Cudf: [FEA] Efficient concatenate for many partitions
Is your feature request related to a problem? Please describe.
In a large distributed system like Spark, the data often needs to be partitioned into many parts to allow a task's data to fit in GPU memory. Each downstream task fetches its corresponding input partition from each upstream task's output partitions. Usually these input partitions are concatenated together to form a large input table for the task.
If there are many partitions and/or many columns, the current concatenation approach is slow on the CPU. For example, if there are 100 input partitions and each partition contains 10 columns then the concatenation operation performs 1000 calls to cudaMemcpy to concatenate the data vectors in the final table. The overhead of all these CUDA calls on the CPU can be significant.
Describe the solution you'd like
Ideally there would be an API that takes a vector of tables and emits a new table. The implementation would leverage a kernel that could stitch together N column fragments into a single column, much like gdf_mask_concat uses a single kernel to concatenate N mask fragments.
One concern with a table-based API is if table building itself is relatively expensive. For 100 input partitions, we would need to construct 100 tables in order to call the table concatenate API. If constructing a table calls a CUDA API for every column in the table then we're back to the original problem of too many CUDA API calls to handle the input partitions.
Describe alternatives you've considered
If table construction is too costly when there are many input partitions with many columns then a column-based API would be more useful. We currently bulk-transfer each input partition's data to the device in one cudaMemcpy call and can construct the column metadata relatively cheaply.
All 30 comments
We could use this within cuDF as well so that we can concatenate many dataframes / columns at once instead of pairwise.
gdf_column_concat concatenates multiple columns. The only thing is that it has never been updated to operate on table inputs.
@jlowe Constructing tables should be relatively cheap -- the table construction should not invoke CUDA functions -- it's a host-side vector of column pointers.
gdf_column_concat concatenates multiple columns. The only thing is that it has never been updated to operate on table inputs.
gdf_column_concat does concatenate multiple columns per call, and we are already calling it to do that. However the implementation has a CPU-side for loop which calls cudaMemcpy for each partition:
for (int i = 0; i < num_columns; ++i) {
std::size_t bytes = column_byte_width * columns_to_concat[i]->size;
CUDA_TRY( cudaMemcpy(target, columns_to_concat[i]->data, bytes, cudaMemcpyDeviceToDevice) );
target += bytes;
output_column->null_count += columns_to_concat[i]->null_count;
}
That's what leads to the (partitions * columns) CUDA call count issue. Therefore this ask is a bit more than a table API veneer on the existing multi-column concatenate.
the table construction should not invoke CUDA functions
Totally agree. I only brought it up because at one point the table constructor was building a list of device-side vectors, e.g.: column data pointers, column dtypes, column byte widths. It looks like the current table constructor no longer does this, so as long as that holds we're fine.
I only brought it up because at one point the table constructor was building a list of device-side vectors, e.g.: column data pointers, column dtypes, column byte widths. It looks like the current table constructor no longer does this, so as long as that holds we're fine.
You're thinking of device_table, which is not meant to be a publicly used data structure.
I actually was thinking of the original gdf_table which was public but no longer exists in that form.
Update on this issue.
The goal is to optimize the concatenate implementation to avoid needing to do a kernel call/memcpy per partition to be concatenated and instead concatenate n partitions in a single kernel.
See https://github.com/rapidsai/cudf/blob/branch-0.13/cpp/src/column/column.cu#L258
@trevorsm7 the first task here should be to make some benchmarks for the existing implementation.
It looks like the existing concatenate for table should be fine as-is, as it uses the above concatenate for column internally.
