I came across an interesting approach to speeding up spatial join operations in a PostGIS workflow: use ST_Subdivide to split large, complex geometries into smaller, simpler ones that can be indexed much quicker.
Steve Vance elaborates on this topic in his blog post (which was inspired by a presentation given by Paul Ramsey) and produced this image illustrating the approach:

ST_Subdivide strikes me as useful addition to sf’s quiver of geometrical operations - especially for giving a speed boost to the operations that rely on indexing (e.g., the maximum overlap overlay recently added to st_join() in #578).
All 13 comments
Would st_subdivide also be a nice way to chunk up operations in a memory-efficient way for sending jobs to arbitrary types of workers using, for example, future_lapply?
Something like this?
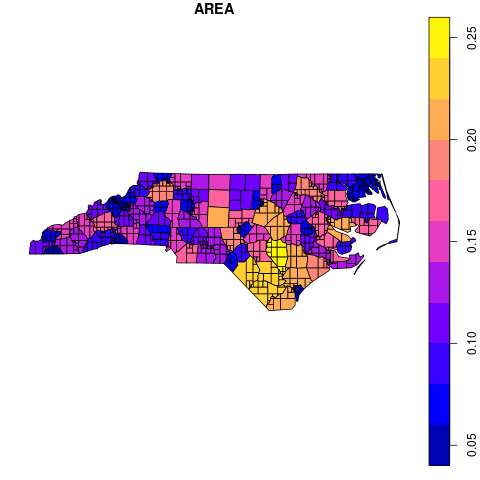
Yes that looks like the output I would expect for something like st_subdivide(nc, max_vertices = 10)
So, this gives back 100 features with GEOMETRYCOLLECTION geometries; to gain the speed on large geometries, I'm not sure whether they're fine like this, or whether you'd still have to pull them through st_collection_extract (or st_cast without a to argument) to get a feature for each subdivision. Benchmarks welcome!
st_subdivide() works the way I expected it to and your tip about using st_collection_extract() is very helpful.
Unfortunately, I'm not seeing the 30x speed up mentioned in Paul Ramsey's presentation (see here). I'll close the issue and keep playing around with it.
Here's a benchmark using nc and a watershed dataset:
# SETUP ----
library(lwgeom) # devtools::install_github('r-spatial/lwgeom)
library(tidyverse)
library(sf)
library(esri2sf) # devtools::install_github('yonghah/esri2sf')
library(rbenchmark)
# LOAD DATA ----
# NC counties
nc <- read_sf(system.file("shape/nc.shp", package = "sf"))
nc <- st_transform(nc, 32119) # NC state plane, m
# NC watersheds
url <- "https://services.nconemap.gov/secure/rest/services/NC1Map_Watersheds/MapServer/2"
nc_wtr <- esri2sf(url)
##
## Attaching package: 'jsonlite'
## The following object is masked from 'package:purrr':
##
## flatten
## [1] "Feature Layer"
## [1] "esriGeometryPolygon"
nc_wtr <- st_transform(nc_wtr, 32119) # NC state plane, m
# PLOT OVERLAY ----
plot(nc_wtr[, 1])
plot(nc[, 1], col = "transparent", add = TRUE)

# BENCHMARK: ST_SUBDIVIDE IMPACT ON ST_INTERSECTS() ----
cols <- c("replications", "elapsed", "user.self", "sys.self")
nc_wtr_subdvd <- nc_wtr %>%
st_subdivide(max_vertices = 256) %>%
st_collection_extract()
