Sparklyr: mutate_at doesn't work with spark udf functions
Hi, I need to load a csv file into spark via spark_read_csv and correct the date time columns to the right data type. Since I have a couple of date time columns, I want to use mutate_at to apply the conversion on all of them at once.
tbl_data %>%
mutate_at(vars(starts_with('col_dt')),to_date) %>%
select(starts_with('col_dt')) %>%
head()
This gives me the error below:
Error in is_fun_list(.funs) : object 'to_date' not found

Note that I'm able to use this to_date spark udf function in mutate like this:
tbl_data %>%
mutate(col_dt = to_date(col_dt)) %>%
select(col_dt) %>%
head()
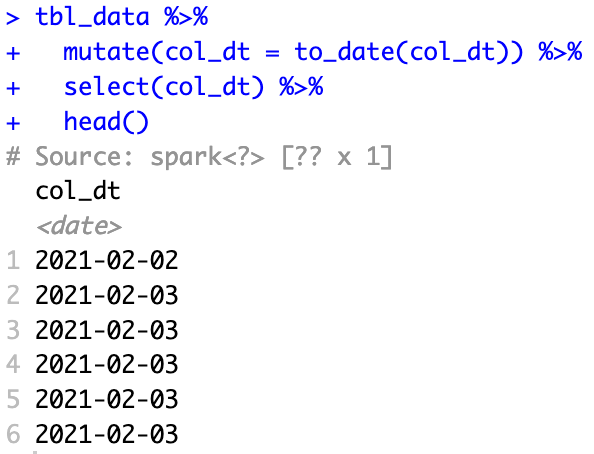
I see exactly the same problem for to_timestamp as well, where it can be recognized in mutate() but not mutate_at().
I'm using sparklyr 1.4.0 & dplyr 1.0.2.
Session Info:
> sessionInfo()
R version 3.6.1 (2019-07-05)
Platform: x86_64-conda_cos6-linux-gnu (64-bit)
Running under: Red Hat Enterprise Linux 8.2 (Ootpa)
Matrix products: default
BLAS/LAPACK: /opt/conda/envs/R-3.6/lib/R/lib/libRblas.so
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C LC_TIME=en_US.UTF-8
[4] LC_COLLATE=en_US.UTF-8 LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] dplyr_1.0.2 sparklyr_1.4.0 ibmwsrspark_0.1.3 reticulate_1.18 httr_1.4.2
loaded via a namespace (and not attached):
[1] Rcpp_1.0.5 pillar_1.4.6 compiler_3.6.1 dbplyr_1.4.4 r2d3_0.2.3
[6] base64enc_0.1-3 tools_3.6.1 digest_0.6.27 jsonlite_1.7.1 lifecycle_0.2.0
[11] tibble_3.0.4 lattice_0.20-41 pkgconfig_2.0.3 rlang_0.4.8 Matrix_1.2-18
[16] cli_2.1.0 DBI_1.1.0 rstudioapi_0.11 yaml_2.2.1 parallel_3.6.1
[21] withr_2.3.0 generics_0.0.2 htmlwidgets_1.5.2 vctrs_0.3.4 askpass_1.1
[26] rappdirs_0.3.1 rprojroot_1.3-2 grid_3.6.1 tidyselect_1.1.0 glue_1.4.2
[31] forge_0.2.0 R6_2.4.1 fansi_0.4.1 purrr_0.3.4 tidyr_1.1.2
[36] blob_1.2.1 magrittr_1.5 backports_1.1.10 ellipsis_0.3.1 htmltools_0.5.0
[41] assertthat_0.2.1 config_0.3 utf8_1.1.4 openssl_1.4.3 crayon_1.3.4
All 3 comments
@wanting0wang I think the error you ran into is expected because dplyr (by design) needs to evaluate the function expression in a mutate_at() call within some specific R env.
But there is a workaround: you can instead have a one-sided formula such as ~ to_date(.x) to avoid dplyr trying to evaluate to_date as an R expression, e.g., the following will work as expected in sparklyr 1.6:
library(sparklyr)
library(dplyr)
sc <- spark_connect(master = "local")
tbl_data <- copy_to(
sc,
data.frame(num = 1, string = "tom", col_dt_1 = "1970-01-01", col_dt_2 = "1970-01-02")
)
tbl_data %>% mutate(across(starts_with("col_dt"), ~ to_date(.x))) %>% print()
## # Source: spark<?> [?? x 4]
## num string col_dt_1 col_dt_2
## <dbl> <chr> <date> <date>
## 1 1 tom 1970-01-01 1970-01-02
@yitao-li Thank you for the example! Yes mutate(across()) works perfectly for me. So I suppose spark udf functions are just not supported in mutate_at(), and mutate(across()) is a more flexible alternative to mutate_at() and can correctly parse these functions.
@wanting0wang just a heads up, the *_at(), *_if(), and *_all() dplyr verbs are all actually getting deprecated for the across() constructions. So things like mutate(across()) and summarise(across()) should cover any cases where you'd previously use mutate_at(), mutate_if(), summarise_all(), etc.
Most helpful comment
@wanting0wang just a heads up, the
*_at(),*_if(), and*_all()dplyr verbs are all actually getting deprecated for theacross()constructions. So things likemutate(across())andsummarise(across())should cover any cases where you'd previously usemutate_at(),mutate_if(),summarise_all(), etc.