Cudf: Make a plan for sort_values/set_index
It would be nice to be able to use the set_index method to sort the dataframe by a particular column.
There are currently two implementations for this, one in dask.dataframe and one in dask-cudf which uses a batcher sorting net. While most dask-cudf code has been removed in favor of the dask.dataframe implementations this sorting code has remained, mostly because I don't understand it fully, and don't know if there was a reason for this particular implementation.
Why was this implementation chosen? Was this discussed somewhere? Alternatively @sklam, do you have any information here?
cc @kkraus14 @randerzander
All 80 comments
NVIDIA folks have asked if there is some way to integrate an MPI or NCCL enabled multi-gpu sort into Dask for improved efficiency. My initial reaction to this is that it's likely to be difficult to integrate smoothly in a way that respects other Dask management like resilience, spilling to disk, load balancing, and so on. Lets expand on this.
First, if we have a multi-node sorting algorithm, we can always treat it how we treat XGBoost. Dask gives up control to some other higher performance system, it does its thing, we claim control back. If anything fails during this stage then we just retry the whole thing. We give up on any kind of memory management or load balancing during this process and just hope that the external system can handle things well without blowing up.
Second question is if we just have a single-node multi-GPU system, maybe we can use that? This is also a bit tricky currently, but we might be able to make structural changes to Dask to make it less tricky. The cost-benefit analysis of those changes might make this undesirable though. Currently the approach most people seem to be using with Dask and GPUs is to have one Dask worker per GPU. Currently Dask workers don't have any knowledge of other Dask workers on the same node, so there isn't anything built up to handle local collective action. We would be doing something similar to what is done above where we would more or less stop Dask from doing its normal task-scheduling thing, hand-write a bunch of control flow, hope nothing breaks, run custom code, and then have Dask take back control when we're done.
Both are totally doable, but would require us to build something like dask-xgboost, and raise general concerns around memory management, resilience, diagnostics, load balancing, spilling to disk, and so forth. We lose a lot of Dask's management when we switch into this mode.
So maybe Dask should start thinking more about collective actions. This is coming up often enough that it probably deserves more attention. That's a large conversation though and probably requires dedicated time from someone somewhat deeply familiar with Dask scheduler internals.
I think that, short term, we should continue with the current approach of using efficient single-core/gpu sorts and shuffling techniques currently done in dask.dataframe. We should tune these to the extent that we can both by making the single-gpu sort algorithms faster, the book keeping operations faster, and the communication faster. If this isn't enough then we should investigate collective actions as discussed above, but that should be part of a larger effort than just sorting.
I agree with this approach. Thrust has a good single GPU sorting, and I think UCX should help tremendously. Any thoughts on book keeping, or was that more of a placeholder for if we need it?
There are two forms of book keeping that are relevant here:
- The slicing and concatenation of cudf dataframes. This is largely a test of memory management.
- The administrative tracking of tasks within the scheduler. This becomes more of an issue as we scale out to more nodes.
There are currently two implementations for this, one in dask.dataframe and one in dask-cudf which uses a batcher sorting net. While most dask-cudf code has been removed in favor of the dask.dataframe implementations this sorting code has remained, mostly because I don't understand it fully, and don't know if there was a reason for this particular implementation.
Why was this implementation chosen? Was this discussed somewhere? Alternatively @sklam, do you have any information here?
@sklam can you expand on the motivation behind your use of batcher sort net? Why was this decided on rather than the approach take in the mainline dask dataframe codebase?
@mrocklin, I was trying to avoid the following of the mainline sort/shuffle:
- Storing the data onto the disk
- Output is imbalanced. All equal keys in the same partition. If the key distribution is heavily skewed, GPU memory may run out.
I chose the sorting network because:
- It's very easy to implement.
- The sorting network has a fixed structure given the number of partitions.
- There are more parallel regions. (this one is according to my bad memory)
- The memory usage is predictable. The network can be seen as multiple-stages. Each stage has data-dependency on the previous stage only. Given 1GB of data, the algorithm will roughly need 2GB (1GB for the input and 1GB for the output at each stage).
- It will rebalance the partitions. It may produce fewer partitions and the partition size will not be more than the
max(input_partition_sizes)
OK, so these concerns seem similar to the concerns we had while designing the various shuffle algorithms for dask dataframe/bag. I'm not seeing anything here that is highly specific to GPU computation. (please correct me if I'm wrong).
My inclination then is to try to unify both CPU and GPU around a single implemenatation, probably starting with the one in dask.dataframe, but then maybe we would consider batcher-sorting-networks as an alternative for both, rather than just for dask-cudf, after doing some more extensive benchmarking.
Thanks for the information @sklam , I really appreciate the continuity here.
I recently took a look at Dask dataframe's task-based shuffle and improved docstrings here in order to help others dive in: https://github.com/dask/dask/pull/4674/files
I think that we need the following:
Something like the
pandas.util.hash_pandas_objectfunction, which hashes a pandas object row-by-row returning a Series of integer values. (@kkraus14 do we have this?)A
Series.searchsortedmethod, allowing us to figure out where each row should go. Used here:def set_partitions_pre(s, divisions): partitions = pd.Series(divisions).searchsorted(s, side='right') - 1 partitions[(s >= divisions[-1]).values] = len(divisions) - 2 return partitionsThis helps us to assign a partition number to every row, based on where the future-index value of that row sits relative to the divisions. Example: "Is the value in between the first and second divisions? Great, it goes in the first partition."
The
dask.dataframe.shuffle.shuffle_groupfunction, which splits a pandas dataframe apart into a dict of pandas dataframes based on the value of a particular column provided by the hash values above. We also need to modify this value in a particular way to achieved multi-staged shuffling. This logic is explained in the PR above.We can either rewrite this from scratch or, if cudf is supporting ufuncs we might be able to get away with just dispatching on the
pandas._libs.algos.groupsort_indexerand reuse all of the fancy logic here (which would be nice if cudf supports most ufuncs and out parameters)- Eventually we will also need quantile information. This is also evolving in Dask core at the moment, so I suggest that we wait on this for a bit.
