I just need to clone 12 mammal records into 12 parasite records with relationship parasite of. Possible? How?
We just use the clone by barcode tool: clone
the original into the bulkloader (& edit to change 'ethanol' to 'pinned'),
use the tool to get to the interface to scan all the new barcodes in, click
the button & 1000+ new records are created each with their own barcodes.
Then delete the original since it's an empty jar now, ready to be used for
another batch later.
Part of this sounds procedural - eg, you mentioned leaving the parasite-parts behind before, which still sounds like a good way to end up in this situation to me. MAYBE there's still a good reason to keep those denormalized parts, but if you do I'd recommend doing so with this situation in mind - add verbose remarks, maybe something with lot count, ???
It is taking me approx 20 - 30 min per record, with the risk of mistake at each step.
This would be exceptionally useful documentation (and/or a publication). I get the impression that everyone who finds a way to catalog the item of scientific interest early ends up wishing they'd have done so sooner, but I don't think that's ever been quantified. "30 minutes for data we don't trust---->5 seconds and a barcode for data we DO trust" seems like something that collections could take to their administrators and get real change from.
I'm also having difficulty imagining how specimens with no identifiers being sent out as loans with no documentation are producing replicable science.
An early application of https://github.com/ArctosDB/arctos/issues/1966 would make this situation much worse - it'd result in a rat and a whole bunch of parasites incorrectly sharing yet another authoritative-sounding identifier.
Yes, this is a very difficult situation:
"specimens with no identifiers being sent out as loans with no
documentation " given the identifiers they do have are things like "H115-1"
which can be entered variously as "H 115", "H115", "H115 1" etc. which is
why I am spending so much time double, triple checking against all possible
forms of data entry into Arctos as well as 10 year old excel file
inventories and previous publications etc. At least in the above, they are
all from Russia from one collector, so I can sort of narrow it down.
I am, however, down to the reasonably more searchable ones with NK
numbers. I have a list of NK numbers with the correct host data that I've
verified. Is there tool I can use to clone these in bulk vs one at a time?
On Mon, Jun 15, 2020 at 10:08 AM dustymc notifications@github.com wrote:
- [EXTERNAL]*
Part of this sounds procedural - eg, you mentioned leaving the
parasite-parts behind before, which still sounds like a good way to end up
in this situation to me. MAYBE there's still a good reason to keep those
denormalized parts, but if you do I'd recommend doing so with this
situation in mind - add verbose remarks, maybe something with lot count, ???It is taking me approx 20 - 30 min per record, with the risk of mistake at
each step.This would be exceptionally useful documentation (and/or a publication). I
get the impression that everyone who finds a way to catalog the item of
scientific interest early ends up wishing they'd have done so sooner, but I
don't think that's ever been quantified. "30 minutes for data we don't
trust---->5 seconds and a barcode for data we DO trust" seems like
something that collections could take to their administrators and get real
change from.I'm also having difficulty imagining how specimens with no identifiers
being sent out as loans with no documentation are producing replicable
science.An early application of #1966
https://github.com/ArctosDB/arctos/issues/1966 would make this
situation much worse - it'd result in a rat and a whole bunch of parasites
incorrectly sharing yet another authoritative-sounding identifier.—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/2239#issuecomment-644227208,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBCVTUSHWF5TFIPNEALRWZBO3ANCNFSM4IRR54BQ
.
Is there tool I can use to clone these in bulk vs one at a time?
Not a tool, but a strategy? Get a download of all the mammal records you need to clone for the parasites and use it to seed a bulkload file for the parasites, make appropriate edits and add appropriate other identifiers, then bulkload parasite records.
I'd probably do what Teresa suggested, it comes with more room for review
NK numbers...tool
Not exactly, and there probably can't/shouldn't be - NKs aren't unique, so feeding such a tool "NK1" and then figuring out that NK1 had been used on 500 parasites and a rat would save you 10 minutes and then cost you a week of cleanup
You can search by a list of NKs, get barcodes from that, and use those in the clonebybarcode. That could be a tool - for now, if you want to send me CSV of the NKs I could add barcodes.
Where is clobebybarcode?
On Mon, Jun 15, 2020, 12:37 PM dustymc notifications@github.com wrote:
- [EXTERNAL]*
I'd probably do what Teresa suggested, it comes with more room for review
NK numbers...tool
Not exactly, and there probably can't/shouldn't be - NKs aren't unique, so
feeding such a tool "NK1" and then figuring out that NK1 had been used on
500 parasites and a rat would save you 10 minutes and then cost you a week
of cleanupYou can search by a list of NKs, get barcodes from that, and use those in
the clonebybarcode. That could be a tool - for now, if you want to send me
CSV of the NKs I could add barcodes.—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/2239#issuecomment-644308012,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBD2L56D2HTFONHWTTLRWZS7VANCNFSM4IRR54BQ
.