Cudf: [BUG] Romberg Numeric Integration kernel compile fails with NvvmError in 0.7, runs fine in 0.6
Description
When I tried to run a test kernel in Rapids 0.7, I get an NvvmError when compiling the kernel. The code works correctly in 0.6.
The code is a kernel to perform Romberg Numeric Integration on a user defined function. This is a common scenario in Bayesian methods where a user may wish to estimate the integral of a probability density function over a given period.
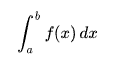
I have tested it with the natural logarithm function and compared the estimate with the actual value given by the definite integral.
Code
Libraries
import cudf
from cudf.dataframe import DataFrame
import numpy as np
import math
import numba
from numba import cuda
Romberg Integration Numba CUDA JIT function
@cuda.jit(device=True)
def integrate(a, b):
steps = 10
table = numba.cuda.local.array(shape=(steps,steps),dtype=numba.float64)
for i in range(steps):
for j in range(steps):
table[i][j] = 0.0
h = (b - a)
table[0, 0] = h * (fn(a) + fn(b)) / 2
for j in range(1, steps):
h /= 2
# extended trapezoidal rule
table[j, 0] = table[j - 1, 0] / 2
s = 0
for i in range(1, 2 ** j + 1, 2):
s += fn(a + i * h)
table[j, 0] += h * s
# richardson extrapolation
for k in range(1, j + 1):
table[j, k] = table[j, k - 1] + \
(table[j, k - 1] - table[j - 1, k - 1]) / (4.0**k - 1.0)
return table[steps-1, steps-1]
Create test dataframe
nelem = 100000000
df = DataFrame()
df['a'] = np.arange(nelem).astype(np.float64)+1
df['b'] = (np.arange(nelem).astype(np.float64)+2.0)*2
print(df)
Kernel Function (estimate returned in e, definite integral returned in y)
def romberg(a, b, e, y):
for i, (A, B) in enumerate(zip(a,b)):
e[i] = integrate(A,B)
y[i] = (B * math.log(B) - B) - (A * math.log(A) - A)
Apply Kernel
%%time
df = df.apply_rows(romberg,incols=['a', 'b'],outcols=dict(e=np.float64,y=np.float64),kwargs=dict())
Expected behavior
Worked without problem in previous versions of Rapids cuDF including 0.5 and 0.6
Environment details (please complete the following information):
- Docker container (unmodified) runtime 0.7 CUDA 10.0 Ubuntu 18.04
- Method of cuDF install: Docker
docker pull rapidsai/rapidsai:0.7-cuda10.0-runtime-ubuntu18.04-gcc7-py3.7 docker run --runtime=nvidia --name rapids0.7 --rm -it -p 8888:8888 -p 8787:8787 -p 8786:8786 -v /home/john/Data/:/data/ -v /home/john/Source/Notebooks:/rapids/notebooks/work rapidsai/rapidsai:0.7-cuda10.0-runtime-ubuntu18.04-gcc7-py3.7
Additional context
Error output
~~~~
NvvmError Traceback (most recent call last)
/conda/envs/rapids/lib/python3.7/site-packages/cudf/dataframe/dataframe.py in apply_rows(self, func, incols, outcols, kwargs, cache_key)
1972 """
1973 return applyutils.apply_rows(self, func, incols, outcols, kwargs,
-> 1974 cache_key=cache_key)
1975
1976 @applyutils.doc_applychunks()
/conda/envs/rapids/lib/python3.7/site-packages/cudf/utils/applyutils.py in apply_rows(df, func, incols, outcols, kwargs, cache_key)
60 applyrows = ApplyRowsCompiler(func, incols, outcols, kwargs,
61 cache_key=cache_key)
---> 62 return applyrows.run(df)
63
64
/conda/envs/rapids/lib/python3.7/site-packages/cudf/utils/applyutils.py in run(self, df, launch_params)
103 bound = self.sig.bind(args)
104 # Launch kernel
--> 105 self.launch_kernel(df, bound.args, **launch_params)
106 # Prepare output frame
107 outdf = df.copy()
/conda/envs/rapids/lib/python3.7/site-packages/cudf/utils/applyutils.py in launch_kernel(self, df, args)
122 blksz = 64
123 blkct = cudautils.optimal_block_count(len(df) // blksz)
--> 124 self.kernelblkct, blksz
125
126
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/compiler.py in __call__(self, args)