Also, just to direct people, I think that the core function that we'll have to make work is rearrange_by_column_tasks. I think that all of set_index/merge/sort_values can be made to rely on functionality there.
@mrocklin @datametrician I have a vested interest in seeing this work succeed, and will begin by implementing the solution path you've outlined above. If we _need_ to get into the specifics of MPI/NCCL, we can cross that bridge later. I think a general solution which puts Dask first is going to help the ecosystem the most.
A minimal test would look something like the following:
import cudf, dask.dataframe as dd
# Make a dataframe, we'd like to divide the data by y
df = cudf.DataFrame({'x': [1, 2, 3, 4, 5, 6], 'y': [0, 4, 0, 4, 0, 4]})
# Split it up into a few partitions with Dask
ddf = dd.from_pandas(df, npartitions=3)
# Try to create a new dataframe with two partitions sorted on the y column, split by y 0->2 and 2->4
out = dd.shuffle.rearrange_by_divisions(ddf, column='y', divisions=[0, 2, 4], shuffle='tasks')
# compute in a single thread so it's easy to use %pdb and %debug
out.compute(scheduler='single-threaded')
I just tried this and ran into an minor problem of cudf.DataFrame.drop not supporting the axis= keyword. As a suggestion, these errors are easier to identify if you remove some of Dask's error reporting with the following diff
diff --git a/dask/dataframe/core.py b/dask/dataframe/core.py
index 6a08af9..894fba6 100644
--- a/dask/dataframe/core.py
+++ b/dask/dataframe/core.py
@@ -3736,8 +3736,8 @@ def _emulate(func, *args, **kwargs):
Apply a function using args / kwargs. If arguments contain dd.DataFrame /
dd.Series, using internal cache (``_meta``) for calculation
"""
- with raise_on_meta_error(funcname(func), udf=kwargs.pop('udf', False)):
- return func(*_extract_meta(args, True), **_extract_meta(kwargs, True))
+ kwargs.pop('udf')
+ return func(*_extract_meta(args, True), **_extract_meta(kwargs, True))
I imagine that, like with the groupby aggregations work, this will end up triggering many small PRs in cudf.
Potential fix here: https://github.com/rapidsai/cudf/pull/1396
Next thing I run into, searchsorted
In [1]: import cudf, dask.dataframe as dd
...:
...: # Make a dataframe, we'd like to divide the data by y
...: df = cudf.DataFrame({'x': [1, 2, 3, 4, 5, 6], 'y': [0, 4, 0, 4, 0, 4]})
...:
...: # Split it up into a few partitions with Dask
...: ddf = dd.from_pandas(df, npartitions=3)
...:
...: # Try to create a new dataframe with two partitions sorted on the y column, split by y 0->2 and 2->4
...: dd.shuffle.rearrange_by_divisions(ddf, column='y', divisions=[0, 2, 4], shuffle='tasks')
Out[1]: <dask_cudf.DataFrame | 40 tasks | 2 npartitions>
In [2]: _.compute(scheduler='single-threaded')
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
... <removed for clarity>
TypeError: can't compute boolean for <class 'cudf.dataframe.series.Series'>
In [3]: debug
> /home/nfs/mrocklin/cudf/python/cudf/dataframe/series.py(325)__bool__()
323 into a boolean.
324 """
--> 325 raise TypeError("can't compute boolean for {!r}".format(type(self)))
326
327 def values_to_string(self, nrows=None):
ipdb> up
> /home/nfs/mrocklin/miniconda/envs/cudf/lib/python3.7/site-packages/pandas/core/series.py(2337)searchsorted()
2335 sorter = ensure_platform_int(sorter)
2336 result = self._values.searchsorted(Series(value)._values,
-> 2337 side=side, sorter=sorter)
2338
2339 return result[0] if is_scalar(value) else result
ipdb>
> /home/nfs/mrocklin/dask/dask/dataframe/shuffle.py(434)set_partitions_pre()
432
433 def set_partitions_pre(s, divisions):
--> 434 partitions = pd.Series(divisions).searchsorted(s, side='right') - 1
435 partitions[(s >= divisions[-1]).values] = len(divisions) - 2
436 return partitions
ipdb>
Searchsorted work is happening here: https://github.com/rapidsai/cudf/pull/2156
Just a minor update here - After #2156 goes through, the next issue we run into is the very next line of set_partitions_pre:
~/workspace/cudf-dask-devel/dask/dask/dataframe/shuffle.py in set_partitions_pre(s, divisions)
546
547 def set_partitions_pre(s, divisions):
548 partitions = pd.Series(divisions).searchsorted(s, side='right') - 1
--> 549 partitions[(s >= divisions[-1]).values] = len(divisions) - 2
550 return partitions
AttributeError: 'Series' object has no attribute 'values'
Since values is not a cudf.Series property, perhaps the solution is as simple as making it one (and just returning the column values on host)?
Adding a simple values property to cudf.Series does seem to get us through the small test.
Changes to cudf/python/cudf/cudf/dataframe/series.py:
@property
def values(self):
return self._values
@property
def _values(self):
return self.to_pandas().values
Test:
In [1]: import cudf, dask.dataframe as dd
...:
...: # Make a dataframe, we'd like to divide the data by y
...: df = cudf.DataFrame({'x': [1, 2, 3, 4, 5, 6], 'y': [0, 4, 0, 4, 0, 4]})
...:
...: # Split it up into a few partitions with Dask
...: ddf = dd.from_pandas(df, npartitions=3)
...:
...: # Try to create a new dataframe with two partitions sorted on the y column, split by y 0->2 and 2->4
...: out = dd.shuffle.rearrange_by_divisions(ddf, column='y', divisions=[0, 2, 4], shuffle='tasks')
...:
...: # compute in a single thread so it's easy to use %pdb and %debug
...: print(out.compute(scheduler='single-threaded'))
x y
0 1 0
2 3 0
4 5 0
1 2 4
3 4 4
5 6 4
@kkraus14 - Is there a reason we might want to avoid adding a values property to Series (especially one that returns a non-device array)?
