Cudf: [BUG] .isnull() not counting nan w/o None present
Describe the bug
cudf.Series().isnull() not counting nan values the same as pandas, unless None is present, or only nan values are present.
Steps/Code to reproduce bug
This code outputs (0, 2)
import numpy
import cupy
import cudf
df = cudf.DataFrame()
df['a'] = [numpy.nan, 0, 'null', cupy.nan]
cudf_count = sum(df.a.isnull())
df = cudf.DataFrame().to_pandas()
df['a'] = [numpy.nan, 0, 'null', cupy.nan]
pd_count = sum(df.a.isnull())
cudf_count, pd_count
This code (substitute None for 0) outputs (3, 3)
import numpy
import cupy
import cudf
df = cudf.DataFrame()
df['a'] = [numpy.nan, None, 'null', cupy.nan]
cudf_count = sum(df.a.isnull())
df = cudf.DataFrame().to_pandas()
df['a'] = [numpy.nan, None, 'null', cupy.nan]
pd_count = sum(df.a.isnull())
cudf_count, pd_count
This code (substitute 0 for 'null', still with None) outputs (3, 3) as well
import numpy
import cupy
import cudf
df = cudf.DataFrame()
df['a'] = [numpy.nan, None, 0, cupy.nan]
cudf_count = sum(df.a.isnull())
df = cudf.DataFrame().to_pandas()
df['a'] = [numpy.nan, None, 0, cupy.nan]
pd_count = sum(df.a.isnull())
cudf_count, pd_count
This code (nans only) outputs (2, 2)
import numpy
import cupy
import cudf
df = cudf.DataFrame()
df['a'] = [numpy.nan, cupy.nan]
cudf_count = sum(df.a.isnull())
df = cudf.DataFrame().to_pandas()
df['a'] = [numpy.nan, cupy.nan]
pd_count = sum(df.a.isnull())
cudf_count, pd_count
Expected behavior
The first code to output: (2, 2)
Environment overview (please complete the following information)
- Environment location: app.blazingsql.com (RAPIDS Stable)
- Method of cuDF install: conda
Environment details
Click here to see environment details
**git***
commit ad15332f33020f213b81134861f91a2677873765 (HEAD -> feature/taxi.ipynb, origin/feature/taxi.ipynb)
Author: Winston A Robson <[email protected]>
Date: Tue Jun 30 00:08:22 2020 +0000
shrunk taxi urls list
**git submodules***
***OS Information***
DISTRIB_ID=Ubuntu
DISTRIB_RELEASE=18.04
DISTRIB_CODENAME=bionic
DISTRIB_DESCRIPTION="Ubuntu 18.04.3 LTS"
NAME="Ubuntu"
VERSION="18.04.3 LTS (Bionic Beaver)"
ID=ubuntu
ID_LIKE=debian
PRETTY_NAME="Ubuntu 18.04.3 LTS"
VERSION_ID="18.04"
HOME_URL="https://www.ubuntu.com/"
SUPPORT_URL="https://help.ubuntu.com/"
BUG_REPORT_URL="https://bugs.launchpad.net/ubuntu/"
PRIVACY_POLICY_URL="https://www.ubuntu.com/legal/terms-and-policies/privacy-policy"
VERSION_CODENAME=bionic
UBUNTU_CODENAME=bionic
Linux ip-172-31-12-139 4.15.0-1065-aws #69-Ubuntu SMP Thu Mar 26 02:17:29 UTC 2020 x86_64 x86_64 x86_64 GNU/Linux
***GPU Information***
Failed to initialize NVML: Driver/library version mismatch
***CPU***
Architecture: x86_64
CPU op-mode(s): 32-bit, 64-bit
Byte Order: Little Endian
CPU(s): 4
On-line CPU(s) list: 0-3
Thread(s) per core: 2
Core(s) per socket: 2
Socket(s): 1
NUMA node(s): 1
Vendor ID: GenuineIntel
CPU family: 6
Model: 85
Model name: Intel(R) Xeon(R) Platinum 8259CL CPU @ 2.50GHz
Stepping: 7
CPU MHz: 3192.167
BogoMIPS: 4999.99
Hypervisor vendor: KVM
Virtualization type: full
L1d cache: 32K
L1i cache: 32K
L2 cache: 1024K
L3 cache: 36608K
NUMA node0 CPU(s): 0-3
Flags: fpu vme de pse tsc msr pae mce cx8 apic sep mtrr pge mca cmov pat pse36 clflush mmx fxsr sse sse2 ss ht syscall nx pdpe1gb rdtscp lm constant_tsc rep_good nopl xtopology nonstop_tsc cpuid aperfmperf tsc_known_freq pni pclmulqdq ssse3 fma cx16 pcid sse4_1 sse4_2 x2apic movbe popcnt tsc_deadline_timer aes xsave avx f16c rdrand hypervisor lahf_lm abm 3dnowprefetch invpcid_single pti fsgsbase tsc_adjust bmi1 avx2 smep bmi2 erms invpcid mpx avx512f avx512dq rdseed adx smap clflushopt clwb avx512cd avx512bw avx512vl xsaveopt xsavec xgetbv1 xsaves ida arat pku ospke avx512_vnni
***CMake***
***g++***
/usr/bin/g++
g++ (Ubuntu 7.5.0-3ubuntu1~18.04) 7.5.0
Copyright (C) 2017 Free Software Foundation, Inc.
