Mask_rcnn: Error while loading my own dataset(images) #KeyError: 'regions'
hi every body, i made a dataset, but has error when i load data
Traceback (most recent call last):
File "catvsdog.py", line 373, in
train(model)
File "catvsdog.py", line 196, in train
dataset_val.load_cat_dog(args.dataset, "val")
File "catvsdog.py", line 117, in load_cat_dog
annotations = [a for a in annotations if a['regions']]
File "catvsdog.py", line 117, in
annotations = [a for a in annotations if a['regions']]
KeyError: 'regions'
i reference https://github.com/matterport/Mask_RCNN/issues/1045
i try to editor my dataset code
{"1.jpeg4055": {"filename": "25.jpg","size": 22699,"regions":[{"shape_attributes":{"name":"rect","x":68,"y":23,"width":106,"height":156},"region_attributes": {"head": "head"}}],"file_attributes": {}}}
but error still had
All 12 comments
hi @TwinkleStars1029 its lool like your JSON file doensn't the same formatting as the required. Did you try to change it?
it helped me
Hi @flit86
Could you please share which part of your JSON did you change to match the formatting required by the code?
I am trying to utilize the Mask RCNN to identify buildings from images.
This is the error that keeps popping up:
2019-07-08 19:04:21.815598: W tensorflow/compiler/jit/mark_for_compilation_pass.cc:1412] (One-time warning): Not using XLA:CPU for cluster because envvar TF_XLA_FLAGS=--tf_xla_cpu_global_jit was not set. If you want XLA:CPU, either set that envvar, or use experimental_jit_scope to enable XLA:CPU. To confirm that XLA is active, pass --vmodule=xla_compilation_cache=1 (as a proper command-line flag, not via TF_XLA_FLAGS) or set the envvar XLA_FLAGS=--xla_hlo_profile.
Traceback (most recent call last):
File "dbuilding.py", line 358, in
train(model)
File "dbuilding.py", line 175, in train
dataset_train.load_dbuilding(args.dataset, "train")
File "dbuilding.py", line 109, in load_dbuilding
annotations = [a for a in annotations if a['regions']]
File "dbuilding.py", line 109, in
annotations = [a for a in annotations if a['regions']]
KeyError: 'regions'
Would really appreciate it if you could provide a detailed explanation of the changes that you made!
hi @zulfiqarbolt,
first, u need to check a program by which you annotate, because in VIA 2.0, regions was changed from a dict to a list!!
second, try to modify your JSON file, ex. delete this part:

im not sure, but my file and file @waleedka has this format:
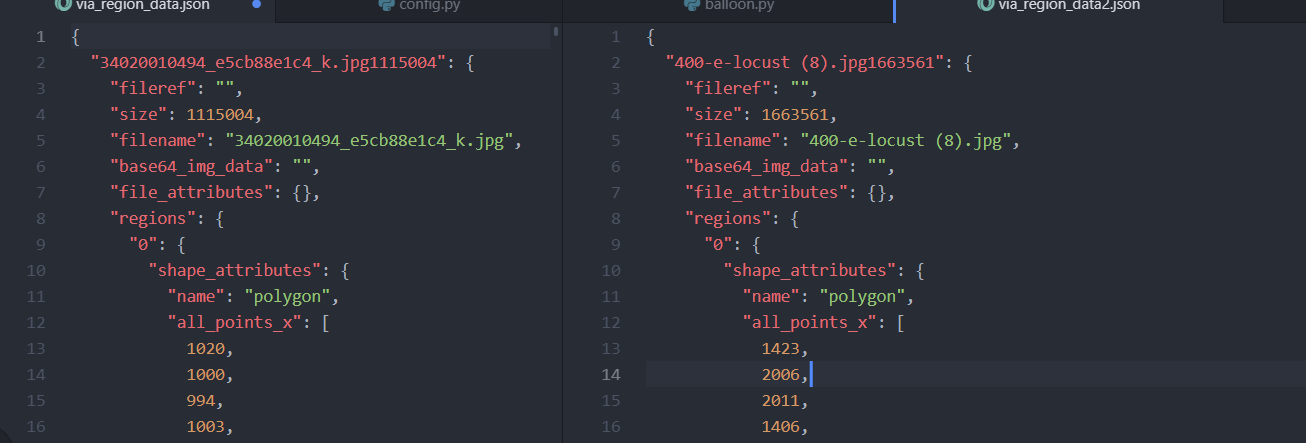
hi @zulfiqarbolt,
first, u need to check a program by which you annotate, because in VIA 2.0, regions was changed from a dict to a list!!
second, try to modify your JSON file, ex. delete this part:
im not sure, but my file and file @waleedka has this format:
Thanks @flit86. I shifted to the VIA annotation tool version 1 for annotating the images in JSON format and now the model works perfectly fine!
However, I observed in the balloon.py file, that the load_Balloon( ) function actually converts the JSON dictionary file into a list eventually. If it eventually converts it into a list, then why doesn't the JSON file annotated from VIA version 2.0 (which changes the regions from a dictionary to a list) work with the model?