# Without st_subdivided
set.seed(7890)
benchmark(columns = cols, replications = 100, {
st_intersects(nc_wtr, nc)
})
## replications elapsed user.self sys.self
## 1 100 18.45 18.22 0.24
# With st_subdivided
benchmark(columns = cols, replications = 100, {
st_intersects(nc_wtr_subdvd, nc)
})
## replications elapsed user.self sys.self
## 1 100 20.49 20.34 0.08
Session info
devtools::session_info()
## Session info -------------------------------------------------------------
## setting value
## version R version 3.4.2 (2017-09-28)
## system x86_64, mingw32
## ui RTerm
## language (EN)
## collate English_United States.1252
## tz America/Los_Angeles
## date 2017-12-15
## Packages -----------------------------------------------------------------
## package * version date source
## assertthat 0.2.0 2017-04-11 CRAN (R 3.4.1)
## backports 1.1.1 2017-09-25 CRAN (R 3.4.1)
## base * 3.4.2 2017-09-28 local
## bindr 0.1 2016-11-13 CRAN (R 3.4.1)
## bindrcpp 0.2 2017-06-17 CRAN (R 3.4.1)
## bitops 1.0-6 2013-08-17 CRAN (R 3.4.1)
## broom 0.4.3 2017-11-20 CRAN (R 3.4.3)
## cellranger 1.1.0 2016-07-27 CRAN (R 3.4.1)
## class 7.3-14 2015-08-30 CRAN (R 3.4.2)
## classInt 0.1-24 2017-04-16 CRAN (R 3.4.2)
## cli 1.0.0 2017-11-05 CRAN (R 3.4.2)
## colorspace 1.3-2 2016-12-14 CRAN (R 3.4.1)
## compiler 3.4.2 2017-09-28 local
## crayon 1.3.4 2017-11-16 Github (r-lib/crayon@b5221ab)
## curl 3.0 2017-10-06 CRAN (R 3.4.2)
## datasets * 3.4.2 2017-09-28 local
## DBI 0.7 2017-06-18 CRAN (R 3.4.1)
## devtools 1.13.4 2017-11-09 CRAN (R 3.4.2)
## digest 0.6.12 2017-01-27 CRAN (R 3.4.1)
## dplyr * 0.7.4 2017-09-28 CRAN (R 3.4.2)
## e1071 1.6-8 2017-02-02 CRAN (R 3.4.2)
## esri2sf * 0.1.0 2017-12-15 Github (yonghah/esri2sf@23867c0)
## evaluate 0.10.1 2017-06-24 CRAN (R 3.4.2)
## forcats * 0.2.0 2017-01-23 CRAN (R 3.4.1)
## foreign 0.8-69 2017-06-22 CRAN (R 3.4.2)
## ggplot2 * 2.2.1.9000 2017-12-13 Github (tidyverse/ggplot2@bfff1d8)
## glue 1.2.0 2017-10-29 CRAN (R 3.4.2)
## graphics * 3.4.2 2017-09-28 local
## grDevices * 3.4.2 2017-09-28 local
## grid 3.4.2 2017-09-28 local
## gtable 0.2.0 2016-02-26 CRAN (R 3.4.1)
## haven 1.1.0 2017-07-09 CRAN (R 3.4.1)
## hms 0.4.0 2017-11-23 CRAN (R 3.4.3)
## htmltools 0.3.6 2017-04-28 CRAN (R 3.4.1)
## httr * 1.3.1 2017-08-20 CRAN (R 3.4.1)
## jsonlite * 1.5 2017-06-01 CRAN (R 3.4.1)
## knitr 1.17 2017-08-10 CRAN (R 3.4.2)
## lattice 0.20-35 2017-03-25 CRAN (R 3.4.2)
## lazyeval 0.2.1 2017-10-29 CRAN (R 3.4.2)
## lubridate 1.7.1 2017-11-03 CRAN (R 3.4.2)
## lwgeom * 0.1-1 2017-12-15 Github (r-spatial/lwgeom@04d8cd6)