@rjzamora Just tried a small example of set_index this morning and saw the same issue of missing the values property. #2395
@kkraus14 - Is there a reason we might want to avoid adding a
valuesproperty to Series (especially one that returns a non-device array)?
So copying an entire column from device to host? That would be very expensive.
548 partitions = pd.Series(divisions).searchsorted(s, side='right') - 1One issue I see above is a hard dependency on pandas.
549 partitions[(s >= divisions[-1]).values] = len(divisions) - 2
If partitions were to be a cudf series instead of a pandas series , values would not need to be a non-device array
@jrhemstad Sorry - That was a silly question!
@ayushdg - Right. I guess I am just unsure of the best way (if possible) to handle both pandas and cudf using the same logic in Dask, but I haven't exactly tried much yet :)
Some options to explore:
- Can
.valuesreturn something like a cupy array (if it is installed)? Do we strictly need to use
.values? Is this because the result ofsearchsortedis a numpy array rather than a Series?def set_partitions_pre(s, divisions): partitions = pd.Series(divisions).searchsorted(s, side='right') - 1 partitions[(s >= divisions[-1]).values] = len(divisions) - 2 return partitions
I agree that it would be good not to call .to_pandas() here and try to keep things on the device.
See also some conversation about Series.values here: https://github.com/rapidsai/cudf/issues/1824
Also see: #2373 which attempts to add .values support by moving to host. Not sure if it's the best option. There is some discussion echoing this point on that pr as well.
@mrocklin I explored some options to avoid copying the entire column from device to host. I made this simple gist with some experiments.
For a dataframe with 1e8 rows, the dask+pandas version takes about 13.5s, while the dask+cudf version takes about 7.6s. If I use a to_pandas()-based values property, the operation takes ~11s
Note that the gist also shows that the cudf version of searchsorted is a bit different than the pandas version (it returns a series, rather than a numpy array). However, this seems reasonable to me..
7.6s sounds very high, what's taking the time in this situation? Could you share a profile?
7.6s sounds very high, what's taking the time in this situation? Could you share a profile?
@kkraus14 agreed - I'll take a closer look and collect a profile
Note that the gist also shows that the cudf version of searchsorted is a bit different than the pandas version (it returns a series, rather than a numpy array). However, this seems reasonable to me..
So lets take a look at the set_partitions_pre function for a moment
def set_partitions_pre(s, divisions):
partitions = pd.Series(divisions).searchsorted(s, side="right") - 1
partitions[(s >= divisions[-1]).values] = len(divisions) - 2
return partitions
The reason that .values has come up is in the second line, where presumably we're trying to index into partitions with something. If it's a numpy array (as is the case when s is a pandas.Series) then we apparently need a .values call (although I would be a surprised if NumPy couldn't figure things out here). (for background, I think that this is saying "Hey! Every row that is targetting something beyond the last parittion, go to the last partition instead!" I have no idea why we need that, but there is probably some test for it that would fail that comes up in some corner case (this is worth checking to make sure though).
So, as you say, we might have a type issue in that cudf returns a series rather than an array-like. This seems to me, as you say, a sensible choice of cudf's part. So if we focus on just this function we might do a few things:
- Implement
.valuesand coerce things to an array-like - Not implement
.valuesbut keep things as a series all the way through, perhaps with anif is_arraylike(partitions)branch - ??
To figure out the right approach we probably need to zoom out a bit and look at the context in which this is called, and what happens next to these objects. Are the next operations also array/dataframe agnostic, or will we need to coerce at some point anyway?
It looks like the next thing that happens is that they get assigned back into the dataframe. So if we go the array route it would be good to ensure that this is possible. If we go the series route then we might not care.
df2 = df.assign(_partitions=partitions)
I'm curious how much works if we add an is_arraylike condition like the following:
def set_partitions_pre(s, divisions):
partitions = pd.Series(divisions).searchsorted(s, side="right") - 1
index = s > divisions[-1]
if is_arraylike(partitions):
index = index.values
partitions[index] = len(divisions) - 2
return partitions
Or if we can remove the .values call entirely, or if this causes something to break, what is it, and should we fix that instead?
def set_partitions_pre(s, divisions):
partitions = pd.Series(divisions).searchsorted(s, side="right") - 1
partitions[s > divisions[-1]] = len(divisions) - 2
return partitions
Thanks for the feedback @mrocklin! Just to provide a bit more information: For the "cupy" experiment I shared, I am using a custom set_partitions_pre_cupy_series function, which takes in divisions as a series (e.g. cupy.Series):
def set_partitions_pre_cupy_series(s, divisions):
partitions = (divisions.searchsorted(s, side="right") - 1).values_cupy
partitions[(s >= divisions.iloc[-1]).values_cupy] = len(divisions) - 2
return partitions
This ultimately returns partitions as a cupy array. To do this, I added a (rough) values_cupy property to Series:
@property
def values_cupy(self):
try:
import cupy
return cupy.asarray(self._column._data.mem)
except ImportError:
return self.to_array()
Overall, I am thinking it is likely best to do something like the is_arraylike(partitions) branch you mentioned. It may also make sense to remove the explicit pandas dependency by changing the divisions input type to be a series. The question I am struggling with at the moment is how to efficienly reset the partitions elements in a general way
I have no idea why we need that, but there is probably some test for it that would fail that comes up in some corner case (this is worth checking to make sure though).
This actually happens for every value belonging to the last partition here. So, might not be much of a corner case.
Regarding the values property: #2373 added this today. So, we don't have to worry about implementing a host-based values definition if it is needed.
