Mask_rcnn: Test on own dataset / No instances to display
Hi everyone,
I'm currently using this repo for an object detection problem with a lot of classes.
I trained on my own dataset (37 classes) and I used the splash balloon effect which works but I would like bounding boxes and scores confidence to see which object is detected.
That is why I have tried to test with the "demo.ipynb" code, to load my weights and to test the detection on my own dataset.
There is no error but I got this message : "NO INSTANCE TO DISPLAY"
I am a beginner in the computer vision field but I think I'm close to have results.
Here is my curves of my training :
But I think there is a problem when the weights are loaded.
In the original "demo.ipynb", I have an error with these lines which load the weights:
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR, config=config)
model.load_weights(COCO_MODEL_PATH, by_name=True)
My error disappears when I replace with this line:
model.load_weights(COCO_MODEL_PATH, by_name=True, exclude=[ "mrcnn_class_logits", "mrcnn_bbox_fc", "mrcnn_bbox", "mrcnn_mask"])
But "NO INSTANCE TO DISPLAY"
Maybe somebody can help me with this ?
Regards,
Antoine
All 18 comments
Better check the nucleus demo. There, you will see many ways to use the methods from visualize.py
Hey! I have been experiencing this NO INSTANCE TO DISPLAY, but actually I notice that I had the DETECTION_MIN_CONFIDENCE parameter (in config.py) that was equal to 0.7, hence all the prediction below this threshold was not displayed. Puting this to 0.5 allowed to display more instances for example. Dont know if this helps.
@fastlater Thanks for your answer ! Are you talking about this repo:
https://github.com/wanwanbeen/maskrcnn_nuclei ?
@camillemontalcini Thanks for having shared your feedbacks ! I tried to change the value to 0.5 and it displays instances but the detections are not good at all :'(
(I forgot to mention that I have used VGG Image Annotator tool for my groundtruth data, and I don't know if the .json output is different in structure from the COCO .json annotations...)
@camillemontalcini https://github.com/matterport/Mask_RCNN/tree/master/samples/nucleus
You said: there is a problem when the weights are loaded. What's exact problem?
@AliceDinh Here is my jupyter notebook code (the demo.ipynb) modified for my needs. I have 37 classes and I load the weights "mask_rcnn_f100_0150.h5". Firstly, I test the detection on images of the training set "imagesf100".
Then, here is my config:
Here I have my 37 classes but I don't know if it is necessary like COCO, and I load my weights which are not trained on MS-COCO dataset.
Here is my detection and "No instances to display" even if I have a DETECTION_MIN_CONFIDENCE of 0.7 or 0.5.
Am I wrong somewhere ?
@nayzen I reckoned that there is nothing wrong with the notebook. However, probably your annotation, what took did you use to create the annotation (VIA?) the format of annotation? and your code F100.py, you should have look these again.
I used the VIA tool to do my annotations. The format is .json and this is the organisation:
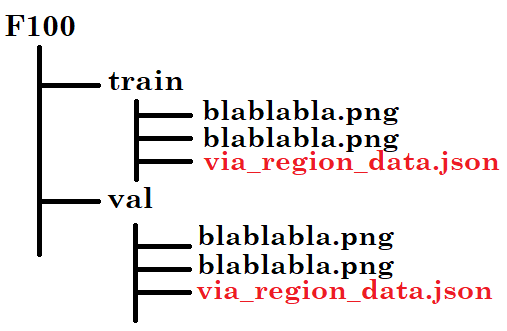
For the code "F100.py", I listened to your advices by email.
