Seurat: JackStraw producing insignificant results with SCTintegration
Hi-
Firstly, just wanted to note how useful SCTransform has been and thank you for that tool. I'm posting in regards to the new developer version with the PrepSCTintegration pipeline. I've integrated 14 datasets (~30 k) cells with it and when I previously integrated the datatsets (prior to 3.1), Jackstraw was indicating ~50 significant PC's. Now with SCT, it produces probabilities of 1 as early as PC 7. Here is what I ran:
for (i in 1:length(object.list)) {
object.list[[i]] <- SCTransform(object.list[[i]], vars.to.regress ="percent.mt", verbose = F)
}
int.features <- SelectIntegrationFeatures(object.list = object.list, nfeatures = 3000)
object.list<- PrepSCTIntegration(object.list = object.list, anchor.features = int.features)
int.anchors <- FindIntegrationAnchors(object.list = object.list, anchor.features = int.features)
integrated.object <- IntegrateData(anchorset = int.anchors, normalization.method = "SCT")
After integration, I ran PCA --> Jackstraw --> tSNe as I normally would without ScaleData as demonstrated in the vignette.
All 6 comments
@jad362
Similar issues, I have fewer significant PCs using SCTransform and I don't know whether I should choose PC based on Jackstraw.
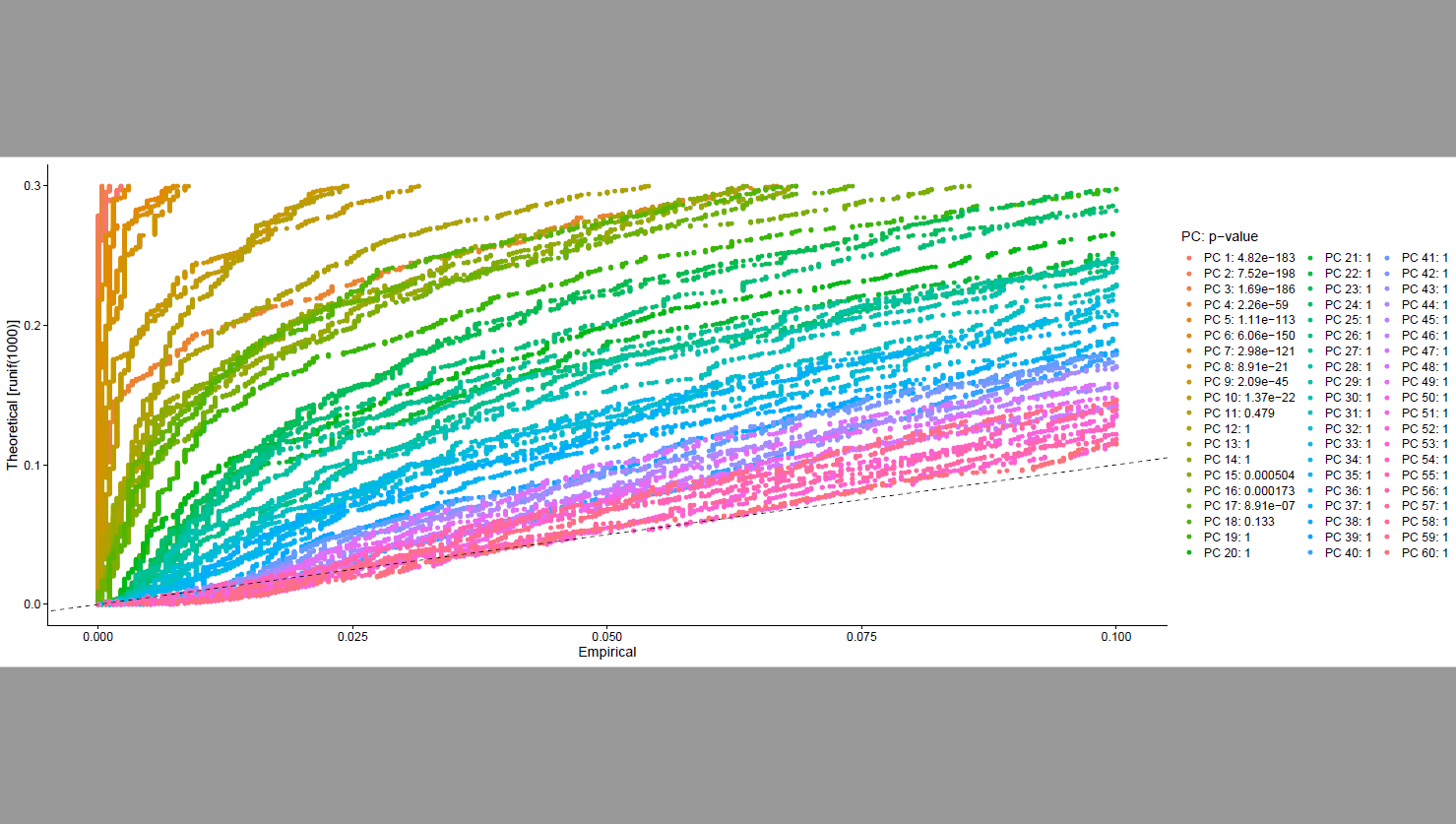
One thing which confuse me, PCs which are above the dashed line seems to be significant PC but p-value are 1.
Similar issue. I got nearly 1/3 significant PC number compared to the regular protocol in almost every dataset.
One thing which confuse me, PCs which are above the dashed line seems to be significant PC but p-value are 1.
I got a similar result which also confuses me. How does pc14 have a p-value of 1 then pc15 turns out to be significant..?
Apologies that it took us so long to respond to this. Unfortunately the JackStraw procedure as implemented does not work well on SCT-transformed data. This is because JackStraw, as we implemented it, assumes each gene has equal variance - this is not true for SCT normalization (where gene variance is weighted by biological heterogenetiy). We are disabling the ability to perform JackStraw on SCT-normalized data (@timoast)
This is now updated on the develop branch
Most helpful comment
One thing which confuse me, PCs which are above the dashed line seems to be significant PC but p-value are 1.