Mmdetection: inference error when use config file ms_rcnn_x101_64x4d_fpn_1x.py
Thanks for your error report and we appreciate it a lot.
Checklist
1.I'm not find the bug has been fixed
- en...... the amount of the issues is so much
Describe the bug
an error occured when i inference use ms_rcnn_x101_64x4d_fpn_1x model,the detail of the error is as follows:
Traceback (most recent call last):
File "infer_oct_binary_ms.py", line 389, in
dic_img_name_to_disease_list=inference_OCT(config_file,checkpoint_file,out_file,test_img_dir,score_thr,normal_img_dir)
File "infer_oct_binary_ms.py", line 27, in inference_OCT
out_file=out_file+'/{}'.format(img_name))
File "/home/lwc/GitHub/mmdetection/mmdet/apis/inference.py", line 136, in show_result
img[mask] = img[mask] * 0.5 + color_mask * 0.5
IndexError: boolean index did not match indexed array along dimension 2; dimension is 3 but corresponding boolean dimension is 2
I found that the shape of the mask is (494, 765, 2) ,and the shape of the img is (494, 765, 3) in ms ,for comparision, the inference for cascade mask rcnn model is runned again,and the shape of the mask is (494, 765)
Reproduction
- What command or script did you run?
A placeholder for the command.
- Did you make any modifications on the code or config? Did you understand what you have modified?
I just add return for the result without modifying the code. - What dataset did you use?
None
Environment
I think it's none of the environment's business
- OS: [e.g., Ubuntu 16.04.6]
- GCC [e.g., 5.4.0]
- PyTorch version [e.g., 1.1.0]
- How you installed PyTorch [e.g., pip, conda, source]
- GPU model [e.g., 1080Ti, V100]
- CUDA and CUDNN version: CUDA 10.1 CUDNN:7.1
- [optional] Other information that may be related (such as
$PATH,$LD_LIBRARY_PATH,$PYTHONPATH, etc.)
Error traceback
If applicable, paste the error trackback here.
dic_img_name_to_disease_list=inference_OCT(config_file,checkpoint_file,out_file,test_img_dir,score_thr,normal_img_dir)
File "infer_oct_binary_ms.py", line 27, in inference_OCT
out_file=out_file+'/{}'.format(img_name))
File "/home/lwc/GitHub/mmdetection/mmdet/apis/inference.py", line 136, in show_result
img[mask] = img[mask] * 0.5 + color_mask * 0.5
IndexError: boolean index did not match indexed array along dimension 2; dimension is 3 but corresponding boolean dimension is 2
Bug fix
None
All 5 comments
def inference_OCT(config_file,checkpoint_file,out_file,test_img_dir,score_thr=0.7,normal_img_dir=''):
model = init_detector(config_file, checkpoint_file, device='cuda:0')
img_detect=0
glob_list=glob.iglob(test_img_dir + '/*.' + '*')
img_list=[]
img_list.extend(glob_list)
dic_img_name_to_disease_list=defaultdict(list)
if os.path.exists(normal_img_dir):
glob_list_normal=glob.iglob(normal_img_dir+'/*.*')
img_list.extend(glob_list_normal)
for i, result in enumerate(inference_detector(model, img_list)):
img_name=os.path.basename(img_list[i])
img,bboxes,labels=### show_result(img_list[i], result, model.CLASSES,score_thr,\
out_file=out_file+'/{}'.format(img_name))
pre_label_lst=imshow_bboxes(img,bboxes,labels,model.CLASSES,score_thr)
base_img_name=os.path.basename(img_list[i])
dic_img_name_to_disease_list[base_img_name]=pre_label_lst
img_detect+=1
print('img_detected:',img_detect,' ',img_list[i],'-----',pre_label_lst)
result_file=os.path.join(out_file,'dic_img_name_to_disease_name_list.pk')
with open(result_file,'wb') as f:
pkl.dump(dic_img_name_to_disease_list,f)
print('done!')
return dic_img_name_to_disease_list
show_result in mmdetection/mmdet/apis/inference.py
def show_result(img,
result,
class_names,
score_thr=0.3,
wait_time=0,
out_file=None):
"""Visualize the detection results on the image.