I think you have to try with lower confidence to see any instance, you used 0.5, let's try lower. And try with the images in the training set. That's all I can suggest. Sorry can't help much
Thanks anyway @AliceDinh , for your help :-)
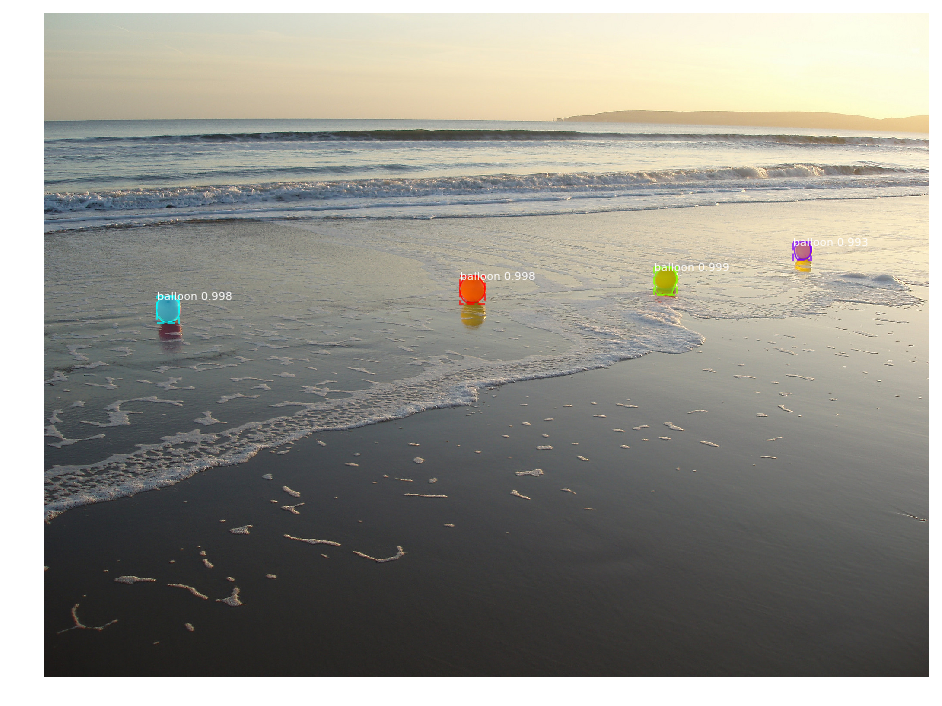
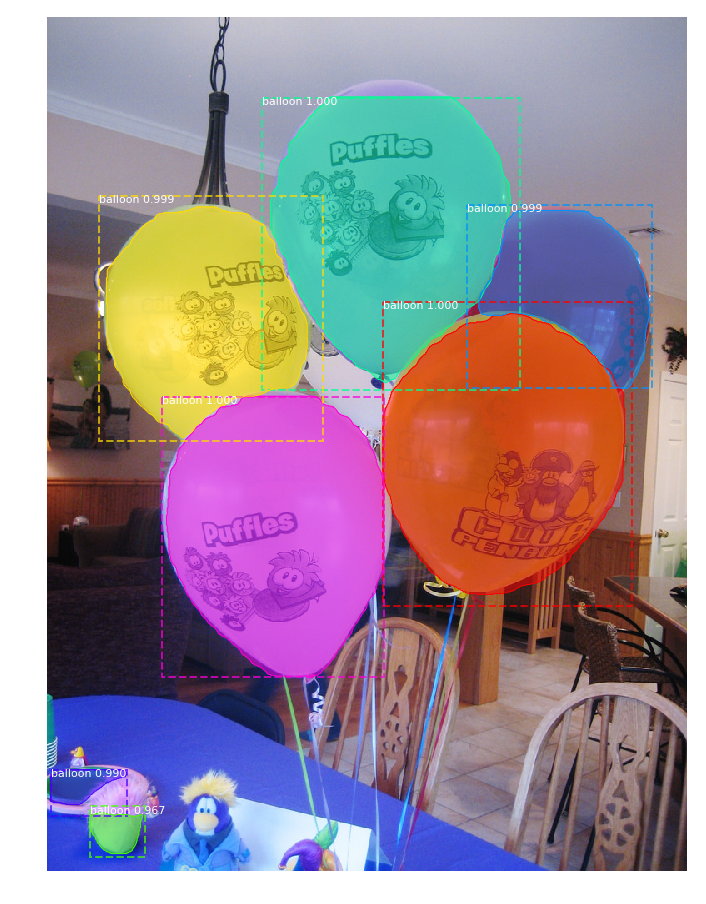
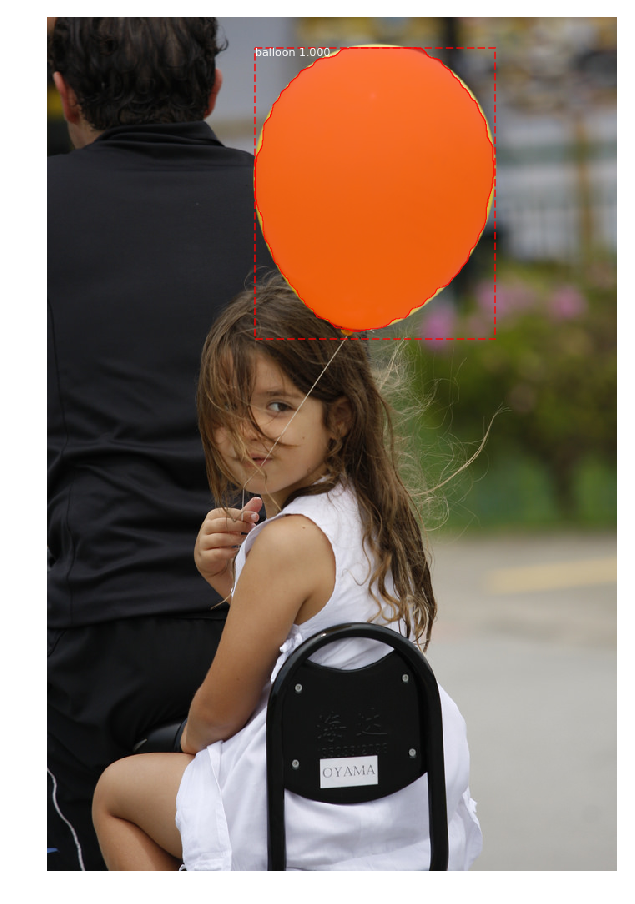
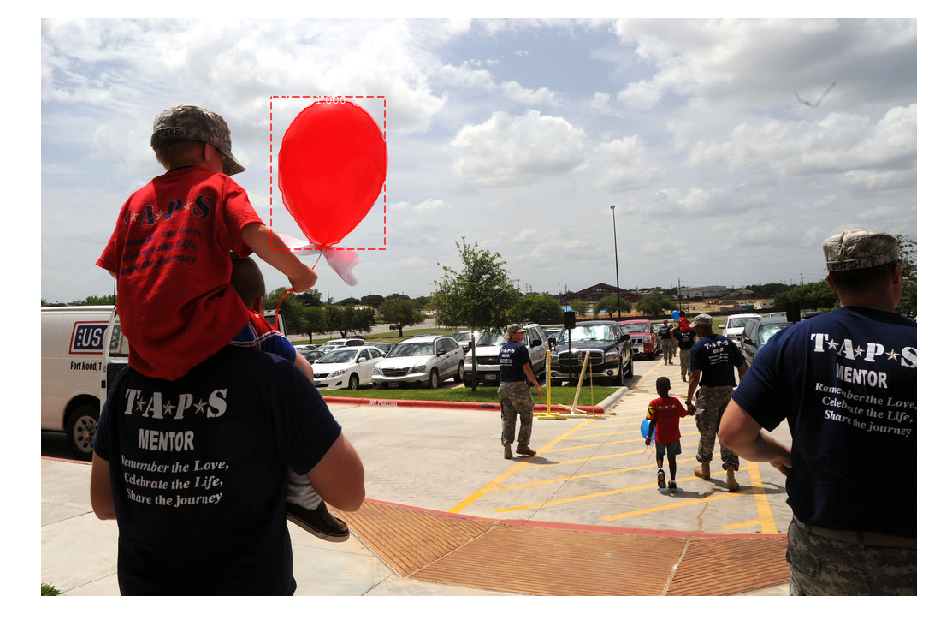
When I test the "demo.ipynb" with balloon weights, I see instances but the detection are not good at all:
I don't understand why. All I can say is that I must exclude some layers otherwise it doesn't work :(

Here is the error : "_ValueError: Dimension 1 in both shapes must be equal, but are 8 and 324. Shapes are [1024,8] and [1024,324]. for 'Assign_1366' (op: 'Assign') with input shapes: [1024,8], [1024,324]._"
If someone could explain to me.. I would be grateful.
I fixed my problem deleting the .h5 file at the root, and copying it again.




@nayzen
Hello!
I met the same question. Only can I get a good result when I put
exclude=[ "mrcnn_class_logits", "mrcnn_bbox_fc", "mrcnn_bbox", "mrcnn_mask"]
in the code.
u said: I fixed my problem deleting the .h5 file at the root, and copying it again.
do u mean that u found a way to get a good result without code like
exclude=[ "mrcnn_class_logits", "mrcnn_bbox_fc", "mrcnn_bbox", "mrcnn_mask"]
thx for reading
