Hello,
I am using st_sample on a polygon, and I realized in the documentation that the sampling size might differ from the requested size because it is sampling from the bounding box of the polygon. What I am not entirely clear is how it is possible to get more samples than the requested size, which I often encounter.
The following is a simple test:
x = st_sfc(st_polygon(list(rbind(c(0,0),c(90,0),c(90,90),c(0,0)))))
plot(x); plot(st_sample(x, 5, type = "random"), add = TRUE)
As for a slightly different question, what would be the best way to ensure that I get the final sample number to the requested size?
Thank you!
All 25 comments
My read of the code is that it is taking your size of 5 and multiplying it by the ratio of your polygon area and the bounding box area. Here, it is sampling 10 from the bounding box:
a0 = st_area(st_make_grid(x, n = c(1,1))) # this is the bounding box
a1 = sum(st_area(x))
size = round(size * a0 / a1)
I suppose you could run st_sample(x, 5, type = "random")[1:5] to always get a maximum of 5 points?
Thank you! This explains a lot!
However, the simple solution of subsetting did not work many times because the number of samples is less than the requested size for roughly half of the time. A histogram of the difference between requested size and resulting size is attached:
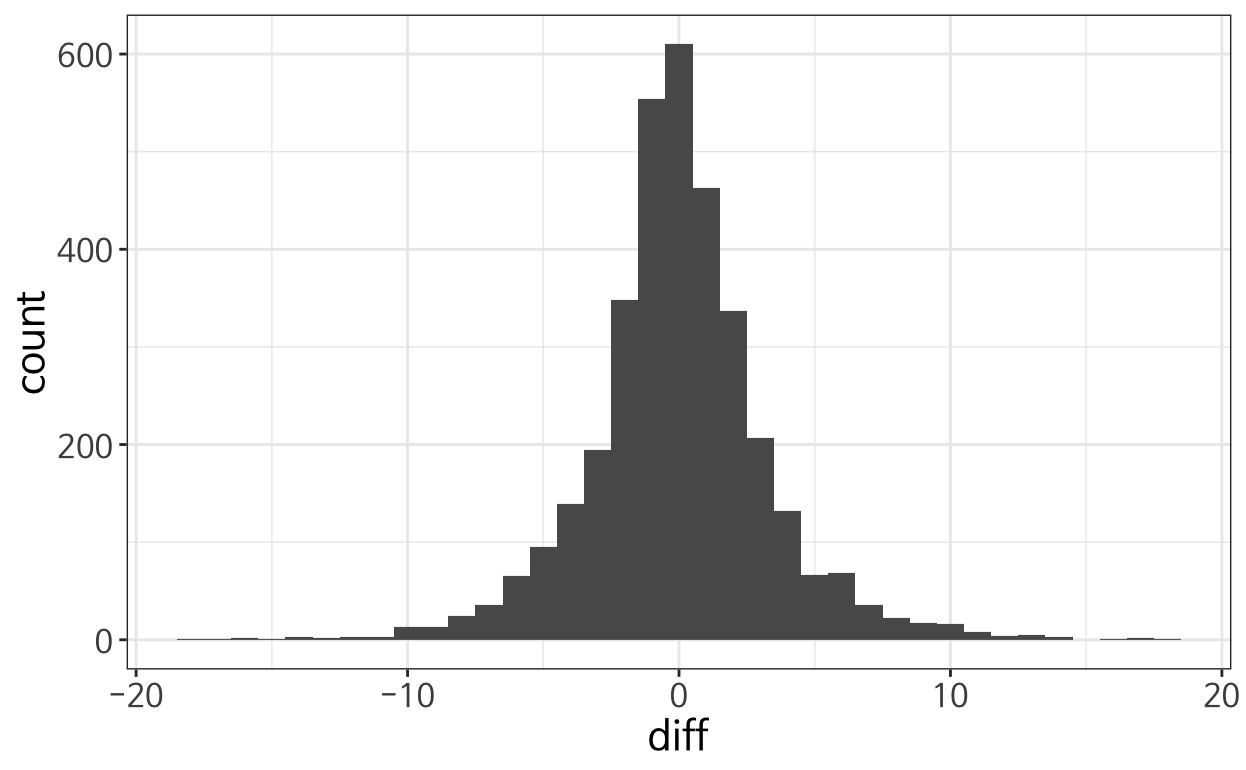
My dirty way of dealing with this was to increase the number of samples such that it is very unlikely to yield less than the requested number (in my case, arbitrarily adding +30 to the st_sample parameter) but if there could be more elegant solution, I would be grateful!
I haven't made a study out of this, but would be happy to hear of pointers to efficient algorithms sampling _exactly_ n points from a (set of) polygon(s). What we now have is a random process, which performs badly when the ratio of polygon area / bounding box area is small (e.g. linear elements represented as narrow polygons).