Args:
img (str or np.ndarray): Image filename or loaded image.
result (tuple[list] or list): The detection result, can be either
(bbox, segm) or just bbox.
class_names (list[str] or tuple[str]): A list of class names.
score_thr (float): The threshold to visualize the bboxes and masks.
wait_time (int): Value of waitKey param.
out_file (str, optional): If specified, the visualization result will
be written to the out file instead of shown in a window.
"""
assert isinstance(class_names, (tuple, list))
img = mmcv.imread(img)
if isinstance(result, tuple):
bbox_result, segm_result = result
else:
bbox_result, segm_result = result, None
bboxes = np.vstack(bbox_result)
# draw segmentation masks
if segm_result is not None:
segms = mmcv.concat_list(segm_result)
inds = np.where(bboxes[:, -1] > score_thr)[0]
for i in inds:
color_mask = np.random.randint(0, 256, (1, 3), dtype=np.uint8)
mask = maskUtils.decode(segms[i]).astype(np.bool)
img[mask] = img[mask] * 0.5 + color_mask * 0.5
# draw bounding boxes
labels = [
np.full(bbox.shape[0], i, dtype=np.int32)
for i, bbox in enumerate(bbox_result)
]
labels = np.concatenate(labels)
mmcv.imshow_det_bboxes(
img.copy(),
bboxes,
labels,
bbox_color='white',
text_color='white',
class_names=class_names,
score_thr=score_thr,
show=out_file is None,
wait_time=wait_time,
out_file=out_file)
return img.copy(),bboxes,labels
def imshow_bboxes(img,bboxes,labels,class_names=None,score_thr=0,
bbox_color='green',
text_color='green',
thickness=1,
font_scale=0.5,
win_name='',
out_file=None):
"""Draw bboxes and class labels (with scores) on an image.
Args:
img (str or ndarray): The image to be displayed.
bboxes (ndarray): Bounding boxes (with scores), shaped (n, 4) or
(n, 5).
labels (ndarray): Labels of bboxes.
class_names (list[str]): Names of each classes.
score_thr (float): Minimum score of bboxes to be shown.
bbox_color (str or tuple or :obj:`Color`): Color of bbox lines.
text_color (str or tuple or :obj:`Color`): Color of texts.
thickness (int): Thickness of lines.
font_scale (float): Font scales of texts.
show (bool): Whether to show the image.
win_name (str): The window name.
wait_time (int): Value of waitKey param.
out_file (str or None): The filename to write the image.
"""
assert bboxes.ndim == 2
assert labels.ndim == 1
assert bboxes.shape[0] == labels.shape[0]
assert bboxes.shape[1] == 4 or bboxes.shape[1] == 5
if score_thr > 0:
assert bboxes.shape[1] == 5
scores = bboxes[:, -1]
inds = scores > score_thr
bboxes = bboxes[inds, :]
labels = labels[inds]
label_list=[]
for bbox, label in zip(bboxes, labels):
bbox_int = bbox.astype(np.int32)
label_text = class_names[
label] if class_names is not None else 'cls {}'.format(label)
if label_text not in label_list:
label_list.append(label_text)
if len(bbox) > 4:
label_text += '|{:.02f}'.format(bbox[-1])
return label_list
when i use high level api ,ms_rcnn_x101_64x4d_fpn_1x model throws error like:
ValueError: cannot reshape array of size 0 into shape (139703259679624,72200400,0)
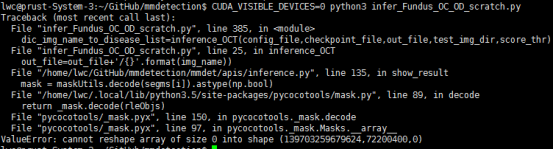
this error still remains after mmdetection is updated.
@tjsongzw
Thanks for reporting the bug. This is the temporary solution.
result = inference_detector(model, img)
result = (result[0], result[1][0])
Did anyone found a solution for this?
I'm facing same issue
when you use mask score R-CNN for inference, extra code should be added after inference_detector function to get the result object, the extra code is result = (result[0], result[1][0]),just as i said before
Most helpful comment
Thanks for reporting the bug. This is the temporary solution.