Regarding performance:
As far as I can tell, a good chunk of time is likely python/dask. Otherwise, there seems to be a good chunk of time (~45%) in Dask's shuffle_group, which does make some pandas and numpy-specific calls.
Just a followup - The test I described above uses cupy to calculate the new partition of each row. Unfortunately, the next phase of actually sorting the dataframe into these new partitions requires everything to be copied to host anyway.
The actual splitting of the dataframe into new partitions (within each of the original partitions) is accmplished in shuffle_group (and shuffle_group_2). My profiling indeed shows that these functions account for much of the rearrange_by_divisions call. Here is shuffle_group (shuffle_group_2 is similar):
def shuffle_group(df, col, stage, k, npartitions):
""" Splits dataframe into groups ..."""
if col == "_partitions":
ind = df[col] # True in our case
else:
ind = hash_pandas_object(df[col], index=False)
c = ind._values # Need `_values` property in cudf series for this
typ = np.min_scalar_type(npartitions * 2)
c = np.mod(c, npartitions).astype(typ, copy=False)
np.floor_divide(c, k ** stage, out=c)
np.mod(c, k, out=c)
indexer, locations = groupsort_indexer(c.astype(np.int64), k) # Numpy/pandas specific
df2 = df.take(indexer)
locations = locations.cumsum()
parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])]
return dict(zip(range(k), parts))
Like set_partitions_pre, this function introduces a couple issues for cudf:
- We are assuming a
_valuesproperty exists for the series/column being used for partitioning. This is not true for cudf (I believe even #2373 left this out). - Pandas'
groupsort_indexerfunction requires that a numpy array be input for the first argument. This is fine if we return a numpy array for_values, but not if we use a zero-copy cupy array
Overall, the groupsort_indexer function is pretty simple. It takes in an array of groups (partition numbers in our case), and the total number of groups (number of partitions). The output is the new group-sorted index, and an array with the number of items in each group. @kkraus Does a similar algorithm already exist in cudf?
cc'ing @kkraus14 (note the 14 in his username)
Repeating the question here:
Overall, the groupsort_indexer function is pretty simple. It takes in an array of groups (partition numbers in our case), and the total number of groups (number of partitions). The output is the new group-sorted index, and an array with the number of items in each group. @kkraus Does a similar algorithm already exist in cudf?
I think that in private conversation Keith also mentioned that there may be
some operation in cudf that does several of these steps all at once, which
may be another approach
On Wed, Jul 31, 2019 at 1:59 PM Richard (Rick) Zamora <
[email protected]> wrote:
Just a followup - The test I described above uses cupy to calculate the
new partition of each row. Unfortunately, the next phase of actually
sorting the dataframe into these new partitions requires everything to be
copied to host anyway.The actual splitting of the dataframe into new partitions (within each of
the original partitions) is accmplished in shuffle_group (and
shuffle_group_2). My profiling indeed shows that these functions account
for much of the rearrange_by_divisions call. Here is shuffle_group (
shuffle_group_2 is similar):def shuffle_group(df, col, stage, k, npartitions):
""" Splits dataframe into groups ..."""
if col == "_partitions":
ind = df[col] # True in our case
else:
ind = hash_pandas_object(df[col], index=False)c = ind._values # Need `_values` property in cudf series for this typ = np.min_scalar_type(npartitions * 2) c = np.mod(c, npartitions).astype(typ, copy=False) np.floor_divide(c, k ** stage, out=c) np.mod(c, k, out=c) indexer, locations = groupsort_indexer(c.astype(np.int64), k) # Numpy/pandas specific df2 = df.take(indexer) locations = locations.cumsum() parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])] return dict(zip(range(k), parts))Like set_partitions_pre, this function introduces a couple issues for
cudf:
- We are assuming a _values property exists for the series/column
being used for partitioning. This is not true for cudf (I believe even
#2373 https://github.com/rapidsai/cudf/pull/2373 left this out).- Pandas' groupsort_indexer function requires that a numpy array be
input for the first argument. This is fine if we return a numpy array for
_values, but not if we use a zero-copy cupy arrayOverall, the groupsort_indexer function
https://github.com/pandas-dev/pandas/blob/c7a1321029e07ee6d7ea30036649b488b2e362f7/pandas/_libs/algos.pyx#L151
is pretty simple. It takes in an array of groups (partition numbers in our
case), and the total number of groups (number of partitions). The output is
the new group-sorted index, and an array with the number of items in each
group. @kkraus https://github.com/kkraus Does a similar algorithm
already exist in cudf?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTBLIIHEKHAFSZ45R2TQCH4JLA5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD3IRHYQ#issuecomment-517018594,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTDDFE3WSK5FT2TUBLLQCH4JLANCNFSM4ID3GGRA
.
We currently only have a hash partitioner function which does the equivalent of hash the input dataframe based on some key column(s), run a modulo against the number of partitions N, split the dataframe into N dataframes based on the hash % N.
This doesn't seem quite the same, no?
No, not quite the same.
At this point we have a dataframe with a new column that has the output partition of each row. We now want to split our dataframe up into many small dataframes, one for each output partition.
We do this by creating an indexer which will effectively sort things by that column, and then calling take to do the reordering, and then calling iloc many times to split things apart (though this could be a bulk operation in preferred
c = ... # the output partition locations
indexer, locations = groupsort_indexer(c.astype(np.int64), k) # link to this above in Rick's comment
df2 = df.take(indexer)
locations = locations.cumsum()
parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])]
At this point we have a dataframe with a new column that has the output partition of each row. We now want to split our dataframe up into many small dataframes, one for each output partition.
So in libcudf we already have a gdf_hash_partition algorithm that does this, but uses the hash values of DF rows to determine the partition mapping. (It would appear the shuffle_group algorithm has the option to do this in the else branch).
It would appear that here you're instead using an arbitrary column of values to determine the partition mapping.
