Seurat: The min expression levevl in a VlnPlot is not zero, but negative value
When type in
VlnPlot(object = pbmc_small, features = 'PC_1', slot = "counts")
you can see what happen. It does bother me. Needding your help...
All 4 comments
Hi,
This is a bit of a misnomer in the plot: you're plotting the embedding values for PC1, not expression of a feature. These values are allowed to be negative (look at a DimPlot with reduciton = 'pca') so it's no concern that the violin plot show negative values. We've made some changes to VlnPlot (and RidgePlot) so that the axis labels more accurately describe what they're showing. These changes are available in the development version of Seurat. To install the development version of Seurat, please see the instructions here.
Hello,
I have a related question. I am plotting gene expression levels using VlnPlot but with some of them I am getting negative values. The data has been normalised and scaled but I don't remember ever specifying any controls - so if these values are log2 fold change, I'm not sure what they are relative to. Could you explain why I sometimes get negative expression levels?
The code I am using is here. To execute it, change line 8 of global_variables.R to the location where you downloaded and then execute explore.R. This should pickup the Seurat object from the RDS file in SavedObjects and plot a bunch of VlnPlot's.
Thanks
Ciaran
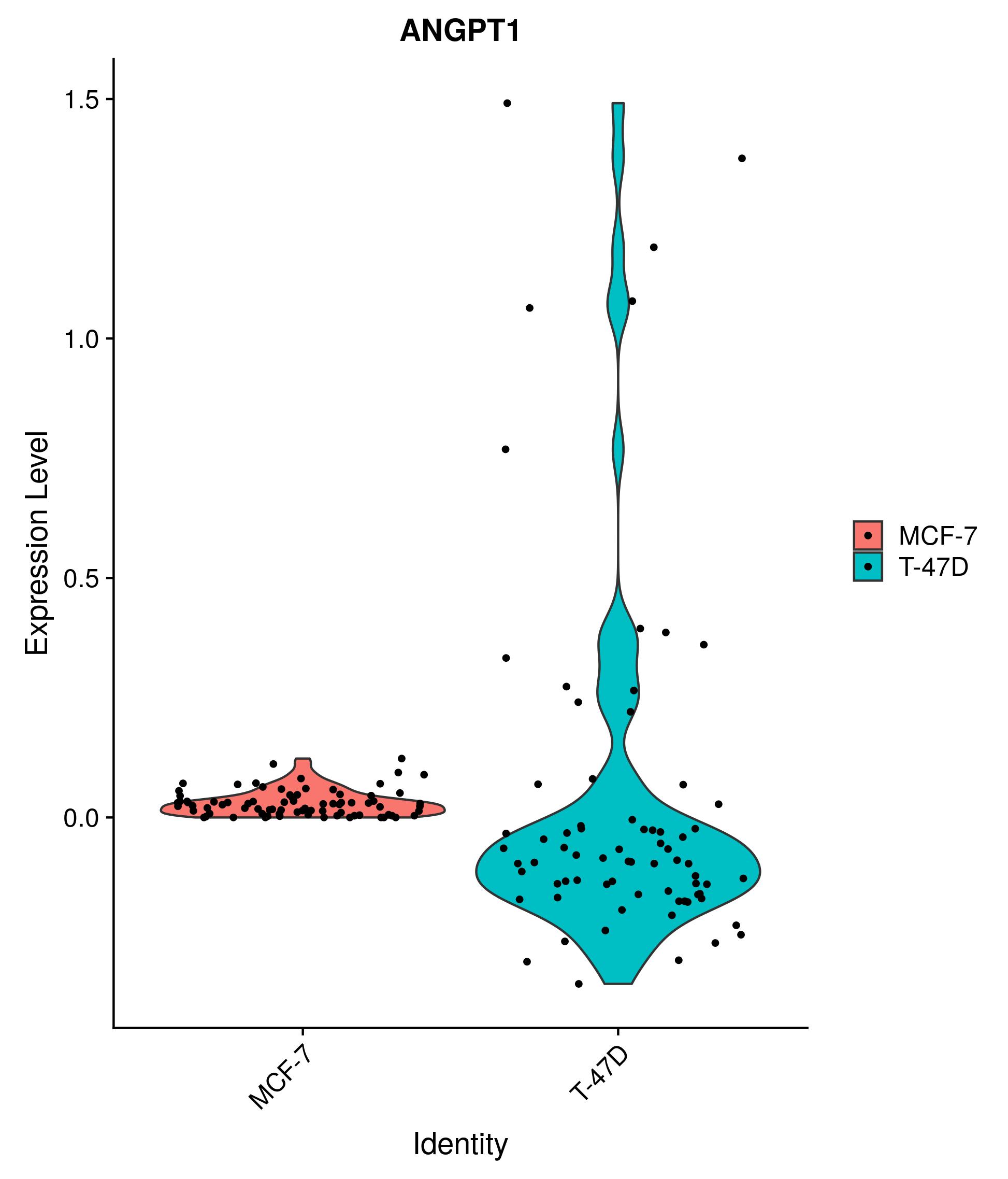
yeah, I have solved this problem by asigning the paramater of assay. Just look at this code: VlnPlot(..., assay = "RNA"), it may help you!
Excellant, thank you.
Most helpful comment
yeah, I have solved this problem by asigning the paramater of assay. Just look at this code: VlnPlot(..., assay = "RNA"), it may help you!