Seurat: Leiden algorithm cast SNN to dense
FindClusters has an option for the leiden algorithm, but as far as I can tell it casts the adjacency matrix to a dense matrix prior to generating the graph. I assume this is required given the way leidenalg is being called currently, but do you have any suggestions for large datasets where this cast to dense is more prohibitive? No worries if not, and realize might be lower on priority list, just wanted to check!
All 16 comments
It's a good point. @TomKellyGenetics do you think there's a way to avoid casting to a dense matrix here?
It should be possible to pass an SNN graph directly to the python package (which takes this as an input). I'm open to implementing this but will take some time to look into how to go about it. At the moment, the "leiden" package in R avoids using the igraph library deliberately to reduce the size of the install.
See issue on the standalone R package: https://github.com/TomKellyGenetics/leiden/issues/1
Currently, the adjacency matrix is passed to python, then the SNN graph is computed in Python which is compatible with the Python leidenalg. For raw adjacency matrices, this was faster than computing SNN graphs in R (but could be reconsidered for Seurat objects). If it is worthwhile for performance and handling large datasets, I'm okay to add support for graph methods and igraph dependancies back in (Seurat has this dependency already). There is good support for basic data structures such as a matrix to be passed between R and Python but I'm not sure it's as simple for the separate igraph implementations. A sparse "dgCmatrix" can also be passed to python (compatible with the current leiden version) but this doesn't seem to improve performance.
On the other hand, if performance is a concern, these changes would be incremental compared to a large overhaul using RCpp to cut out Python dependency altogether (I'm not experienced with C++). This is also being considered and has been discussed with @vtraag and others on social media. I had someone offer to help with this but we've been short on time to implement it: https://github.com/TomKellyGenetics/leiden/issues/4
In short, it's _possible_ but I'm not sure how long it will take to make major changes. Pull Requests are welcome on the standalone package and we can migrate them onto the Seurat version once it is working.
Just revisiting this issue to see if there are any more thoughts about it. Is there an urgent need to optimise the R implementation of Leiden for larger matrices? This would be quite involved but I'm open to suggestions or Pull Requests.
Sorry for the very late reply, I definitely don't have an urgent need for
it, was just looking to try out comparisons of Leiden to louvain for a few
datasets, nothing time sensitive. With current version casting the SNN
matrix to dense results in memory related errors for large datasets even
with fairly large amount of memory available at least when I've tried it,
so mostly opened issue bc of that vs other aspects of performance.
On Fri, Jul 5, 2019 at 1:17 AM Tom Kelly notifications@github.com wrote:
Just revisiting this issue to see if there are any more thoughts about it.
Is there an urgent need to optimise the R implementation of Leiden for
larger matrices? This would be quite involved but I'm open to suggestions
or Pull Requests.—
You are receiving this because you authored the thread.
Reply to this email directly, view it on GitHub
https://github.com/satijalab/seurat/issues/1645?email_source=notifications&email_token=ABER4A2LL5KEKRAUIH5P3K3P53KONA5CNFSM4HUVFDDKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGODZIRZIQ#issuecomment-508632226,
or mute the thread
https://github.com/notifications/unsubscribe-auth/ABER4A4B7XGUAZHYJJ4K76DP53KONANCNFSM4HUVFDDA
.
Thanks for the explanation. I suspect that the R version of the Leiden algorithm will need to be improved to handle larger datasets as they are generate but it will suffice for typical single-cell analyses at the moment. Out of curiosity, how large were the SNN matrices you were trying? Have you tried to run it on the Python version?
I will look into whether SNN graphs can be passed directly to the Python implementation to reduce memory load.
The dataset I was trying it out with was roughly 200k cells and I haven't
tried the python version yet -- will be traveling for a few weeks but can
look at in more detail when get back if would be helpful.
On Sun, Jul 7, 2019, 9:18 PM Tom Kelly notifications@github.com wrote:
Thanks for the explanation. I suspect that the R version of the Leiden
algorithm will need to be improved to handle larger datasets as they are
generate but it will suffice for typical single-cell analyses at the
moment. Out of curiosity, how large were the SNN matrices you were trying?