It would be fairly trivial to split up and generalize the gdf_hash_partiton function to do what you need here.
All of this code could be replaced with a single libcudf function that I'm guessing will be orders of magnitude faster.
c = np.mod(c, npartitions).astype(typ, copy=False)
np.floor_divide(c, k ** stage, out=c)
np.mod(c, k, out=c)
indexer, locations = groupsort_indexer(c.astype(np.int64), k) # Numpy/pandas specific
df2 = df.take(indexer)
locations = locations.cumsum()
parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])]
Right, we're being a little bit careful about where things go. Sometimes
we move them around in multiple stages, or have particular destinations in
mind based on values.
Putting all of those lines into a single operation sounds fine from my
perspective.
On Thu, Aug 1, 2019 at 1:38 PM Jake Hemstad notifications@github.com
wrote:
At this point we have a dataframe with a new column that has the output
partition of each row. We now want to split our dataframe up into many
small dataframes, one for each output partition.So in libcudf we already have a gdf_hash_partition algorithm that does
this, but uses the hash values of DF rows to determine the partition
mapping.It would appear that here you're instead using an arbitrary column of
values to determine the partition mapping.It would be fairly trivial to split up and generalize the
gdf_hash_partiton function to do what you need here.All of this code could be replaced with a single libcudf function that I'm
guessing will be orders of magnitude faster.c = np.mod(c, npartitions).astype(typ, copy=False) np.floor_divide(c, k ** stage, out=c) np.mod(c, k, out=c) indexer, locations = groupsort_indexer(c.astype(np.int64), k) # Numpy/pandas specific df2 = df.take(indexer) locations = locations.cumsum() parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])]—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTH5N4HMEBLJMOFCEHLQCNCV3A5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD3L2LFY#issuecomment-517449111,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTBMW7OP43M7K467QMDQCNCV3ANCNFSM4ID3GGRA
.
Right, we're being a little bit careful about where things go. Sometimes we move them around in multiple stages, or have particular destinations in mind based on values. Putting all of those lines into a single operation sounds fine from my perspective.
I'm a bit confused because that sounds different than what you originally described.
To be clear, what I'm envisioning is an algorithm that does the following:
Inputs:
- A DataFrame with
Nrows - A
partition_mapofNvalues[0, m)
Outputs:
mDataFrames where DFicontains all of the rows from the input DF wherepartition_map[j] == i
This is more generic than I originally envisioned. There's opportunity for optimizing this further by combining in the step of first generating the partition_map.
That sounds good to me
Out of curiosity, how is the partition_map being generated for this particular use case? Obviously it has to do with which partition a row maps to based on the sorting, but I'm curious how that's being determined.
Thanks for the input @jrhemstad @kkraus14 @mrocklin! This is super useful. Sorry - I didn't see this recent discussion until now.
Out of curiosity, how is the partition_map being generated for this particular use case? Obviously it has to do with which partition a row maps to based on the sorting, but I'm curious how that's being determined.
If I follow the question correctly, the partition map (partitions) is currently determined using searchsorted:
partitions = pd.Series(divisions).searchsorted(s, side="right") - 1
where divisions is a list of divisions along the index/column (s) that we want to partition on.
So it sounds like the size of partitions is equal to the number of desired partitions. That's not quite the same thing as the partition_map I described, which is equal in size to the number of rows in the DF being partitioned.
How do you determine the mapping of each row in the DF to a particular partition? (this is what the partition_map I was describing is)
In the case I described, the size of partitions is indeed equal to the number of rows in the DF being partitioned, because s corresponds to a full column.
~What is the size of divisions? I think I'm misunderstanding how searchsorted works.~
Nevermind, I understand now. divisions is equal in size to the number of partitions, and you're using search_sorted to generate the partition_map from divisions.
For my own elucidation, how is divisions determined?
It would actually be very useful if you could just provide a high level description of the distributed sorting algorithm.
It would also be helpful to understand how many partitions there will be, a rough range of orders of magnitude would be sufficient.
Thats right - divisions is just a list with length (npartitions + 1). There are ways divisions can be calculated, but you can really just think of divisions as a user input for now. For example, the user can specify that they want to partition along some column y using divisions = [0,0.3,0.6].
To do this, they might call:
df = pd.DataFrame( {'x': [i for i in range(size)], 'y': np.random.randn(size)} )
ddf = dd.from_pandas(df, npartitions=3)
out = dd.shuffle.rearrange_by_divisions(ddf, column='y', divisions=[0,0.3,0.6], shuffle='tasks')
result_df = out.compute()
rearrange_by_divisions will start by calculating the partition_map (partitions) you described (using search_sorted). The function looks like this:
def set_partitions_pre(s, divisions):
partitions = pd.Series(divisions).searchsorted(s, side="right") - 1
partitions[(s >= divisions[-1]).values] = len(divisions) - 2
return partitions
After we have partitions, we need call rearrange_by_column to perform the actual shuffling (either in disk or in memory). On a high level, the in-memory version (rearrange_by_column_tasks) does two things:
- Uses
shuffle_groupwithin each of the old partitions to split/sort the local dataframe according to the new partitions (Note that this is wheregroupsort_indexercomes into play) - Concatenates the resulting splits into the new divisions
My understanding is that the number of partitions is typically orders of magnitude smaller than the size of the dataframe in practice (in order for each worker/task to have sufficient data/computation for efficiency)
@mrocklin feel free to correct me or expand.
Thanks @rjzamora that's very helpful.
I'm trying to think what the right primitive(s) is for libcudf/cuDF to expose to accelerate this operation.
