Scikit-image: peak_local_max fails to enforce min_distance
Description
I am trying to extract local maxima from a distance transform (made using openCV). When I run: coordinates = peak_local_max(distance_xform, min_distance=100, num_peaks=12)
The following values are in coordinates: [[2775 2647] [944 1540] [919 2420] [1486 1192] [899 2099] [900 2054] [900 2055] [900 2056] [900 2057] [900 2058] [900 2059] [900 2060]]
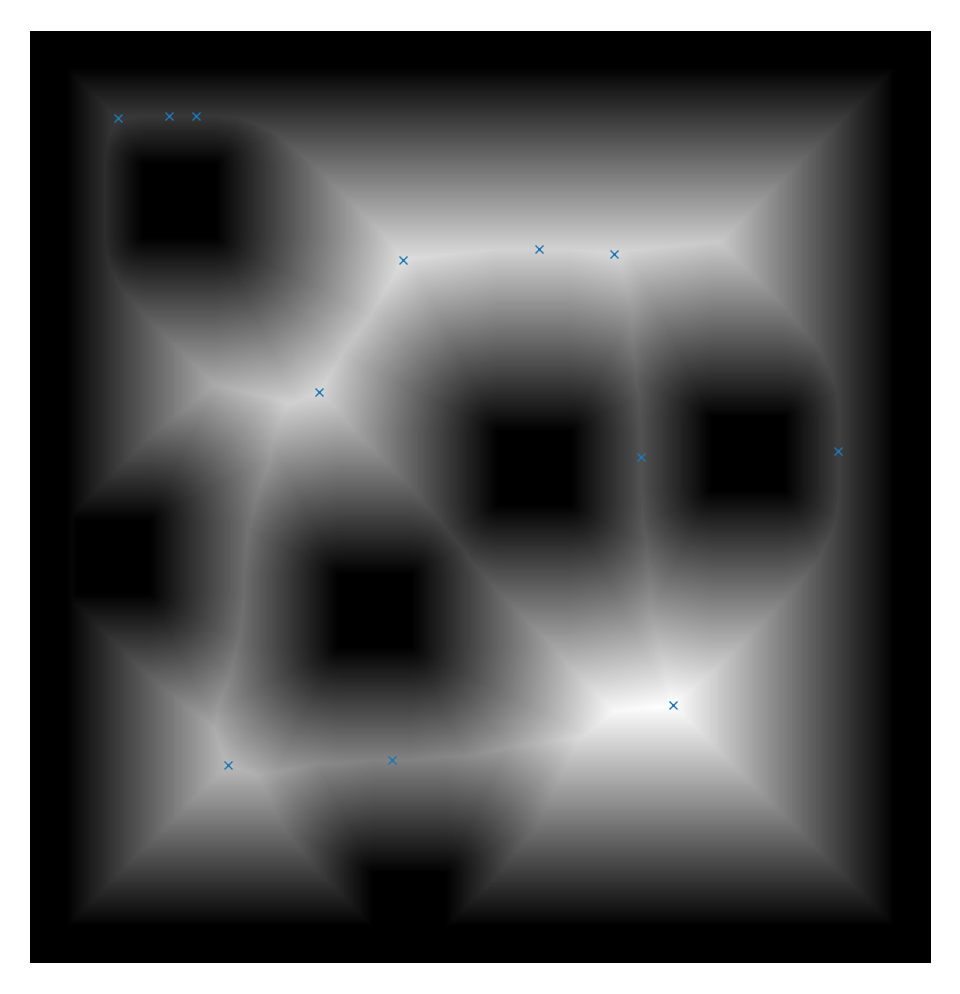
If you look at the distance transform image there is a line one pixel wide on the horizontal axis extending from 1922 to 2228 at 900. All pixels on this line have the same value. It seems this causes peak_local_max to ignore min_distance and pick peaks 1 pixel adjacent to each other along this line.
I am using:
Ubuntu 14.04.2 LTS
skimage 0.9.3
Python 2.7.6
[If reporting a bug, attach the entire traceback from Python.] (No traceback: functional error)
[If proposing an enhancement/new feature, provide links to related articles, reference examples, etc.]