Although the model is working fine, this particular segment in the code perplexes me a bit.
hi @zulfiqarbolt
i don't clarity understand where (the part) you have a problem with code.
can you send the photo or scrin???
Hi,
I am also having this issue, as I annotated with via-2.0.8. Is there any way to modify the balloon.py code so that a via-2.x formatted JSON file can be read in? I don't know enough about JSON parsing sadly.
Like @zulfiqarbolt I have also noticed that below the problem code (line 117) at lines 120-128 there is code for via-1.x and via-2.x formats. It seems this needs to also be done for line 117.
EDIT:
After playing around a bit, I believe this code will remove unlabelled images when the JSON has been formatted with via-2.x. Replace line 117 with the following:
via_1_check = annotations.get('regions')
via_2_check = annotations.get('_via_img_metadata')
# JSON is formatted with VIA-1.x
if via_1_check:
annotations = list(annotations.values())
# JSON is formatted with VIA-2.x
elif via_2_check:
annotations = list(annotations['_via_img_metadata'].values())
# Unknown JSON formatting
else:
raise ValueError('The JSON provided is not in a recognised via-1.x or via-2.x format.')
annotations = [a for a in annotations if a['regions']]
Note this has not been tested on a VIA-1.x formatted file, this is just using the code from the balloon.py file as a reference for how they should be handled. Happy to make this a pull request if people want.
@flit86 this may help you?
Hi,
@Trotts , thanks for your answer.
My problem was in the formatting and errors in the description of the annotation. Now I corrected it. Thank you very much for the advice!!!!! @Trotts
@Trotts i think you should also remove 113 to sort this out.
@Trotts Thank you! You should def make this a pull request!
I was battling the Key Error for 4 days and finally found that the solution was simple.. The key error happens because of incorrect Annotation.
Suppose you have 3 classes (Pizza, Burger and Sandwich). in Annotation , where Pizza = 1, Burger =2 , Sandwich = 3
lets say your region attributes name is "Food .
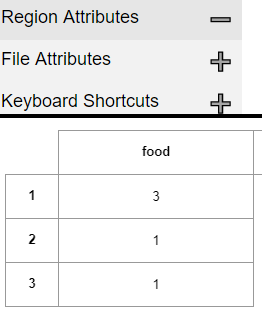
When you load an image with say 2 pizzas and 1 burget, the Annotation should be of this format
The mistake I initially made was :
Food
1 Pizza
2 Pizza
3 Burger
the correct Answer should be
Food
1 1
2 1
3 2
Another important point, if you have loaded an image that did not have annotation and you generated the Json file, it'll show as "regions" : {}. So ensure that you remove the entire line for image where ther is no annotation (by searching for "regions" : {}. and deleting the lines..
I had more than 10 pics without annotation and this was the reason why i had the error..
Hope my answer was clear :)
Hi,
I am also having this issue, as I annotated with via-2.0.8. Is there any way to modify the balloon.py code so that a via-2.x formatted JSON file can be read in? I don't know enough about JSON parsing sadly.
Like @zulfiqarbolt I have also noticed that below the problem code (line 117) at lines 120-128 there is code for via-1.x and via-2.x formats. It seems this needs to also be done for line 117.
EDIT:
After playing around a bit, I believe this code will remove unlabelled images when the JSON has been formatted with via-2.x. Replace line 117 with the following:
via_1_check = annotations.get('regions') via_2_check = annotations.get('_via_img_metadata') # JSON is formatted with VIA-1.x if via_1_check: annotations = list(annotations.values()) # JSON is formatted with VIA-2.x elif via_2_check: annotations = list(annotations['_via_img_metadata'].values()) # Unknown JSON formatting else: raise ValueError('The JSON provided is not in a recognised via-1.x or via-2.x format.') annotations = [a for a in annotations if a['regions']]Note this has not been tested on a VIA-1.x formatted file, this is just using the code from the balloon.py file as a reference for how they should be handled. Happy to make this a pull request if people want.
@flit86 this may help you?
got the error line 117, in load_nail
via_1_check = annotations.get('regions')
AttributeError: 'list' object has no attribute 'get'
Hi, I have similar problem. KeyError: 'region_attributes'
But it happend on;y when I have more then one label. First I taged the images with one label only and it worked, but then when I added another label, it didn't worked.
Most helpful comment
Hi,
I am also having this issue, as I annotated with via-2.0.8. Is there any way to modify the balloon.py code so that a via-2.x formatted JSON file can be read in? I don't know enough about JSON parsing sadly.
Like @zulfiqarbolt I have also noticed that below the problem code (line 117) at lines 120-128 there is code for via-1.x and via-2.x formats. It seems this needs to also be done for line 117.
EDIT:
After playing around a bit, I believe this code will remove unlabelled images when the JSON has been formatted with via-2.x. Replace line 117 with the following:
Note this has not been tested on a VIA-1.x formatted file, this is just using the code from the balloon.py file as a reference for how they should be handled. Happy to make this a pull request if people want.
@flit86 this may help you?