We have a few options:
- We could make a primitive that could entirely replace
rearrange_by_divisions
- You give us a DF and a set of divisions, and we'll partition up the DF
- This would likely be the fastest
- We could keep Dask's
rearrange_by_division, and instead replaceshuffle_group
- That is effectively what I proposed here: https://github.com/rapidsai/cudf/issues/2452
- This is similar to what we did with
gdf_hash_partitionfordask-cudf's distributed Join and it was very effective, see https://github.com/dask/dask/issues/4876 - Dask would still compute the
partition_map, but cuDF would handle doing the actual partitioning - This would be the more general option
Curious what @kkraus14 @harrism think the right primitive to expose is here.
I might be missunderstanding, but since rearrange_by_column_tasks operates on a dask dataframe, the libcudf/cuDF primitives would most likely need to be more fine-grained (along the lines of option 2). It would be great to be wrong about this :)
From my perspective, it seems useful to replace the set_partitions_pre function (shown above) with a libcudf/cuDF primitive, as well as shuffle_group, and possibly shuffle_group_2. The latter two are both based on pandas' groupsort_indexer at the moment. I will look through these functions in detail to get a better idea of if/how the logic can be combined.
In order to avoid centralized overhead, dask.dataframe will sometimes move things around in a couple of stages. What we need is the in-memory operation that occurs in each stage. This operation is shuffle_group. Note that it includes a bit of finicky math here:
c = ind._values
typ = np.min_scalar_type(npartitions * 2)
c = np.mod(c, npartitions).astype(typ, copy=False)
np.floor_divide(c, k ** stage, out=c)
np.mod(c, k, out=c)
And then the actual splitting
indexer, locations = groupsort_indexer(c.astype(np.int64), k)
df2 = df.take(indexer)
locations = locations.cumsum()
parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])]
return dict(zip(range(k), parts))
I think that a single function in cudf that did all of this would be great. I would also understand if cudf only wanted to implement the second part, which is a bit more generally useful.
@mrocklin can you explain the finicky math part?
It sounds like the partition mapping returned by set_partitions_pre isn't the final mapping and requires some additional modification.
I've explained the motivation here as a PR with a nicer docstring. If you
need the specific approach let me know. It'll probably take some time to
write down, so I haven't done it yet. If you're just curious here then
I'll pass for now in the interests of time. If it's important for solving
this problem then I'll prioritize this appropriately.
Next time we're together perhaps we can find a whiteboard, go through it,
and then I can pester you to write it up into docs somewhere? :)
On Fri, Aug 2, 2019 at 8:59 AM Jake Hemstad notifications@github.com
wrote:
@mrocklin https://github.com/mrocklin can you explain the finicky math
part?It sounds like the partition mapping returned by set_partitions_pre isn't
the final mapping and requires some additional modification.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTBFWPMOCJSTLSK53Y3QCRKW5A5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD3OE7FA#issuecomment-517754772,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTFLQF2DZ2TXUT7UBILQCRKW5ANCNFSM4ID3GGRA
.
I forgot to link to the PR: https://github.com/dask/dask/pull/5213
I think that a single function in cudf that did all of this would be great.
I can't develop a single function to do shuffle_group if I don't understand the finicky math part.
So the best I can do is what I described here: https://github.com/rapidsai/cudf/issues/2452 which requires you to materialize the final partition_map for a DF.
Yeah, that looks great. That's effectively the second part of the two parts described here: https://github.com/rapidsai/cudf/issues/2272#issuecomment-517727092
indexer, locations = groupsort_indexer(c.astype(np.int64), k)
df2 = df.take(indexer)
locations = locations.cumsum()
parts = [df2.iloc[a:b] for a, b in zip(locations[:-1], locations[1:])]
return dict(zip(range(k), parts))
The first part, the finicky path part ...
c = ind._values
typ = np.min_scalar_type(npartitions * 2)
c = np.mod(c, npartitions).astype(typ, copy=False)
np.floor_divide(c, k ** stage, out=c)
np.mod(c, k, out=c)
We can probably just do with cupy, assuming that we can get a device-array-like out of a cudf series, perhaps with a .values attribute that does this.
If you wanted to combine the two steps, that would be fine with me, my sense is that we should keep them separate first for the sake of generality, and only proceed to merge them if it's a large bottleneck (or if it's hard for cudf to implement a .values attribute that generates a device-array-like object (which may be the case))
I don't think that you need to actually understand the finicky math here in order to implement it. The c attribute is something like your partition_map. We can't go directly from input partition to output partition all in one go because that generates millions of tasks. So we do this in stages.
We can probably just do with cupy, assuming that we can get a device-array-like out of a cudf series, perhaps with a .values attribute that does this.
Just a note that cupy can (as far as I can tell) be used to produce a device array for the values of a cudf series. I've already tested this by tweaking set_partitions_pre to do something like this:
def set_partitions_pre_cupy(s, divisions):
partitions = (cudf.Series(divisions).searchsorted(s, side="right") - 1).values_cupy
partitions[(s >= divisions[-1]).values_cupy] = len(divisions) - 2
return partitions
where the values_cupy property is just returning cupy.asarray(self._column._data.mem)
We can probably just do with cupy, assuming that we can get a device-array-like out of a cudf series, perhaps with a
.valuesattribute that does this.
If we use CuPy here how do we handle nulls? We can't.
You have .values raise NotImplementedError if there are nulls.
In this case, we won't have any nulls. In the future, maybe we'll figure
out something better.
On Fri, Aug 2, 2019 at 4:52 PM Keith Kraus notifications@github.com wrote:
We can probably just do with cupy, assuming that we can get a
device-array-like out of a cudf series, perhaps with a .values attribute
that does this.If we use CuPy here how do we handle nulls? We can't.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTBTK6SR25ZJCC7LHGDQCTCCBA5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD3PCFNQ#issuecomment-517874358,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTALACOF3TKPSBVL5R3QCTCCBANCNFSM4ID3GGRA
.
If we hit the first branch of the shuffle:
if col == "_partitions":
ind = df[col] # True in our case
ind can absolutely have nulls here and if it does the codepath with CuPy will break. If we just had .values return a cuDF index or a cuDF series we could then use NEP18 to handle the np.mod and np.floordivide calls following it with null support nicely until we get into the actual data movement for partitioning, where our partitioning functions will handle nulls.