Way to reproduce
[If reporting a bug, please include the following important information:]
- [x] Code example
- [x] Relevant images (if any)
- [x] Operating system and version
- [x] Python version
- [x] scikit-image version (run
skimage.__version__)

All 5 comments
@jfflanagan do you have the option to update skimage? That is an old version (3.5 years old; we just released 0.13), and I'm pretty sure that issue was fixed in the interim.
I switched to skimage 0.14dev and problem persists. Box positions are a little different: [[2775 2647] [944 1540] [919 2420] [1486 1192] [900 2125] [900 2050] [900 2052] [900 2053] [900 2055] [900 2056] [900 2057] [900 2058]]
Isn't corner_peaks what we are looking for here, or am I missing something? Can't this issue be closed then?
An even worst illustration of this bug is when min_distance is larger then the "radius" of the footprint:
>>> img = np.zeros((15, 15))
>>> img[5, 5] = 1
>>> img[9, 9] = 2
>>> min_distance = 5
>>> footprint = np.ones((7, 7))
>>> out1 = peak_local_max(img, min_distance=min_distance, indices=False)
>>> out2 = peak_local_max(img, footprint=footprint, indices=False)
>>> out3 = peak_local_max(img, min_distance=min_distance, footprint=footprint, indices=False)
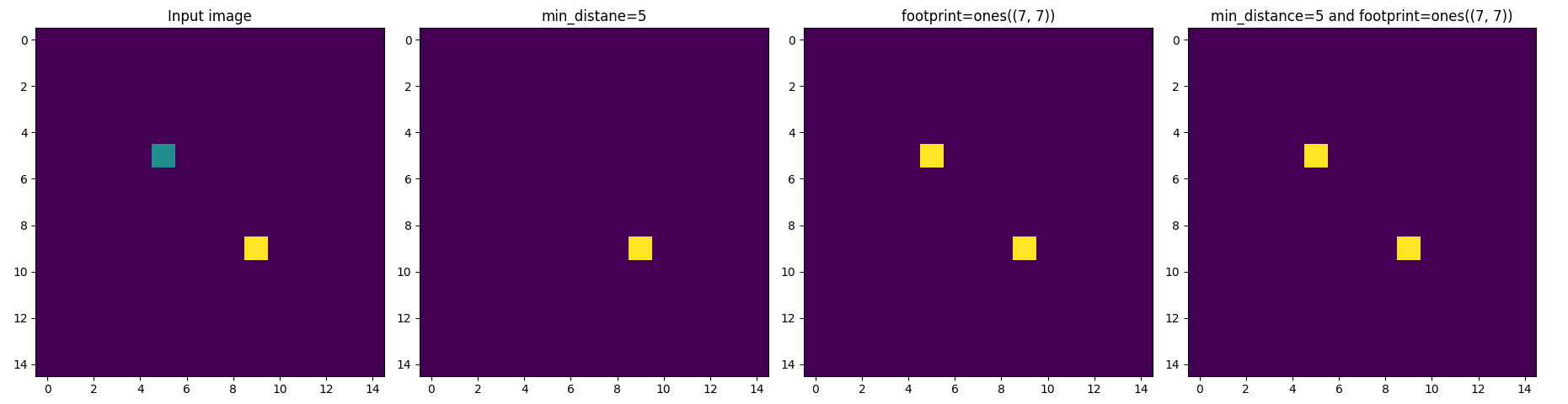
In fact, min_distance is ignored when footprint is not None (please see the _get_peak_mask function)
I wonder if this shouldn't be reported in a separate issue since the same observed behavior is caused by two different reasons...
I performed some experiments:
import imageio
import skimage
from skimage import feature, img_as_float
import numpy as np
from time import time
from matplotlib import pylab as plt
img = imageio.imread('~/Images/test_img.jpg')
t0 = time()
coords = feature.peak_local_max(
img, min_distance=100, num_peaks=12
)
t1 = time()
print(f"version {skimage.__version__}: {t1-t0} seconds")
plt.gray()
plt.imshow(distance)
plt.plot(coords[:, 1], coords[:, 0], "x")
plt.axis('off')
plt.tight_layout()
plt.show()
Here are the results:
version 0.17.2
version 0.17.2: 0.28 seconds

with #4760 refactoring
version 0.18.dev0: 0.79 seconds