## magrittr 1.5 2014-11-22 CRAN (R 3.4.1)
## memoise 1.1.0 2017-04-21 CRAN (R 3.4.2)
## methods * 3.4.2 2017-09-28 local
## mnormt 1.5-5 2016-10-15 CRAN (R 3.4.1)
## modelr 0.1.1 2017-07-24 CRAN (R 3.4.1)
## munsell 0.4.3 2016-02-13 CRAN (R 3.4.1)
## nlme 3.1-131 2017-02-06 CRAN (R 3.4.2)
## parallel 3.4.2 2017-09-28 local
## pillar 0.0.0.9000 2017-12-13 Github (r-lib/pillar@5a082e1)
## pkgconfig 2.0.1 2017-03-21 CRAN (R 3.4.1)
## plyr 1.8.4 2016-06-08 CRAN (R 3.4.1)
## psych 1.7.8 2017-09-09 CRAN (R 3.4.2)
## purrr * 0.2.4.9000 2017-12-13 Github (tidyverse/purrr@62b135a)
## R6 2.2.2 2017-06-17 CRAN (R 3.4.1)
## rbenchmark * 1.0.0 2012-08-30 CRAN (R 3.4.1)
## Rcpp 0.12.14 2017-11-23 CRAN (R 3.4.2)
## RCurl 1.95-4.8 2016-03-01 CRAN (R 3.4.1)
## readr * 1.1.1 2017-05-16 CRAN (R 3.4.1)
## readxl 1.0.0 2017-04-18 CRAN (R 3.4.1)
## reshape2 1.4.3 2017-12-11 CRAN (R 3.4.3)
## rlang 0.1.4 2017-11-05 CRAN (R 3.4.2)
## rmarkdown 1.8 2017-11-17 CRAN (R 3.4.2)
## rprojroot 1.2 2017-01-16 CRAN (R 3.4.2)
## rvest 0.3.2 2016-06-17 CRAN (R 3.4.1)
## scales 0.5.0.9000 2017-11-30 Github (hadley/scales@d767915)
## sf * 0.5-6 2017-12-15 Github (r-spatial/sf@be20ebb)
## stats * 3.4.2 2017-09-28 local
## stringi 1.1.6 2017-11-17 CRAN (R 3.4.2)
## stringr * 1.2.0 2017-02-18 CRAN (R 3.4.1)
## tibble * 1.3.4.9003 2017-12-13 Github (tidyverse/tibble@60281b3)
## tidyr * 0.7.2.9000 2017-12-13 Github (tidyverse/tidyr@efd9ea5)
## tidyverse * 1.2.1 2017-12-13 Github (tidyverse/tidyverse@3769ff2)
## tools 3.4.2 2017-09-28 local
## udunits2 0.13 2016-11-17 CRAN (R 3.4.1)
## units 0.4-6 2017-08-27 CRAN (R 3.4.1)
## utils * 3.4.2 2017-09-28 local
## withr 2.1.0.9000 2017-11-30 Github (jimhester/withr@fe81c00)
## XML 3.98-1.9 2017-06-19 CRAN (R 3.4.1)
## xml2 1.1.1 2017-01-24 CRAN (R 3.4.1)
## yaml 2.1.14 2016-11-12 CRAN (R 3.4.1)
I guess the speed gain is only really there when, like in the blog, you have one or more very large polygons whose bounding box spans most things you're trying to intersect: a lake, river, riverbed or road comes to mind. nc is already a bunch of square blocks.
Good point, @edzer .
@kendonB - That seems like an excellent use of this method, particularly for spatial operations involving high-resolution polygons that shouldn't be simplified.
Here's two more benchmarks - this time testing the speed of st_intersects() with a large sfc_POINT object and:
- the North Carolina state boundary
- watersheds in the state.
The first tests show modest speed improvements (~3x) while the second test shows _dramatic_ improvement (~300x).