This is free software; see the source for copying conditions. There is NO
warranty; not even for MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
***nvcc***
***Python***
/usr/bin/python
Python 2.7.17
***Environment Variables***
PATH : /usr/local/sbin:/usr/local/bin:/usr/sbin:/usr/bin:/sbin:/bin
LD_LIBRARY_PATH :
NUMBAPRO_NVVM :
NUMBAPRO_LIBDEVICE :
CONDA_PREFIX : /opt/conda-environments/rapids-stable
PYTHON_PATH :
conda not found
***pip packages***
/usr/bin/pip
DEPRECATION: The default format will switch to columns in the future. You can use --format=(legacy|columns) (or define a format=(legacy|columns) in your pip.conf under the [list] section) to disable this warning.
asn1crypto (0.24.0)
backports.functools-lru-cache (1.6.1)
boto3 (1.12.18)
botocore (1.15.18)
certifi (2019.11.28)
chardet (3.0.4)
click (7.1.2)
cryptography (2.1.4)
cycler (0.10.0)
docutils (0.15.2)
enum34 (1.1.6)
futures (3.3.0)
idna (2.9)
ipaddress (1.0.17)
jmespath (0.9.5)
joblib (0.14.1)
kaggle (1.5.6)
keyring (10.6.0)
keyrings.alt (3.0)
mysqlclient (1.3.10)
numpy (1.16.6)
nvidia-ml-py (375.53.1)
pandas (0.24.2)
pip (9.0.1)
pycrypto (2.6.1)
pygobject (3.26.1)
python-dateutil (2.8.1)
python-slugify (4.0.0)
pytz (2020.1)
pyxdg (0.25)
requests (2.23.0)
s3fs (0.2.2)
s3transfer (0.3.3)
scipy (1.2.3)
SecretStorage (2.3.1)
setuptools (39.0.1)
six (1.14.0)
text-unidecode (1.3)
tqdm (4.43.0)
urllib3 (1.25.8)
wheel (0.30.0)
Additional context
^^ conda not found ^^
conda list cu
```
packages in environment at /opt/conda-environments/rapids-stable:
#
Name Version Build Channel
cudatoolkit 10.0.130 0 nvidia
cudf 0.14.0 py37_0 rapidsai
cudnn 7.6.0 cuda10.0_0 nvidia
cugraph 0.14.0 py37_0 rapidsai
cuml 0.14.0 cuda10.0_py37_0 rapidsai
cupy 7.5.0 py37h658377b_0 conda-forge
curl 7.68.0 hf8cf82a_0 conda-forge
cuspatial 0.14.0 py37_0 rapidsai
cuxfilter 0.14.0 py37_0 rapidsai
dask-cuda 0.14.0 py37_0 rapidsai
dask-cudf 0.14.0 py37_0 rapidsai
docutils 0.15.2 py37_0 conda-forge
icu 64.2 he1b5a44_1 conda-forge
libcudf 0.14.0 cuda10.0_0 rapidsai
libcugraph 0.14.0 cuda10.0_0 rapidsai
libcuml 0.14.0 cuda10.0_0 rapidsai
libcumlprims 0.14.1 cuda10.0_0 nvidia
libcurl 7.68.0 hda55be3_0 conda-forge
libcuspatial 0.14.0 cuda10.0_0 rapidsai
ncurses 6.1 hf484d3e_1002 conda-forge
This code breaks:
```python
import numpy
import cupy
import cudf
df = cudf.DataFrame()
df['a'] = [numpy.nan, 0, 'null', cupy.nan, None]
cudf_count = sum(df.a.isnull())
May relate to: https://github.com/rapidsai/cudf/issues/5555
All 7 comments
I think this all comes down to what we translate [numpy.nan, 0, 'null', cupy.nan] to.