The strings concatenate that @davidwendt worked on is already doing a vectorized copy using thrust::for_each_n and a device functor concatenate_fn (and a thrust::inclusive_scan to compute the offsets). Does this look efficient enough to tackle the spirit of this issue? If so, this could be a good starting point for the fixed-width implementation.
https://github.com/rapidsai/cudf/blob/a5a258d73bb271e1bd905b271c13e25e17fe9173/cpp/src/strings/copying/concatenate.cu#L147-L150
https://github.com/rapidsai/cudf/blob/a5a258d73bb271e1bd905b271c13e25e17fe9173/cpp/src/strings/copying/concatenate.cu#L52-L85
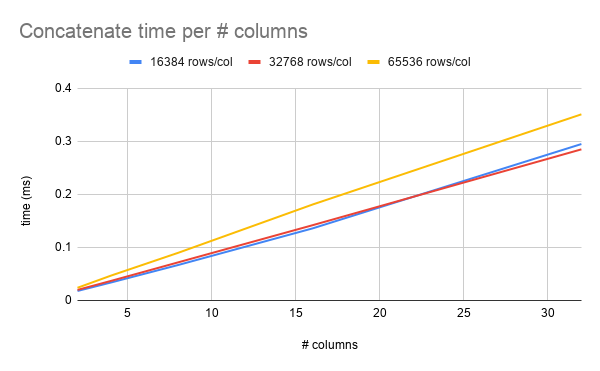
I just picked some arbitrary ranges to start with. Here's a graph of concatenate time per number of columns, where each column is the same number of rows. Here it looks pretty linear, but if I push up the number of columns and decrease the number of rows, maybe the overhead from cudaMemcpy may start to show.
The strings concatenate that @davidwendt worked on is already doing a vectorized copy using
thrust::for_each_nand a device functorconcatenate_fn(and athrust::inclusive_scanto compute the offsets). Does this look efficient enough to tackle the spirit of this issue? If so, this could be a good starting point for the fixed-width implementation.
That implementation is going to use 1 thread per partition. That's going to be slow. In fact, I believe that caused a performance regression: https://github.com/rapidsai/cudf/pull/4215
That is correct. The thrust::for_each approach I implemented in strings::detail::concatenate proved to be very bad for performance and is to be reverted here in #4215
I suspect that this would scale better for a large number of columns with very small rows but it certainly performs poorly for a small number of columns with a large number of rows.
So there's two possible algorithms we can try here.
For n partitions, with the size of partition i indicated by sizes[i]:
- Compute
offsetsvia an exclusive scan ofsizes - Start a
partition_mapvector (size == result.size()) by scatteringito positionoffset[i] - Compute a max-scan over
partition_mapto fill in the holes.partition_map[i]will now tell you which partitionjthatresult[i]comes from - You can then do a
thrust::copyusing atransform_iteratorfor the input that does:
auto get_element = [offsets, partition_map] __device__ (auto i){
auto partition_num = partition_map[i];
return arrays[partition_num][i - offsets[partition_num]]
}
OR
- Compute
offsetsvia an exclusive scan ofsizes thrust::copywith a transform iterator that does:
auto get_element = [offsets] __device__ (auto i){
auto partition_num = thrust::lower_bound(thrust::seq, offsets, offsets + num_partitions, i);
return arrays[*partition_num][i - offsets[*partition_num]]
}
The only difference between the two is the first algorithm materializes the mapping of a thread i to the input partition j that it will read from. Alternatively, the second algorithm computes this mapping on the fly with a binary search.
My bias is toward the second algorithm as it requires less intermediate materializations which reduces memory pressure. offsets is generally going to be "small" (O(10-100)), so it will likely be cached and the binary search should be pretty fast.
As @kaatish pointed out to me, the concatenate_bitmask_kernel is pretty much already doing the second algorithm I proposed above.
As such, it may actually be better to choose the first algorithm as we can reuse the partition_map between the data and the bitmask kernel. What's more, we can reuse the partition_map for each column in a table if we're concatenating multiple columns.
So now I'm leaning towards the first algorithm but it could go either way.
@trevorsm7 It should be pretty easy to implement both algorithms. So let's try both and benchmark to compare.
@jrhemstad With the second approach, wouldn't I need upper_bound(...) - 1 to get the correct partition_num? I'm thinking that lower_bound would incorrectly point me to the next partition, except for the first element of each partition which would be correct.