# Without st_subdivided
benchmark(columns = cols, replications = 100, {
st_intersects(pts_NCSP, nc_bound_NCSP)
})
## replications elapsed user.self sys.self
## 1 100 9.3 9.25 0.02
# With st_subdivided
benchmark(columns = cols, replications = 100, {
st_intersects(pts_NCSP, nc_subd_NCSP)
})
## replications elapsed user.self sys.self
## 1 100 3.87 3.86 0.01

# Without st_subdivided
benchmark(columns = cols, replications = 10, {
st_intersects(pts_NCSP, nc_wtr_union_NCSP)
})
## replications elapsed user.self sys.self
## 1 10 203.47 203.1 0.02
# With st_subdivided
benchmark(columns = cols, replications = 10, {
st_intersects(pts_NCSP, nc_wtr_union_subd_NCSP)
})
## replications elapsed user.self sys.self
## 1 10 0.78 0.78 0
Reprex + Session info
# SETUP ----
library(lwgeom) # devtools::install_github('r-spatial/lwgeom)
library(tidyverse)
library(sf)
library(esri2sf) # devtools::install_github('yonghah/esri2sf')
library(rbenchmark)
# LOAD DATA ----
# NC counties
nc <- read_sf(system.file("shape/nc.shp", package = "sf"))
nc <- st_transform(nc, 4326)
nc_bound <- st_union(nc)
nc_subd <- nc_bound %>%
st_subdivide(max_vertices = 10) %>%
st_collection_extract()
# NC points
set.seed(3775)
bb <- st_bbox(nc_bound)
pts_tbl <- tibble(x = runif(1e4, min = bb$xmin, max = bb$xmax),
y = runif(1e4, min = bb$ymin, max = bb$ymax))
pts_sf <- pts_tbl %>%
transpose %>%
map(~ st_point(c(.x$x,.x$y), dim = "XY")) %>%
st_sfc %>%
st_set_crs(4326)
# NC watersheds
url <- "https://services.nconemap.gov/secure/rest/services/NC1Map_Watersheds/MapServer/2"
nc_wtr <- esri2sf(url)
nc_wtr <- st_transform(nc_wtr, 4326)
nc_wtr_union <- st_union(nc_wtr)
nc_wtr_union_subd <- nc_wtr_union %>%
st_subdivide( max_vertices = 256) %>%
st_collection_extract()
# PLOT OVERLAY ----
par(mar = rep(0,4))
plot(pts_sf, col=rgb(0, 0, 0, 0.33))
plot(nc_subd, col = "transparent",border = "red", add = TRUE)
plot(nc_bound, lwd = 2, border = "black", add = TRUE)
title("NC State Boundary", line = -5)
plot(pts_sf, col=rgb(0, 0, 0, 0.33))
plot(nc_wtr_union_subd, col = "transparent",border = "red", add = TRUE)
plot(nc_wtr_union, lwd = 2, col = "transparent",border = "black", add = TRUE)
plot(nc_bound, lwd = 2, border = "black", add = TRUE)
title("NC Watersheds", line = -5)
# BENCHMARK: ST_SUBDIVIDE IMPACT ON ST_INTERSECTS() ----
cols <- c("replications", "elapsed", "user.self", "sys.self")
# transform to North Caroline State Plane (meters)
nc_subd_NCSP <- st_transform(nc_subd, 32119)
nc_bound_NCSP <- st_transform(nc_bound, 32119)
nc_wtr_union_subd_NCSP <- st_transform(nc_wtr_union_subd, 32119)
nc_wtr_union_NCSP <- st_transform(nc_wtr_union, 32119)
pts_NCSP <- st_transform(pts_sf, 32119)
set.seed(7890)