Have you tried to run it on the Python version?I will look into whether SNN graphs can be passed directly to the Python
implementation to reduce memory load.—
You are receiving this because you authored the thread.
Reply to this email directly, view it on GitHub
https://github.com/satijalab/seurat/issues/1645?email_source=notifications&email_token=ABER4A3FEPIOFGXAIYXEOU3P6KIXDA5CNFSM4HUVFDDKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGODZLXHZY#issuecomment-509047783,
or mute the thread
https://github.com/notifications/unsubscribe-auth/ABER4A45YW2XXNLI7B2AXCDP6KIXDANCNFSM4HUVFDDA
.
Thanks, it would be good to make sure that the Python package can handle datasets this large. I'm busy with other projects but I'll try make time to look into it. I suspect datasets that large will become more common as the HCA datasets start to roll out.
Apologies for hijacking this thread but I have a question related to the leidenalg algorithm aswell. I was wondering in Seurat V3 if it was possible/correct to pass the SNN graph values (object@graphs$RNA_snn@x) as weights or whether this was already done automatically using the default settings. I am not as well versed in the algebra or the programming language.
I hope to hear from you soon.
regards
At the time that the Leiden algorithm PR was submitted, the R version did not support weights and only run unweighted SNN graphs (same as the Python defaults). This was added merge merging to update it with improved passing arguments to the Python library library including weights (but this hasn't been extensively tested). At the moment, weights are optional to run Leiden and are not computed by Seurat. Please see the documentation for the Python version for more detail on this parameter: https://leidenalg.readthedocs.io/en/latest/reference.html#modularityvertexpartition
Quick update on this. I’ve figured out how to pass vertices and edges (possibly weights) directly to the python. I’ll implement this as methods in the stand-alone package first. I’d propose to migrate the RunLeiden internal function to calling the leiden R package as a dependency as it is now on CRAN. I’ll submit Issues and PRs specifically about these proposals in due time.
Thank you for the quick response. Before using leiden algorithm in Seurat V3 I was using it in python's "scanpy" package. The module uses weights by default . If I am not wrong this gets computed from the neighbors graph and connectivities represent the weights .
I have been trying to reproduce my python analysis in R using Seurat but wasn't able to do so most likely because of this issue.
regards
