Seurat: v3.0 Problem removing Legend and Axes in FeaturePlot
Hello,
I am trying to remove the legends and axes from the FeaturePlot using v3.0
This is what I tried:
plot <- FeaturePlot(object = seuratobject, features = gene.list1) + NoLegend() + NoAxes()
print(plot)
But it still return a plot that includes legends and axes.
Thanks in advance for any help.
All 5 comments
When you plot >1 feature, FeaturePlot combines the plots using cowplot::plot_grid. After the plots have been combined you can't change things like the legend and axes. Instead you can use the combine=FALSE argument, so that the plots are not combined, then change each individual plot, then combine the altered plots. Here is an example with the pbmc dataset:
library(Seurat)
p <- FeaturePlot(pbmc_small, head(VariableFeatures(pbmc_small)), combine = FALSE)
for(i in 1:length(p)) {
p[[i]] <- p[[i]] + NoLegend() + NoAxes()
}
cowplot::plot_grid(plotlist = p)
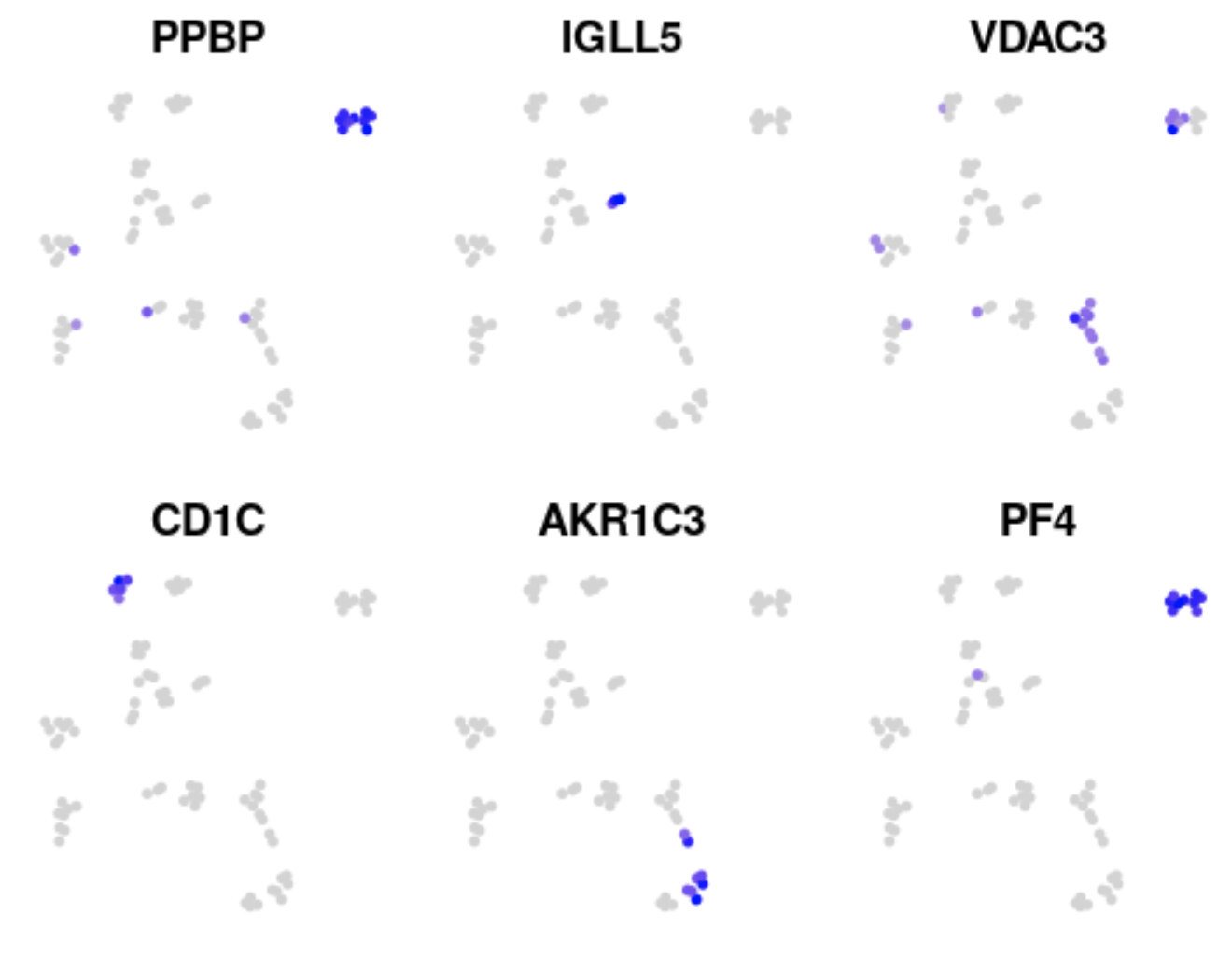
Thank you
Would it be possible to bring back the no.axes and no.legend arguments to FeaturePlot() and DimPlot()? Both seem like a much more elegant way to solve this problem.
I was able to use
FeaturePlot(object = object, features = features, combine = TRUE) & NoLegend() & NoAxes()
Yes, we now use patchwork for combining plots which allows easier modification of the individual panels using the & operator
Most helpful comment
Would it be possible to bring back the no.axes and no.legend arguments to FeaturePlot() and DimPlot()? Both seem like a much more elegant way to solve this problem.