763 Specialize and invoke this kernel with *args.
764 '''
--> 765 kernel = self.specialize(args)
766 cfg = kernel[self.griddim, self.blockdim, self.stream, self.sharedmem]
767 cfg(args)
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/compiler.py in specialize(self, *args)
774 argtypes = tuple(
775 [self.typingctx.resolve_argument_type(a) for a in args])
--> 776 kernel = self.compile(argtypes)
777 return kernel
778
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/compiler.py in compile(self, sig)
793 self.definitions[(cc, argtypes)] = kernel
794 if self.bind:
--> 795 kernel.bind()
796 return kernel
797
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/compiler.py in bind(self)
501 Force binding to current CUDA context
502 """
--> 503 self._func.get()
504
505 @property
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/compiler.py in get(self)
379 cufunc = self.cache.get(device.id)
380 if cufunc is None:
--> 381 ptx = self.ptx.get()
382
383 # Link
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/compiler.py in get(self)
350 arch = nvvm.get_arch_option(cc)
351 ptx = nvvm.llvm_to_ptx(self.llvmir, opt=3, arch=arch,
--> 352 *self._extra_options)
353 self.cache[cc] = ptx
354 if config.DUMP_ASSEMBLY:
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/cudadrv/nvvm.py in llvm_to_ptx(llvmir, opts)
500 cu.add_module(libdevice.get())
501
--> 502 ptx = cu.compile(opts)
503 # XXX remove debug_pubnames seems to be necessary sometimes
504 return patch_ptx_debug_pubnames(ptx)
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/cudadrv/nvvm.py in compile(self, **options)
231 for x in opts])
232 err = self.driver.nvvmCompileProgram(self._handle, len(opts), c_opts)
--> 233 self._try_error(err, 'Failed to compile\n')
234
235 # get result
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/cudadrv/nvvm.py in _try_error(self, err, msg)
249
250 def _try_error(self, err, msg):
--> 251 self.driver.check_error(err, "%s\n%s" % (msg, self.get_log()))
252
253 def get_log(self):
/conda/envs/rapids/lib/python3.7/site-packages/numba/cuda/cudadrv/nvvm.py in check_error(self, error, msg, exit)
139 sys.exit(1)
140 else:
--> 141 raise exc
142
143
NvvmError: Failed to compile
NVVM_ERROR_COMPILATION
~~~~
All 10 comments
@sklam any ideas what would cause this error?
I'm unable to replicate the problem using the same docker setup.
Here's the code that I am trying.
import cudf
from cudf.dataframe import DataFrame
import numpy as np
import math
import numba
from numba import cuda
@cuda.jit(device=True)
def fn(x):
return x
@cuda.jit(device=True)
def integrate(a, b):
steps = 10
table = numba.cuda.local.array(shape=(steps,steps),dtype=numba.float64)
for i in range(steps):
for j in range(steps):
table[i][j] = 0.0
h = (b - a)
table[0, 0] = h * (fn(a) + fn(b)) / 2
for j in range(1, steps):
h /= 2
# extended trapezoidal rule
table[j, 0] = table[j - 1, 0] / 2
s = 0
for i in range(1, 2 ** j + 1, 2):
s += fn(a + i * h)
table[j, 0] += h * s
# richardson extrapolation
for k in range(1, j + 1):
table[j, k] = table[j, k - 1] + \
(table[j, k - 1] - table[j - 1, k - 1]) / (4.0**k - 1.0)
return table[steps-1, steps-1]
nelem = 100000
df = DataFrame()
df['a'] = np.arange(nelem).astype(np.float64)+1
df['b'] = (np.arange(nelem).astype(np.float64)+2.0)*2
print(df)
def romberg(a, b, e, y):
for i, (A, B) in enumerate(zip(a,b)):
e[i] = integrate(A,B)
y[i] = (B * math.log(B) - B) - (A * math.log(A) - A)
df = df.apply_rows(romberg,incols=['a', 'b'],outcols=dict(e=np.float64,y=np.float64),kwargs=dict())
print(df)
Note, I have to make two modifications:
- insert an identity function for the missing
fn. - reduce
nelemto fit on my testing machine.
Since I cannot replicate, the problem may lie within the definition of fn. @MurrayData, can you provide the definition of fn that caused the error?
Oh, this is a heisenbug. I can segfault libnvvm sometimes. I will need to investigate more.
It turns out the issue is caused by out-of-sync versions of numba and llvmlite in the docker image.
The llvmlite in the docker image is:
# Name Version Build Channel
llvmlite 0.27.0dev0 py37hf484d3e_19 numba
It is a nightly dev build and not an official release.