We've assigned _partitions ourselves. It's a unsigned integer column
with no nulls.
On Fri, Aug 2, 2019 at 4:59 PM Keith Kraus notifications@github.com wrote:
If we hit the first branch of the shuffle:
if col == "_partitions": ind = df[col] # True in our caseind can absolutely have nulls here and if it does the codepath with CuPy
will break. If we just had .value return a cuDF index or a cuDF series we
could then use NEP18 to handle the np.mod and np.floordivide calls
following it with null support nicely until we get into the actual data
movement for partitioning, where our partitioning functions will handle
nulls.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTDPOT6TCOXSPSF35LLQCTC7ZA5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD3PCNDQ#issuecomment-517875342,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTCQ4XJCCAPBKEK3JRLQCTC7ZANCNFSM4ID3GGRA
.
Okay, so it sounds like if we can resolve the CuPy packaging issues with regards to cuML / NCCL then we can use it for the .values return in this situation. Thanks for the clarifications @mrocklin.
Should be relatively straightforward especially in using #2433 to construct the CuPy array with zero copy.
Should be relatively straightforward
@kkraus14 no pressure (well, a little) but do you have an expected timeline on this? This affects some planning on how to handle this issue.
Just to summarize the status here:
There is currently a PR in Dask with the goal of supporting cudf-based dask data frames for
rearrange_by_divisionsandset_index. This PR does not aim for an optimal solution, but for simple changes to dask. Hopefully we can build upon the existing changes there to produce a real solutionBoth
rearrange_by_divisionsandset_indexrely on two general steps: (step 1) The generation of a partition map, which labels the new partition for each row in the dataframe, and (step 2) The actual sorting/rearranging of the data frame into the new partitionsThe generation of the partition map (step 1 above) relies on the
set_partitions_prefunctionThe sorting/rearranging (step 2 above) relies on
rearrange_by_column_tasks, which is often dominated byshuffle_groupandshuffle_group_2. The functions, in turn, use pandas’groupsort_indexerto split each old partition into the pieces that belong to each new partition
The current plan: Accelerate the
shuffle_groupfunctions by providing a gpu-accelerated replacement forgroupsort_indexer, and to avoid moving data to disk by leaning on cupy when possible... Do I have this right?
Accelerate the shuffle_group functions by providing a gpu-accelerated replacement for groupsort_indexer, and to avoid moving data to disk by leaning on cupy when possible... Do I have this right?
OK, so it looks like we're currently blocked on
- a cupy
.valuesattribute (probably shortly after https://github.com/rapidsai/cudf/pull/2464) - partitioning a table based on a partition-map https://github.com/rapidsai/cudf/issues/2452
@kkraus14 @harrism for scheduling do you have an estimate about when those might finish? If they're likely to take a while (everyone has lots of work to do) are they things that someone like @rjzamora could solve, or does it make more sense to have someone more familiar with that code handle them?
Some more information that is likely important, and may motivate a more-complete gpu version of shuffle_group... Here is a snakeviz profile of shuffle_group during a rearrange_by_divisions operation on a dataframe with 1e7 rows and 3 partitions:
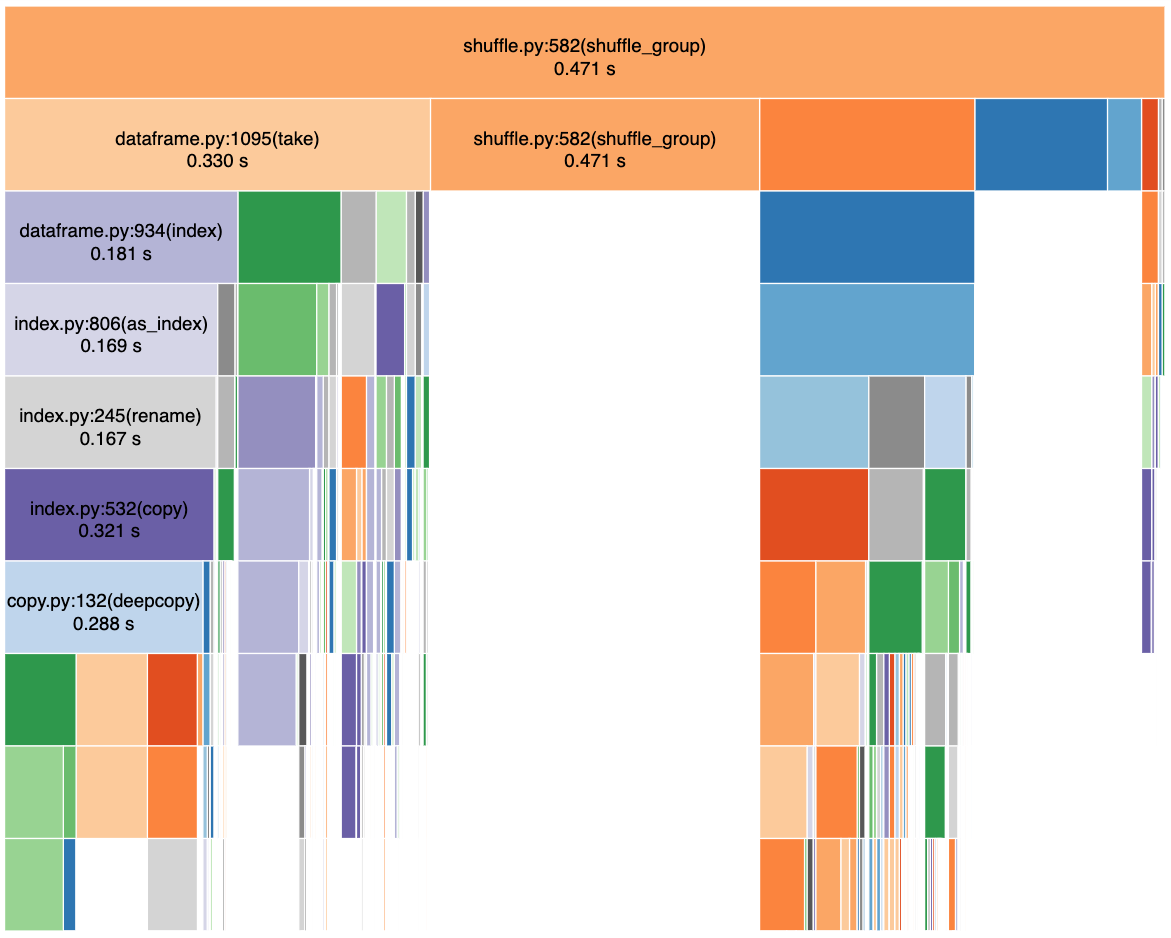
Notes (counting rows from top to bottom):
- The dark blue block in the second row is
groupsort_indexer - The dark orange block in second row (and below) is cudf indexing (
_getitem_tuple_arg) within list comprehension in dask
@rjzamora any chance you can dump an html file that will allow us to explore this data ourselves? Curious to see what else is in this callstack.