I've got a method of the igraph objects working on the development branch of the "leiden" package. This passes edges between igraph objects in R and Python. Hopefully it will be more more efficient for memory with large sparse matrices/SNNs. Unfortunately, there it is slower on small matrices (this is a test on the Zachary data from igraph). Anyway, both methods are setup now so it's more flexible, I'll get an update on CRAN before we decide how to go about calling it within Seurat.
@heeroena this is mainly directed at the memory-issues raised above but the update should enable weighted graph objects as well (it passes weights from the graph object to python leidenalg).
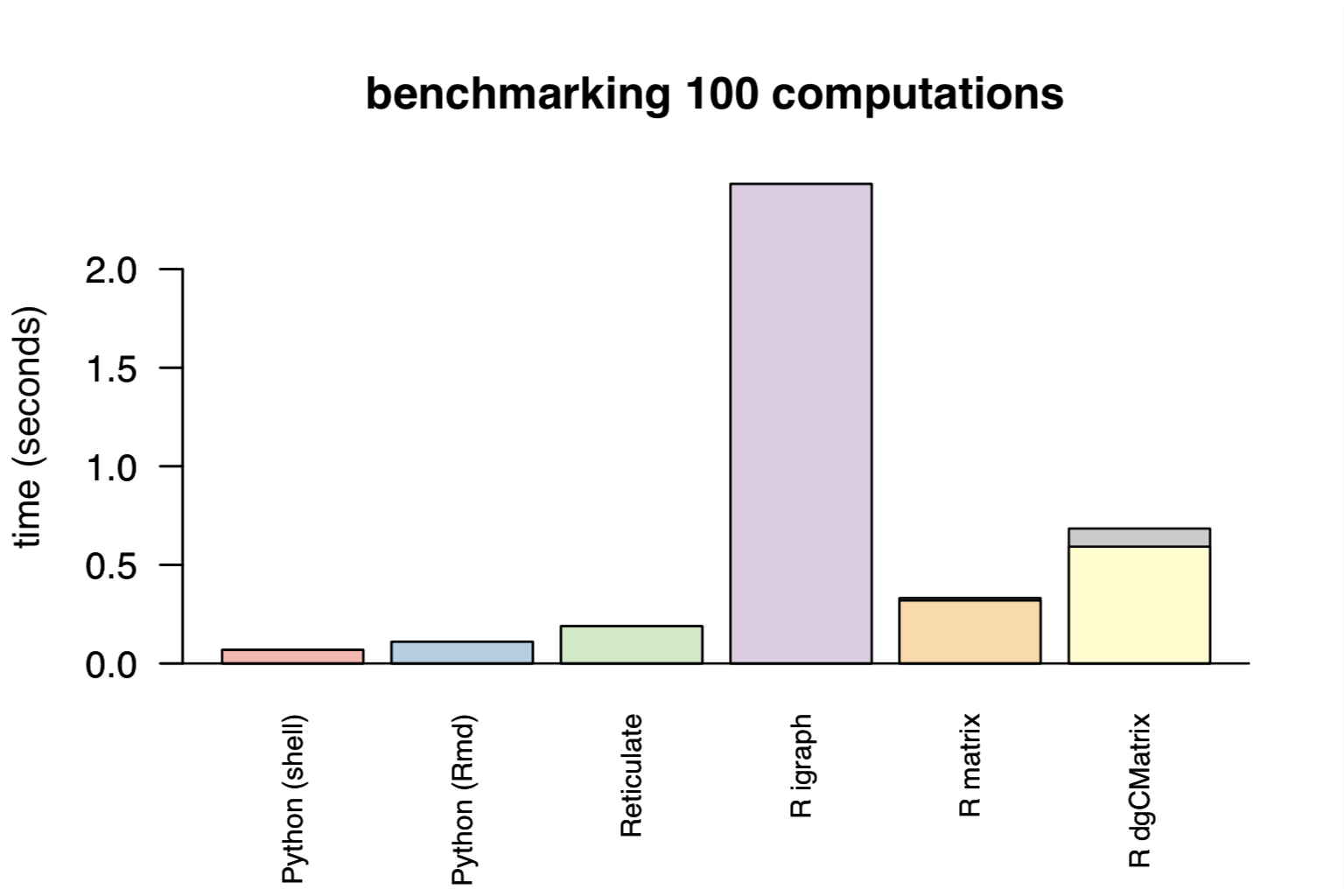
The leiden 0.3.0 R package has been submitted to CRAN implementing these features. I would recommend calling this as a dependency rather than the internal RunLeiden function (that I wrote). While it is slower, it will avoid computing dense matrices and memory issues by passing edges Python (without casting to dense). I will submit a PR to the Seurat development branch once it is accepted on CRAN.
The compatible update (version 0.3.1) is up on CRAN, it should be available for windows in a few days. I've submitted a PR to call in by FindClusters (#1858). This version allows setting the random.seed and and n.iters (just like the RunModularityClusters). The release also supports methods for sparse matrices.
Please note that for small matrices, it is faster to cast to dense and pass an R matrix to a numpy.array. This is still the default behaviour but sparse matrices and graph objects can be passed to python without casting to dense (with methods for dgCMatrix and igraph) to avoid memory issues.
@andrewhill157 I think we can close this now. It's available in the develop branch of Seurat and will be up in the next release. Thanks for prodding me to revisit it. 😄
Also heard that @vtraag is working on adding cluster_leiden to the C igraph library (and by extension will be available in python-igraph or r-igraph). This version may may better performance in the future.
This should be fixed in the development version of Seurat. To install the development version of Seurat, please see the instructions here.
Most helpful comment
Quick update on this. I’ve figured out how to pass vertices and edges (possibly weights) directly to the python. I’ll implement this as methods in the stand-alone package first. I’d propose to migrate the RunLeiden internal function to calling the leiden R package as a dependency as it is now on CRAN. I’ll submit Issues and PRs specifically about these proposals in due time.