Cudf: groupby() super slow on branch-0.7[BUG]
I found the groupby() is super slower on branch-0.7 than that on branch-0.6. The time output is attached.
df = DataFrame({'cat' : df_v,'cnt' : cnt ,'id' : id, 'cat1': df_v,'doc_length':doc_length})
%timeit df1 = df.groupby(['cat1','cat','id','doc_length'], method='hash').count()
Time outputs
#branch-0.7
17 s ± 443 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
#branch-0.6
34.1 ms ± 3.76 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
All 35 comments
I've heard reports of the same thing from @trxcllnt and @lmeyerov. I believe the issue is isolated to categorical columns.
Furthermore, I suspect the issue must be in the Python layer because as far as the C++ is concerned, categoricals are no different from int32.
Yep full repro in https://github.com/rapidsai/cudf/issues/1329
I suspect this is related to https://github.com/rapidsai/cudf/issues/1673 as well.
I'll go back and compare some results with branch-0.6. Floats are also stupendously slow. Ints appear to be v. fast:
In [1]: import cudf
...: import numpy as np
...: import pandas as pd
...:
...: _SIZE = 500000
...:
...: randx = np.random.randint(_SIZE)
...: randy = np.random.randint(_SIZE)
...: randz = np.random.randint(_SIZE)
...:
...: gdf = cudf.DataFrame({'x': randx, 'y': randy, 'z': randz})
...:
...: %timeit gdf.groupby(['x', 'y']).mean()
...:
46 ms ± 1.29 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
It may be related to multi-column groupby(). I test the single column and two columns in groupby().
df2 = df[['cat1','cnt']]
df3 = df[['cat1','cat','cnt']]
%timeit df2 = df2.groupby(['cat1']).agg({'cnt': 'count'})
%time df3 = df3.groupby(['cat1','cat']).agg({'cnt': 'count'})
Time output:
#branch-0.7
#df2
55.8 ms ± 1.67 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
#df3
CPU times: user 13.2 s, sys: 40 ms, total: 13.2 s
Wall time: 13.2 s
#branch-0.6
#df2
17.8 ms ± 816 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
#df3
CPU times: user 12 ms, sys: 4 ms, total: 16 ms
Wall time: 30.6 ms
I bet you'll have bad numbers on df3.groupby(['cat1']).agg({'cnt': 'count'}) as well: if you are grouping on 1 index, just having the additional columns in there slows things down.
@lmeyerov the performance is acceptable on df3.groupby(['cat1']).agg({'cnt': 'count'})
#branch-0.7
73.4 ms ± 2.33 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
#branch-0.6
17.7 ms ± 546 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
df2.groupby(['cat1']).agg({'cnt': 'count'}) and df3.groupby(['cat1']).agg({'cnt': 'count'}) have the same performance in the branch-0.6.
Mike, I think you just showed the reverse --- 0.7 shows a 5X slowdown . The more columns the df has, even if unused, the slower it goes.
- Leo
Hi @thomcom, I try the 0.7 again. It improves the performance in small dataset. When I try the larger dataset, it is still slow. And when I print the df, it generates errors.
#Branch-0.7
len(df)
62914500
%time df1 = df.groupby(['cat1','cat','id','doc_length'], method='hash').count()
CPU times: user 17.7 s, sys: 12.3 s, total: 30 s
Wall time: 30.3 s
print(df1)
/conda/lib/python3.7/site-packages/cudf-0.7.0.dev0+1567.gffac3e1.dirty-py3.7-linux-x86_64.egg/cudf/dataframe/dataframe.py in __str__(self)
448 nrows = settings.formatting.get('nrows') or 10
449 ncols = settings.formatting.get('ncols') or 8
--> 450 return self.to_string(nrows=nrows, ncols=ncols)
451
452 def __repr__(self):
/conda/lib/python3.7/site-packages/cudf-0.7.0.dev0+1567.gffac3e1.dirty-py3.7-linux-x86_64.egg/cudf/dataframe/dataframe.py in to_string(self, nrows, ncols)
405 if isinstance(self.index, cudf.dataframe.multiindex.MultiIndex) or\
406 isinstance(self.columns, cudf.dataframe.multiindex.MultiIndex):
--> 407 raise TypeError("You're trying to print a DataFrame that contains "
408 "a MultiIndex. Print this dataframe with "
409 ".to_pandas()")
TypeError: You're trying to print a DataFrame that contains a MultiIndex. Print this dataframe with .to_pandas()
#Branch-0.6
len(df)
62914500
%time df1 = df.groupby(['cat1','cat','id','doc_length'], method='hash').count()
CPU times: user 584 ms, sys: 236 ms, total: 820 ms
Wall time: 857 ms
print(df1)
cat id doc_length cnt
cat1
0 0 1 1 59
0 0 2 4 56
0 0 3 2 58
0 0 4 1 59
0 0 5 10 50
0 0 6 13 47
0 0 7 10 50
0 0 8 10 50
0 0 9 10 50
0 0 10 10 50
[8769167 more rows]
@MikeChenfu Can you try passing as_index=False as another parameter to the groupby call? We're working on resolving the MultiIndex performance issues and that should workaround it for the time being.
Thanks @kkraus14. The performance is great. It is about 1s.
@MikeChenfu glad to hear the workaround worked well for you. I'm going to keep this open until we resolve the performance issues in using a MultiIndex.