Rather than dumping an HTML file I recommend producing a pstats output file
by using %%prun in a notebook or the cProfile module from the command line
with the -D flag . That file can then be shared and others can run
snakeviz on that file.
On Mon, Aug 12, 2019 at 12:47 PM Keith Kraus notifications@github.com
wrote:
@rjzamora https://github.com/rjzamora any chance you can dump an html
file that will allow us to explore this data ourselves? Curious to see what
else is in this callstack.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTHRCMSJ2CJT3HULJSLQEGHYNA5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD4DEBBY#issuecomment-520503431,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTEE2RHXDJ7RJ552KITQEGHYNANCNFSM4ID3GGRA
.
See
https://ipython.readthedocs.io/en/stable/interactive/magics.html#magic-prun and
-D
On Mon, Aug 12, 2019 at 12:49 PM Matthew Rocklin mrocklin@gmail.com wrote:
Rather than dumping an HTML file I recommend producing a pstats output
file by using %%prun in a notebook or the cProfile module from the command
line with the -D flag . That file can then be shared and others can run
snakeviz on that file.On Mon, Aug 12, 2019 at 12:47 PM Keith Kraus notifications@github.com
wrote:@rjzamora https://github.com/rjzamora any chance you can dump an html
file that will allow us to explore this data ourselves? Curious to see what
else is in this callstack.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/rapidsai/cudf/issues/2272?email_source=notifications&email_token=AACKZTHRCMSJ2CJT3HULJSLQEGHYNA5CNFSM4ID3GGRKYY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOD4DEBBY#issuecomment-520503431,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AACKZTEE2RHXDJ7RJ552KITQEGHYNANCNFSM4ID3GGRA
.
Thanks for following up on this @kkraus14, and thanks @mrocklin for the suggestion - I am about to run some new tests pretty soon. I will share the raw profile that I collect.
We're probably ready to remove some of the older dask-cudf implementations of set_index/merge/join. I've raised this as a separate issue here: https://github.com/rapidsai/cudf/issues/2598
I thought I'd poke this. @harrism @jrhemstad do either of you have thoughts on what a timeline for the partition_map splitting operation could look like? We're quickly reaching a state where this will be our only remaining blocker.
cc @randerzander
@mrocklin yes, @jrhemstad and I have been a plan. A "fastest time to solution, not necessarily fastest solution" plan is described in #2677. Please review.
Now that all of the issues mentioned above have closed, and cupy packaging has been resolved, are we able to continue deprecating the custom dask_cudf sort code?
My hope is doing so will yield performance and scalability improvements. Several workflows have recently had problems with the dask_cudf implementation of set_index and sort_values.
If it's useful, I'm happy to open new issues with example unexpected behavior, but a quick read of the above makes me feel like you all have a good handle on the problem.
...are we able to continue deprecating the custom dask_cudf sort code?... My hope is doing so will yield performance and scalability improvements. Several workflows have recently had problems with the dask_cudf implementation of set_index and sort_values.
@randerzander Can you clarify the pieces that are still custom to dask_cudf? dask_cudf should now be using the main-line dask implementations of set_index and merge/join. Since sort_values is not supported in dask, we never really made a plan to remove the cudf-specific implementation.
If your comment is more about performance improvements/optimizations, then I can say we are certainly in the process of exploring ways to (1) Improve merge/join performance to better match the expected/available performance, and (2) improve the stability of operations like set_index.
Note that I am currently working on a simple (but experimental) modification to the upstream-dask shuffle and set_partition code. It seems that we can use hashing to moderately reduce some of the memory consumption and and data-copy overhead.
To what extent is this issue satisfied by the current work on providing multi-column distributed sorting primitives (table quantiles, k-way merge) in libcudf?
To what extent is this issue satisfied by the current work on providing multi-column distributed sorting primitives (table quantiles, k-way merge) in libcudf?
100%, this is the primary target for that work.
@kkraus14 both of those new primitives were added. Does this need to stay open?
I defer to @rjzamora if he'd like to keep this open for future discussion or if this can now be closed.
We should probably close this once #4308 is merged. That should provide dask_cudf with the general sorting/set_index functionality targeted by this discussion. After that, the remaining work will be to push as mach dask_cudf code as possible into main-line dask (which can probably be tracked separately).
Now that #4308 is merged, it probably makes sense to close this. We can open new issues in the future to discuss any performance/functionality challenges that come up.
Also, the task of moving the new sort_values and repartition_by_hash functionality into upstream dask can/should be tracked elsewhere.
@randerzander - Feel free to reopen this if the current sort_values solution does not end up meeting your needs. You can also open a more-targeted issue.