# Without st_subdivided
benchmark(columns = cols, replications = 100, {
st_intersects(pts_NCSP, nc_bound_NCSP)
})
## replications elapsed user.self sys.self
## 1 100 9.3 9.25 0.02
# With st_subdivided
benchmark(columns = cols, replications = 100, {
st_intersects(pts_NCSP, nc_subd_NCSP)
})
## replications elapsed user.self sys.self
## 1 100 3.87 3.86 0.01
# Without st_subdivided
benchmark(columns = cols, replications = 10, {
st_intersects(pts_NCSP, nc_wtr_union_NCSP)
})
## replications elapsed user.self sys.self
## 1 10 203.47 203.1 0.02
# With st_subdivided
benchmark(columns = cols, replications = 10, {
st_intersects(pts_NCSP, nc_wtr_union_subd_NCSP)
})
## replications elapsed user.self sys.self
## 1 10 0.78 0.78 0
devtools::session_info()
## Session info -------------------------------------------------------------
## setting value
## version R version 3.4.0 (2017-04-21)
## system x86_64, mingw32
## ui RTerm
## language (EN)
## collate English_United States.1252
## tz America/Los_Angeles
## date 2017-12-16
## Packages -----------------------------------------------------------------
## package * version date source
## assertthat 0.2.0 2017-04-11 CRAN (R 3.4.2)
## backports 1.1.0 2017-05-22 CRAN (R 3.4.0)
## base * 3.4.0 2017-04-21 local
## bindr 0.1 2016-11-13 CRAN (R 3.4.2)
## bindrcpp 0.2 2017-06-17 CRAN (R 3.4.2)
## bitops 1.0-6 2013-08-17 CRAN (R 3.4.0)
## broom 0.4.3 2017-11-20 CRAN (R 3.4.3)
## cellranger 1.1.0 2016-07-27 CRAN (R 3.4.2)
## class 7.3-14 2015-08-30 CRAN (R 3.4.0)
## classInt 0.1-24 2017-04-16 CRAN (R 3.4.2)
## cli 1.0.0 2017-11-05 CRAN (R 3.4.2)
## colorspace 1.3-2 2016-12-14 CRAN (R 3.4.2)
## compiler 3.4.0 2017-04-21 local
## crayon 1.3.4 2017-10-30 Github (r-lib/crayon@b5221ab)
## curl 3.0 2017-10-06 CRAN (R 3.4.2)
## datasets * 3.4.0 2017-04-21 local
## DBI 0.7 2017-06-18 CRAN (R 3.4.2)
## devtools 1.13.2 2017-06-02 CRAN (R 3.4.0)
## digest 0.6.12 2017-01-27 CRAN (R 3.4.0)
## dplyr * 0.7.4 2017-09-28 CRAN (R 3.4.2)
## e1071 1.6-8 2017-02-02 CRAN (R 3.4.2)
## esri2sf * 0.1.0 2017-12-12 Github (yonghah/esri2sf@81d211f)
## evaluate 0.10 2016-10-11 CRAN (R 3.4.0)
## forcats * 0.2.0 2017-01-23 CRAN (R 3.4.2)
## foreign 0.8-67 2016-09-13 CRAN (R 3.4.0)
## ggplot2 * 2.2.1.9000 2017-12-02 Github (tidyverse/ggplot2@7b5c185)
## glue 1.2.0.9000 2017-12-05 Github (tidyverse/glue@69bc72c)
## graphics * 3.4.0 2017-04-21 local
## grDevices * 3.4.0 2017-04-21 local
## grid 3.4.0 2017-04-21 local
## gtable 0.2.0 2016-02-26 CRAN (R 3.4.2)
## haven 1.1.0 2017-07-09 CRAN (R 3.4.2)
## hms 0.4.0 2017-11-23 CRAN (R 3.4.3)
## htmltools 0.3.6 2017-04-28 CRAN (R 3.4.0)
## httr * 1.3.1 2017-08-20 CRAN (R 3.4.2)
## jsonlite * 1.5 2017-06-01 CRAN (R 3.4.0)
## knitr 1.16 