Also, should this be return arrays[partition_num - offsets][i - *partition_num] since dereferencing partition_num should result in the offset?
auto get_element = [offsets] __device__ (auto i){
auto partition_num = thrust::lower_bound(thrust::seq, offsets, offsets + num_partitions, i);
return arrays[*partition_num][i - offsets[*partition_num]]
}
With the changes I suggested, assuming offsets is the inclusive_scan of sizes:
auto get_element = [offsets] __device__ (auto i){
auto partition_num = thrust::upper_bound(thrust::seq, offsets, offsets + num_partitions, i) - 1;
return arrays[partition_num - offsets][i - *partition_num]
}
Ah, concatenate_bitmask_kernel is doing the same using upper_bound() - 1, so I'll go with that.
Yeah, your changes look correct to me.
That's what you get when coding with pen/paper :)
With the changes I suggested, assuming offsets is the inclusive_scan of sizes:
offsets should be the exclusive scan, no?
Edit: I think the choice of lower/upper bound depends on if you're using an inclusive/exclusive scan.
Hm, I think if you useexclusive_scan then you'd have trouble with the [i - *partition_num] part because you expect the 0th offset to be 0, not the size of the first partition.
Hm, I think if you use
exclusive_scanthen you'd have trouble with the[i - *partition_num]part because you expect the 0th offset to be 0, not the size of the first partition.
offset[0] == 0 implies an exclusive scan.
[3, 5, 2, 7, 9]
inclusive scan: [3, 8, 10, 17, 26]
exclusive scan: [0, 3, 8, 10, 17]
Oh duh! You're right, I was thinking of inclusive_scan. Yes, exclusive_scan is what I want
I am a little surprised to see that concatenate_bitmask is using an inclusive scan. That doesn't seem right?
Oh duh! You're right, I was thinking of
inclusive_scan. Yes,exclusive_scanis what I want
I always struggled to remember the difference until someone told me what the "inclusive" and "exclusive" part actually means.
"Exclusive" means that scan[i] _excludes_ input[i]
"Inclusive" means that scan[i] _includes_ input[i]
Maybe that was obvious to everyone else, but someone had to tell me before it finally clicked.
Aha, concatenate_bitmask is using inclusive scan, but the first element of the view_offsets vector is reserved as zero, so the final result looks like an exclusive scan with an extra element at the end for the total size.
Aha,
concatenate_bitmaskis using inclusive scan, but the first element of theview_offsetsvector is reserved as zero, so the final result looks like an exclusive scan with an extra element at the end for the total size.
Ah, okay, that makes more sense. That's probably necessary in your algorithm as well then. Ideally we just want to share the same offset vector for the data/bitmask kernels.
@trevorsm7 since you're familiar with Cython porting now, you could probably tackle https://github.com/rapidsai/cudf/issues/3926 too. That is unless @mt-jones hasn't already started.
@jrhemstad I wrote up an implementation of the partition map based algorithm that I think should work, but I've run myself into the mud with thrust somewhere and am getting some eldritch errors that I can't decrypt. Mind taking a look? https://github.com/rapidsai/cudf/pull/4224#issuecomment-589925268
@jlowe I'd like to extend my benchmark to cover a range of parameters relevant to Spark. Specifically, for a single call to concatenate, how many tables are being concatenated, how many columns per table, and home many rows in each column? What's a typical range or a tricky case that we like to benchmark?
The number of columns varies widely depending on the query. For most queries I've seen, it's pretty reasonable due to Spark's SQL optimizer pruning columns. 2-10 columns for most cases. There are cases with a lot more. One example is an input repartition and transcode case (e.g.: CSV to Parquet transformation, preserving all data and balancing across files). All columns are needed, so it could easily be 20+ columns per table in those cases.
As for number of tables, we concatenate input partitions until we hit either a target size (to make it worth running that batch on the GPU) or all of them if the operation needs all tables (e.g.: sorting). The number of input partitions is user-configurable and also can vary widely, but I'd expect it to usually be between 2 to 200. 200 is Spark's default, so I'd expect to see that in practice quite a bit even if it may not be optimal for GPUs.
As for average number of rows per table, that varies the most of all the factors. It simply depend on what rows survive the upstream filtering, partitioning, etc which depends both on the query and the input values. In practice the number of rows will range from 1 to hundreds of thousands (sometimes millions). Unfortunately I don't have a good rule of thumb for average row count.