Seurat: How to find the difference gene between samples
Hi, I have 3 samples. I use MergeSeurat to add them into one object. How I can find the difference gene between samples, not between different clusters? Thanks a lot.
All 8 comments
Hi,
One way to do this is to use the SetAllIdent() function of Seurat, and attribute to each cell its sample ID as its cluster identity:
# Save current cluster identites in [email protected] under 'clusterID'
# Run only if Seurat::FindClusters() was executed
object <- Seurat::StashIdent(object = object, save.name = "clusterID")
# Randomly attribute a sample identity to each cell
sampleID <- sample(x = c(1,2,3), size = length([email protected]), replace = TRUE)
# Add sample identity to [email protected] under 'sampleID'
[email protected]$sampleID <- sampleID
# Set cell identity to sample identity
object <- Seurat::SetAllIdent(object = object, id = "sampleID")
# Find all sample specific marker genes
markers <- Seurat::FindAllMarkers(object = object)
Hope this helped!
Best,
Leon
Hi, Leon,
Thank you so much. I will check it.
Best,
Lili
@leonfodoulian
Dear Leon,
I try to run it as your suggestion. But it give me error:
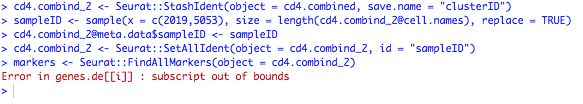
Could you help me to check it? Thanks a lot.
Best,
Lili
Hi Lili,
I don't think you have to use sample() in your case. The code I wrote was just a demonstration. In your case, you should attribute to each cell its real known sample ID. Regarding the error, it is rather related to the FindAllMarkers() function of Seurat. Could you please run traceback() after you get the error to have more information on what went wrong? Also what is the version of Seurat that you have? Sharing your sessionInfo() can also be helpful in this case.
Best,
Leon
Hi,
I might have found the source of the error. I think you have Seurat version 2.0.1 (or maybe earlier). You might want to update to version 2.1.0 or later. Simply said, since you randomly attributed a sample ID to each of the cells, when computing differential expression, at least one of the samples had no upregulated/marker genes. Seurat version 2.0.1 (and earlier) didn't check for this appropriately. This was later updated in Seurat version 2.1.0 and later. The code snippet related to this error can be found in lines 391 to 394 in differential_expression.R. In Seurat version 2.0.1 (and earlier), line 394 preceded lines 391 to 393.
Hope when updating to the latest version of Seurat, this will eventually work.
Best,
Leon
@leonfodoulian
Dear Leon,
Thank you so much. Yes, I have Seurat version 2.0.1. I will update my Seurat and check it. Thanks a lot.
Best,
Lili
We see this question a lot. Leon's answer is great, and you can also look at http://satijalab.org/seurat/immune_alignment.html ("Identify differential expressed genes across conditions")
As this comes up in search and has outdated information, please see @vertesy description for Seurat >v3 here #252 also see #1076
A current solution for v3 would be:
# Setting idents so that DE analysis is performed between samples rather than clusters
Idents(object = mySeruatObject) <- [email protected]$sample
# To shift back to compare clusters rather than samples (you could have clusters named differently,
# something like integrated_snn_res0.4 or seurat_clusters, so take the time to go to your metadata
# file and see what the cluster name is)
# You can see your metadata files by doing [email protected]$ or clicking on your
# Seurat object if you use Rstudio
Idents(object = mySeruatObject) <- [email protected]$clusters
I apologize to Seurat Devs for this is redundant cyclic information.
Most helpful comment
Hi,
One way to do this is to use the
SetAllIdent()function of Seurat, and attribute to each cell its sample ID as its cluster identity:Hope this helped!
Best,
Leon