2017-05-18 CRAN (R 3.4.0)
## lattice 0.20-35 2017-03-25 CRAN (R 3.4.0)
## lazyeval 0.2.1 2017-10-29 CRAN (R 3.4.2)
## lubridate 1.7.1 2017-11-03 CRAN (R 3.4.2)
## lwgeom * 0.1-1 2017-12-16 Github (r-spatial/lwgeom@baf22c6)
## magrittr 1.5 2014-11-22 CRAN (R 3.4.0)
## memoise 1.1.0 2017-04-21 CRAN (R 3.4.0)
## methods * 3.4.0 2017-04-21 local
## mnormt 1.5-5 2016-10-15 CRAN (R 3.4.1)
## modelr 0.1.1 2017-07-24 CRAN (R 3.4.2)
## munsell 0.4.3 2016-02-13 CRAN (R 3.4.2)
## nlme 3.1-131 2017-02-06 CRAN (R 3.4.0)
## parallel 3.4.0 2017-04-21 local
## pillar 0.0.0.9000 2017-12-10 Github (r-lib/pillar@5a082e1)
## pkgconfig 2.0.1 2017-03-21 CRAN (R 3.4.2)
## plyr 1.8.4 2016-06-08 CRAN (R 3.4.2)
## psych 1.7.8 2017-09-09 CRAN (R 3.4.2)
## purrr * 0.2.4.9000 2017-12-05 Github (tidyverse/purrr@62b135a)
## R6 2.2.2 2017-06-17 CRAN (R 3.4.0)
## rbenchmark * 1.0.0 2012-08-30 CRAN (R 3.4.1)
## Rcpp 0.12.14 2017-11-23 CRAN (R 3.4.2)
## RCurl 1.95-4.8 2016-03-01 CRAN (R 3.4.1)
## readr * 1.1.1 2017-05-16 CRAN (R 3.4.2)
## readxl 1.0.0 2017-04-18 CRAN (R 3.4.2)
## reshape2 1.4.2 2016-10-22 CRAN (R 3.4.2)
## rlang 0.1.4 2017-11-05 CRAN (R 3.4.2)
## rmarkdown 1.8 2017-11-17 CRAN (R 3.4.2)
## rprojroot 1.2 2017-01-16 CRAN (R 3.4.0)
## rvest 0.3.2 2016-06-17 CRAN (R 3.4.2)
## scales 0.5.0.9000 2017-12-02 Github (hadley/scales@d767915)
## sf * 0.5-6 2017-12-16 Github (r-spatial/sf@be20ebb)
## stats * 3.4.0 2017-04-21 local
## stringi 1.1.6 2017-11-17 CRAN (R 3.4.2)
## stringr * 1.2.0 2017-02-18 CRAN (R 3.4.0)
## tibble * 1.3.4.9003 2017-12-10 Github (tidyverse/tibble@60281b3)
## tidyr * 0.7.2.9000 2017-12-05 Github (tidyverse/tidyr@efd9ea5)
## tidyverse * 1.2.1 2017-12-10 Github (tidyverse/tidyverse@3769ff2)
## tools 3.4.0 2017-04-21 local
## udunits2 0.13 2016-11-17 CRAN (R 3.4.1)
## units 0.4-6 2017-08-27 CRAN (R 3.4.2)
## utils * 3.4.0 2017-04-21 local
## withr 2.1.0.9000 2017-12-02 Github (jimhester/withr@fe81c00)
## XML 3.98-1.9 2017-06-19 CRAN (R 3.4.1)
## xml2 1.1.1 2017-01-24 CRAN (R 3.4.2)
## yaml 2.1.14 2016-11-12 CRAN (R 3.4.0)
Impressive!
Please note that in PostGIS 2.5 Subdivide algorithm was updated and now reselects pivot not as midpoint but as closest point to it of the polygon, starting from largest holes.
Was this function ever added to sf @edzer ? I can't seem to find it, so I guess not?
Ah, I just found that it is part of r-spatial/lwgeom . Apologies. Leaving this comment here for anyone else searching.
Most helpful comment
Here's two more benchmarks - this time testing the speed of
st_intersects()with a largesfc_POINTobject and:The first tests show modest speed improvements (~3x) while the second test shows _dramatic_ improvement (~300x).
Reprex + Session info