The numba in the docker image is:
# Name Version Build Channel
numba 0.41.0 py37h637b7d7_1000 conda-forge
It's a build from conda-forge.
To fix it, I run conda install -c conda-forge numba=0.41.0 llvmlite to ask conda to reinstall llvmlite for numba version 0.41.0. It will install llvmlite=0.26.0.
To verify, save the code in https://github.com/rapidsai/cudf/issues/1877#issuecomment-497462689 as a .py file and run several times. The script will fail in the unmodified docker image sometimes. (For unknown reason, it's less likely to fail in a IPython notebook). Once llvmlite is in-sync, the script will not fail.
Thanks for triaging @sklam, it's much appreciated. cc @mike-wendt @raydouglass @rlratzel would it be possible to resolve this in the container?
I'll update the Dockerfiles to install that version of llvmlite. I'd normally also add the code provided here to repro the issue in a test suite, but it looks like it doesn't reliably reveal the problem.
I'm unable to replicate the problem using the same docker setup.
Here's the code that I am trying.
import cudf from cudf.dataframe import DataFrame import numpy as np import math import numba from numba import cuda @cuda.jit(device=True) def fn(x): return x @cuda.jit(device=True) def integrate(a, b): steps = 10 table = numba.cuda.local.array(shape=(steps,steps),dtype=numba.float64) for i in range(steps): for j in range(steps): table[i][j] = 0.0 h = (b - a) table[0, 0] = h * (fn(a) + fn(b)) / 2 for j in range(1, steps): h /= 2 # extended trapezoidal rule table[j, 0] = table[j - 1, 0] / 2 s = 0 for i in range(1, 2 ** j + 1, 2): s += fn(a + i * h) table[j, 0] += h * s # richardson extrapolation for k in range(1, j + 1): table[j, k] = table[j, k - 1] + \ (table[j, k - 1] - table[j - 1, k - 1]) / (4.0**k - 1.0) return table[steps-1, steps-1] nelem = 100000 df = DataFrame() df['a'] = np.arange(nelem).astype(np.float64)+1 df['b'] = (np.arange(nelem).astype(np.float64)+2.0)*2 print(df) def romberg(a, b, e, y): for i, (A, B) in enumerate(zip(a,b)): e[i] = integrate(A,B) y[i] = (B * math.log(B) - B) - (A * math.log(A) - A) df = df.apply_rows(romberg,incols=['a', 'b'],outcols=dict(e=np.float64,y=np.float64),kwargs=dict()) print(df)Note, I have to make two modifications:
- insert an identity function for the missing
fn.- reduce
nelemto fit on my testing machine.Since I cannot replicate, the problem may lie within the definition of
fn. @MurrayData, can you provide the definition offnthat caused the error?
@sklam:
@cuda.jit(device=True)
def fn(x):
return math.log(x)
Thanks @sklam, @rlratzel and @kkraus14. I derived a new Docker container, from the existing, with
conda install -c conda-forge numba=0.41.0 llvmlite
added and it works fine now.
cc
I'll update the Dockerfiles to install that version of
llvmlite. I'd normally also add the code provided here to repro the issue in a test suite, but it looks like it doesn't reliably reveal the problem.
We shouldn't have to hardcode a version of llvmlite into Docker, if we remove the numba conda channel I believe all should work as expected. I'm not sure how we got this situation as the numba 0.41.0 package from conda-forge has the following constraint: llvmlite >=0.26.0,<0.27.0a0
We shouldn't have to hardcode a version of
llvmliteinto Docker, if we remove the numba conda channel I believe all should work as expected.
Yeah, I realized removing the numba channel solves the problem too after I commented, and I just committed that change. The next nightly builds should be fixed.
I'm not sure how we got this situation as the numba 0.41.0 package from conda-forge has the following constraint:
llvmlite >=0.26.0,<0.27.0a0
dev < a, so the constraint was satisfied.
Most helpful comment
It turns out the issue is caused by out-of-sync versions of numba and llvmlite in the docker image.
The llvmlite in the docker image is:
It is a nightly dev build and not an official release.
The numba in the docker image is:
It's a build from conda-forge.
To fix it, I run
conda install -c conda-forge numba=0.41.0 llvmliteto ask conda to reinstall llvmlite for numba version 0.41.0. It will install llvmlite=0.26.0.To verify, save the code in https://github.com/rapidsai/cudf/issues/1877#issuecomment-497462689 as a
.pyfile and run several times. The script will fail in the unmodified docker image sometimes. (For unknown reason, it's less likely to fail in a IPython notebook). Once llvmlite is in-sync, the script will not fail.