I developed this workaround by using a while in an ad-hoc function:
get_random_pt <- function(x) {
random_pt <- st_sample(x , size = 1, type = "random")
while (length(random_pt) == 0) {
random_pt <- st_sample(x , size = 1, type = "random")
}
return(random_pt)
}
Then, if you want do it for a data.frame of polygons (polygons) as I needed,wrap this function in a map() and then assign the CRS of the original polygons df (crs_polygons <- st_crs(polygons)) back via st_sfc():
library(purrr)
random_pts <- map(st_geometry(polygons), get_random_pt)
random_pts <- st_sfc(unlist(random_pts, recursive = FALSE),
crs = crs_polygons
)
@damianooldoni you could probably get away with something like the following to include size:
get_random_pt <- function(x, size) {
random_pt <- st_sample(x , size = size, type = "random")
while (length(random_pt) < size) {
random_pt <- c(random_pt, st_sample(x , size = 1, type = "random"))
}
return(random_pt)
}
There is no chance that the result will have more points than size on the first try.
I would re-write it something like:
st_sample_exact <- function(x, size) {
random_pt <- st_sample(x , size = size, type = "random")
while (length(random_pt) < size) {
diff <- size - length(random_pt)
random_pt_new <- st_sample(x , size = diff, type = "random")
random_pt <- c(random_pt, random_pt_new)
}
random_pt
}
Will aim to do some tests and benchmarks...
Results: I think the while solution suggested by @damianooldoni works well but converges quicker if you set size = diff as in my solution below (will update the code above to avoid confusion):
library(sf)
#> Linking to GEOS 3.6.2, GDAL 2.2.3, PROJ 4.9.3
library(spData)
set.seed(2018)
get_random_pt <- function(x, size) {
random_pt <- st_sample(x , size = size, type = "random")
while (length(random_pt) < size) {
random_pt <- c(random_pt, st_sample(x , size = 1, type = "random"))
}
return(random_pt)
}
st_sample_exact <- function(x, size) {
random_pt <- st_sample(x , size = size, type = "random")
while (length(random_pt) < size) {
diff <- size - length(random_pt)
random_pt_new <- st_sample(x , size = diff, type = "random")
random_pt <- c(random_pt, random_pt_new)
}
random_pt
}
n = 1e3
p_st_sample = st_sample(x = nz, size = n)
p_get_random = get_random_pt(nz, n)
p_st_sample_exact = st_sample_exact(nz, n)
length(p_st_sample)
#> [1] 950
length(p_get_random)
#> [1] 1000
length(p_st_sample_exact)
#> [1] 1000
bench::mark(check = FALSE,
st_sample(x = nz, size = n),
get_random_pt(nz, n),
st_sample_exact(nz, n)
)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
#> # A tibble: 3 x 10
#> expression min mean median max `itr/sec`
#> <bch:expr> <bch:tm> <bch:tm> <bch:tm> <bch:tm> <dbl>
#> 1 st_sample(x = nz, size = n) 277.54ms 286.56ms 286.56ms 295.57ms 3.49
#> 2 get_random_pt(nz, n) 4.63s 4.63s 4.63s 4.63s 0.216
#> 3 st_sample_exact(nz, n) 253.18ms 267.25ms 267.25ms 281.32ms 3.74
#> # ... with 4 more variables: mem_alloc <bch:byt>, n_gc <dbl>, n_itr <int>,
#> # total_time <bch:tm>
Created on 2018-11-12 by the reprex package (v0.2.1)
I'm confident that this is a useful addition: had this problem today - would be great to have an option to specify the exact result. A wider issue is that currently st_sample() can fails silently. I think there should be a warning message if length(result) < size.
The function demonstrated above could be made more sophisticated by measuring the area of the polygon vs the area of the bounding box and setting the size accordingly but that would also take time. My question is: assuming this is something that would be useful where should it live and what should it be called?
Tentative suggestion: create a new function st_sample_exact() to avoid changing the API of the existing st_sample() function. Happy to give that a go pending general nod of approval.
In terms of an algorithm to do this, see: https://cs.stackexchange.com/questions/14007/random-sampling-in-a-polygon
That relies on triangulation and then sampling from each triangle in proportion with its size.