@gumdropsteve in the first example what is the dtype of df['a'] and what does printing it show?
@kkraus14 this is the output of the first example
For cudf
>>> df['a']
0 null
1 null
2 0
3 null
Name: a, dtype: int64
for pandas
>>> df['a']
0 NaN
1 0
2 null
3 NaN
Name: a, dtype: object
@taureandyernv can you double check you used [numpy.nan, 0, 'null', cupy.nan] as input for cuDF as well?
The third element being 0 instead of the second element doesn't make sense to me
I'm getting object as dtype for both cuDF & pandas, @kkraus14.
Here's a screenshot with cuDF & pandas for the first example;
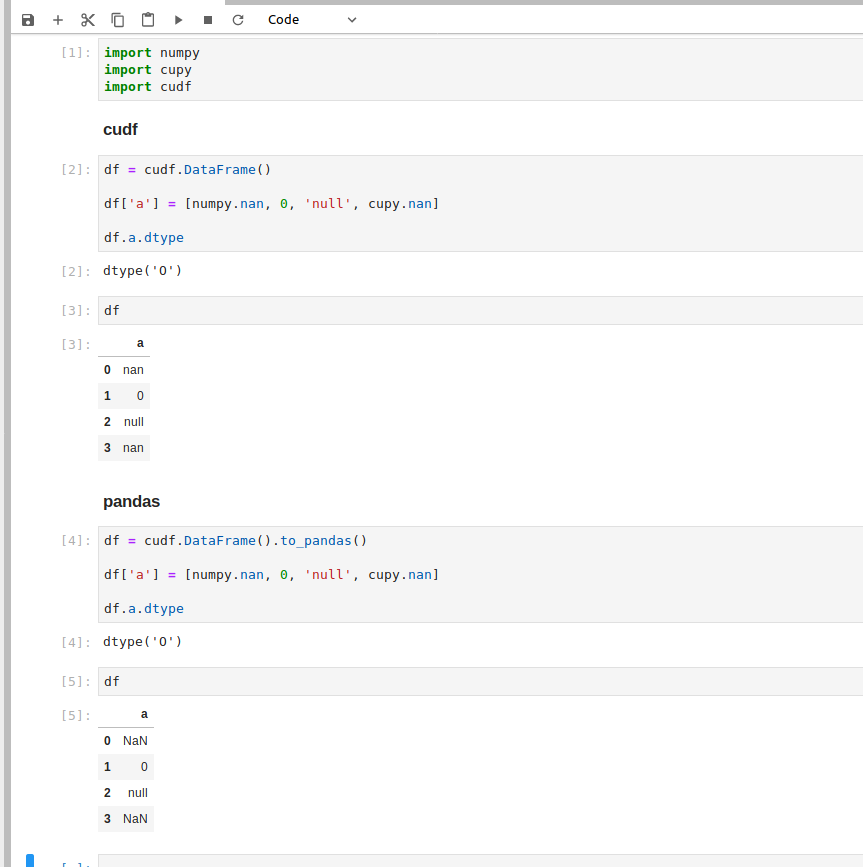
@gumdropsteve thanks for posting that, this makes sense now. What's happening is that cuDF doesn't have a true object type similar to Pandas since we can't handle arbitrary Python objects on the GPU, so it's converting everything to strings. This means cuDF is effectively getting ["nan", "0", "null", "nan"] as its data, which means there's no NA values.
On the other hand, Pandas can have arbitrary Python objects, so it gets [NaN, 0, "null", NaN] which gives it two NA values.
The expected behavior for cuDF in this case feels ambiguous to me, where maybe we should just throw instead of casting everything to strings.
@gumdropsteve thanks for posting that, this makes sense now. What's happening is that cuDF doesn't have a true
objecttype similar to Pandas since we can't handle arbitrary Python objects on the GPU, so it's converting everything to strings. This means cuDF is effectively getting["nan", "0", "null", "nan"]as its data, which means there's no NA values.On the other hand, Pandas can have arbitrary Python objects, so it gets
[NaN, 0, "null", NaN]which gives it two NA values.The expected behavior for cuDF in this case feels ambiguous to me, where maybe we should just throw instead of casting everything to strings.
In that case, should we consider [1, "1.0"] -> ["1", "1.0"] is also wrong ?
I would argue that the behavior is incorrect as it differs from Pandas object expected behavior, so we should probably throw instead of casting everything to string.
Most helpful comment
I would argue that the behavior is incorrect as it differs from Pandas
objectexpected behavior, so we should probably throw instead of casting everything to string.