@greebear
Hello ! yes it could seem weird but it didn't work without the "exclude" part.
I deleted the .h5 where they were loaded, then I copied the .h5 again at the same place.. And it works with the original "demo code", that is to say : without the "exclude layers..." :)
@greebear
PS: And don't forget to include the background for the first class name ! ('BG', 'balloon', '...',...)
I've solved this problem by modifying utils.py line 866 from
shift = np.array([0, 0, 1, 1])
to
shift = np.array([0, 0, 1., 1.])
It's simply a data type error which will cause float number becomes zero, leading to divide by zero error
how can i get the loss and val loss curve?
@lapetite123 Hello,
You can visualize during training (or after) the loss and val curve opening a prompt command and hit:
tensorboard -logdir=C:.....\Mask_RCNN-master\logs\folder-containing-.h5-file
I've solved this problem by modifying utils.py line 866 from
shift = np.array([0, 0, 1, 1])
to
shift = np.array([0, 0, 1., 1.])
It's simply a data type error which will cause float number becomes zero, leading to divide by zero error
Thank you very much !
Most helpful comment
I've solved this problem by modifying utils.py line 866 from
shift = np.array([0, 0, 1, 1])to
shift = np.array([0, 0, 1., 1.])It's simply a data type error which will cause float number becomes zero, leading to divide by zero error