Update: seems 2018 is a lucky number, and that the st_sample_exact() function defined above does not always return an exact number of points...
set.seed(2019)
n = 1e3
p_st_sample = st_sample(x = nz, size = n)
p_get_random = get_random_pt(nz, n)
p_st_sample_exact = st_sample_exact(nz, n)
length(p_st_sample)
#> [1] 974
length(p_get_random)
#> [1] 1000
length(p_st_sample_exact)
#> [1] 1003
Created on 2018-11-12 by the reprex package (v0.2.1)
Another option, using a new sampling function that deliberately always provides length(result) <= size:
library(sf)
#> Linking to GEOS 3.6.2, GDAL 2.2.3, PROJ 4.9.3
library(spData)
st_poly_sample_simple_random = function(x, size) {
bb = st_bbox(x)
lon = runif(size, bb[1], bb[3])
lat = runif(size, bb[2], bb[4])
m = cbind(lon, lat)
p = st_sfc(lapply(seq_len(nrow(m)), function(i)
st_point(m[i,])), crs = st_crs(x))
p[x]
}
st_sample_exact <- function(x, size) {
random_pt <- st_poly_sample_simple_random(x , size = size)
while (length(random_pt) < size) {
diff <- size - length(random_pt)
random_pt_new <- st_poly_sample_simple_random(x , size = diff)
random_pt <- c(random_pt, random_pt_new)
}
random_pt
}
set.seed(2019)
n = 1e3
p_st_sample = st_sample(x = nz, size = n)
p_st_sample_exact = st_sample_exact(nz, n)
plot(p_st_sample_exact)

length(p_st_sample)
#> [1] 974
length(p_st_sample_exact)
#> [1] 1000
bench::mark(check = FALSE,
st_sample(x = nz, size = n),
st_sample_exact(nz, n)
)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
#> # A tibble: 2 x 10
#> expression min mean median max `itr/sec`
#> <bch:expr> <bch:tm> <bch:tm> <bch:tm> <bch:tm> <dbl>
#> 1 st_sample(x = nz, size = n) 411.28ms 431.28ms 431.28ms 451.27ms 2.32
#> 2 st_sample_exact(nz, n) 1.05s 1.05s 1.05s 1.05s 0.950
#> # ... with 4 more variables: mem_alloc <bch:byt>, n_gc <dbl>, n_itr <int>,
#> # total_time <bch:tm>
Created on 2018-11-12 by the reprex package (v0.2.1)
Looking at that I think the simplest solution may be add something like:
if(length(res) > size) res = res[1:size]
as suggested by @jsta may be the best way forward. I think it would be good if st_sample() worked the same as sample(), always returning the desired number of records, assuming it's not 'too late' to change the behaviour of st_sample(). Otherwise it will have to be adding more arguments/functions I guess.
An alternative approach to https://github.com/Robinlovelace/sf/commit/e3e6f52667137fdbf2c83271a78567885a95bb92 that does not change st_sample() is to make the new st_sample_exact() function always return the right value:
library(sf)
#> Linking to GEOS 3.6.2, GDAL 2.2.3, PROJ 4.9.3
library(spData)
set.seed(2019)
st_sample_exact <- function(x, size) {
random_pt <- st_sample(x , size = size, type = "random")
while (length(random_pt) < size) {
diff <- size - length(random_pt)
random_pt_new <- st_sample(x , size = diff, type = "random")
random_pt <- c(random_pt, random_pt_new)
}
if(length(random_pt ) > size) {
random_pt = random_pt[1:size]
}
random_pt
}
n = 1000
p_st_sample = st_sample(x = nz, size = n)
p_st_sample_exact = st_sample_exact(nz, n)
length(p_st_sample)
#> [1] 974
length(p_st_sample_exact)
#> [1] 1000
bench::mark(check = FALSE,
st_sample(x = nz, size = n),
st_sample_exact(nz, n)
)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
#> # A tibble: 2 x 10
#> expression min mean median max `itr/sec` mem_alloc
#> <bch:expr> <bch> <bch> <bch:> <bch> <dbl> <bch:byt>
#> 1 st_sample(x = nz, size = n) 267ms 274ms 274ms 281ms 3.65 1.66MB
#> 2 st_sample_exact(nz, n) 296ms 314ms 314ms 331ms 3.19 2.12MB
#> # ... with 3 more variables: n_gc <dbl>, n_itr <int>, total_time <bch:tm>
sapply(1:50, function(x) length(st_sample_exact(nz, 1000)))
#> [1] 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000
#> [15] 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000
#> [29] 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000 1000
#> [43] 1000 1000 1000 1000 1000 1000 1000 1000
Created on 2018-11-13 by the reprex package (v0.2.1)
Question for @WWolf and @edzer: which approach is better from a user/developer perspective?
I haven't (yet) read the thread in full detail, but instead of adding a function I'd add a parameter to st_sample, say, exact = TRUE; we could leave it to FALSE now until we decided which algorithm to use for TRUE; when we've agreed about a solution I'd be happy to have TRUE as a default because it is really a nuisance to not get what you ask for, exactly.
Thanks for the quickfire feedback @edzer and sorry for all the brainstorming content. I will try to implement your suggestion to
add a parameter to st_sample, say, exact = TRUE
in a new pull request. More soon.
Heads-up: https://github.com/r-spatial/sf/pull/896 now implements that suggestion.
Results below:
devtools::install_github("robinlovelace/sf", ref = "sample-exact-argument")
#> Skipping install of 'sf' from a github remote, the SHA1 (fefdff7f) has not changed since last install.
#> Use `force = TRUE` to force installation
library(sf)
#> Linking to GEOS 3.6.2, GDAL 2.2.3, PROJ 4.9.3
l1 = purrr::map_int(1:10, ~length(st_sample(spData::nz, 1e3)))
l2 = purrr::map_int(1:10, ~length(st_sample(spData::nz, 1e3, exact = TRUE)))
summary(l1)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> 946 1000 1004 1006 1015 1056
summary(l2)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> 1000 1000 1000 1000 1000 1000
Created on 2018-11-13 by the reprex package (v0.2.1)
Another benchmark checking performance and that the results are roughly the same:
devtools::install_github("robinlovelace/sf", ref = "sample-exact-argument")
#> Skipping install of 'sf' from a github remote, the SHA1 (fefdff7f) has not changed since last install.
#> Use `force = TRUE` to force installation
library(sf)
#> Linking to GEOS 3.6.2, GDAL 2.2.3, PROJ 4.9.3
l1 = purrr::map_int(1:10, ~length(st_sample(spData::nz, 1e3)))
l2 = purrr::map_int(1:10, ~length(st_sample(spData::nz, 1e3, exact = TRUE)))
summary(l1)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> 940.0 982.2 1008.0 1001.9 1023.8 1035.0
summary(l2)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> 1000 1000 1000 1000 1000 1000
bench::mark(check = FALSE,
exactFALSE = {p1 = st_sample(spData::nz, 1e3)},
exactTRUE = {p2 = st_sample(spData::nz, 1e3, exact = TRUE)}
)
#> Warning: Some expressions had a GC in every iteration; so filtering is
#> disabled.
#> # A tibble: 2 x 10
#> expression min mean median max `itr/sec` mem_alloc n_gc n_itr
#> <bch:expr> <bch> <bch> <bch:> <bch> <dbl> <bch:byt> <dbl> <int>
#> 1 exactFALSE 370ms 371ms 371ms 373ms 2.69 1.66MB 15 2
#> 2 exactTRUE 287ms 290ms 290ms 293ms 3.45 1.95MB 13 2
#> # ... with 1 more variable: total_time <bch:tm>
par(mfrow = c(1, 2))
plot(p1)
plot(p2)

par(mfrow = c(1, 1))
Created on 2018-11-13 by the reprex package (v0.2.1)
Hi, wow, a long thread! Although I haven't perused very carefully, I think the expected behavior of a naiive user for st_sample would be that it returns the exact number of records all the time. So an option in the st_sample with "exact=TRUE" with the caveat that it will take longer (I guess that is what the algorithm is about?) would be my preference, but I am just a user. BTW, thanks for all the effort!
Thanks for the feedback @WWolf. I agree. Assuming we do go with this exact argument approach it can be introduced incrementally, first as a new argument that does not change the default behaviour (exact = FALSE, as in #896 ) and then, after a while and possibly with messages warning people about upcoming changes, it can switch to TRUE as the default.
Update my side: the build is finally working so if @edzer is happy with it I think that fixes the issue.
@WWolf if you could test the new version of st_sample(..., exact = T) on your data and report back if it works or if you find any issues, that would be appreciated.
I tested your PR on my data. It works. Nice, thanks.
As user, I would go a step further than https://github.com/r-spatial/sf/issues/813#issuecomment-438301331. Parameter exact should be unnecessary as exact = TRUE is what every user would like to use. If you want less points, just reduce size.
Thanks for the feedback @damianooldoni. I think it's worth keeping exact = TRUE but will let @edzer have the last say. Heads-up Edzer: I think this is almost good to merge. Please let me know of anything else you need in there (will look at documentation now):
This can be closed now. @damianooldoni and @WWolf are you happy for this to be closed now? Please let close if so! And let me know if you spot any issues with the new performance.
Thank you very much for your effort!!
OK, please note that in 0.7-3, exact is still FALSE by default, because we found a regression in a reverse dependency check that needs to be resolved first.
Most helpful comment
Hi, wow, a long thread! Although I haven't perused very carefully, I think the expected behavior of a naiive user for
st_samplewould be that it returns the exact number of records all the time. So an option in thest_samplewith "exact=TRUE" with the caveat that it will take longer (I guess that is what the algorithm is about?) would be my preference, but I am just a user. BTW, thanks for all the effort!