I was unable to duplicate the workaround
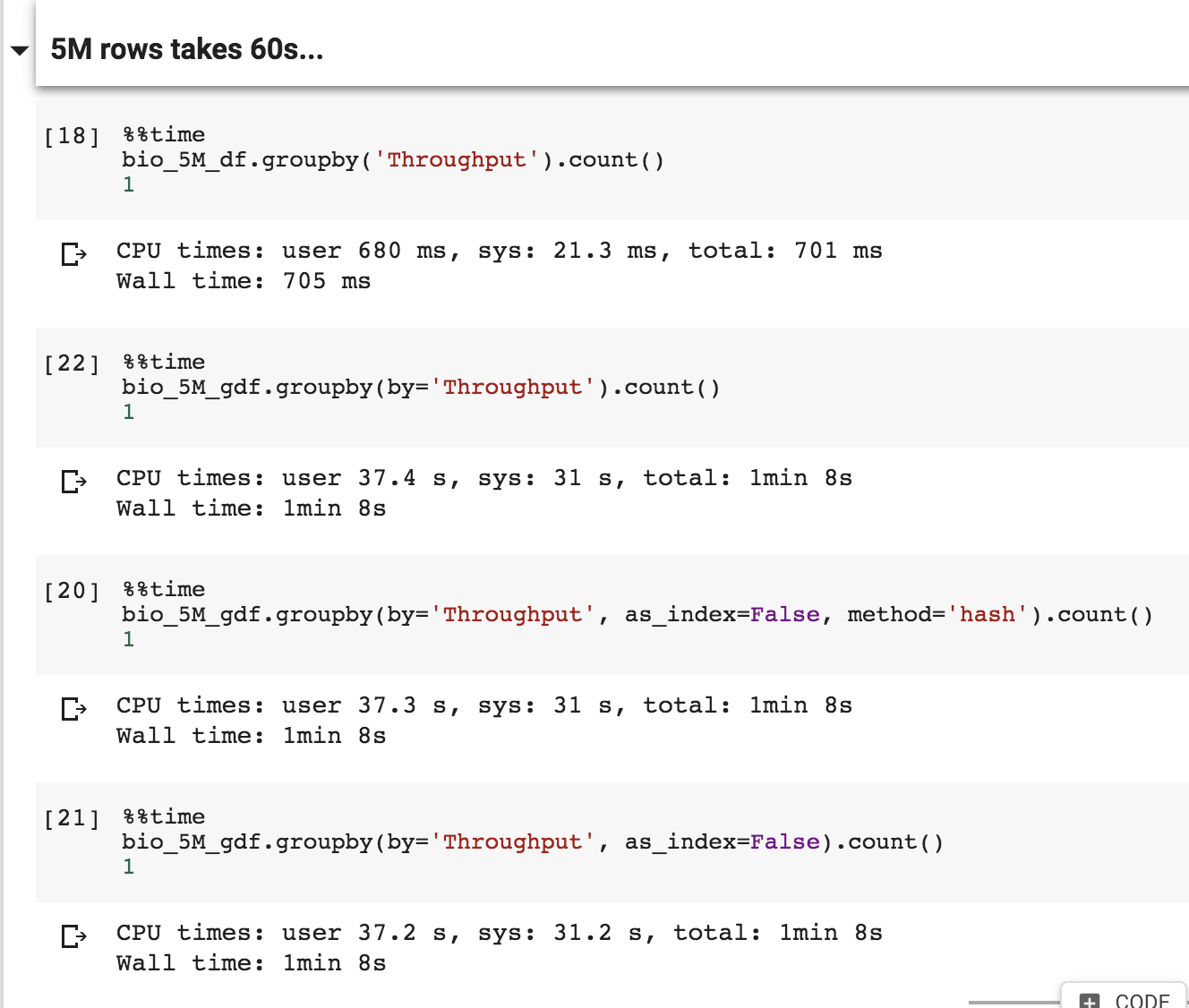
I was unable to duplicate the workaround
That shouldn't be hitting the MultiIndex issues as it's only a single column groupby, will connect offline to see what's going funky.
Added some print statements to the Cython and found the time is actually in the CPP libcudf call:
Before CPP: 0.002177715301513672
After CPP: 49.825637340545654
Before CPP: 49.834495544433594
After CPP: 99.42340469360352
Final: 99.42666935920715
Given there's a major refactor of groupby coming down the pipes, it doesn't make sense to investigate the perf issue in that case until after the refactor has landed.
This should have been autoresolved by https://github.com/rapidsai/cudf/pull/1702
It shows a different performance on the latest master
```
df = DataFrame({'cat' : df_v, 'cnt' : cnt , 'id' : id, 'doc_length' : doc_length})
%time df1 = df.groupby(['cat', 'id', 'doc_length'], as_index=False).count()
gdf = df1[['cat','cnt']]
%time gdf1 = gdf.groupby(['cat'], as_index=False).count()
CPU times: user 140 ms, sys: 120 ms, total: 260 ms
Wall time: 341 ms
CPU times: user 11.5 s, sys: 8.17 s, total: 19.6 s
Wall time: 19.7 s
I test different dataset to show above problem.
In [1]: from cudf import DataFrame
...: a = DataFrame({'b':[1,2,3,4,5,6,7,8] * 100000, 'd':range(800000),'c':range(800000),'cnt':range(800000)})
...: %time a1 = a.groupby(['b','d','c'],as_index=False).count()
...:
...: a2 = a1[['b','cnt']]
...: %time a3 = a2.groupby(['b'],as_index=False).count()
CPU times: user 476 ms, sys: 24 ms, total: 500 ms
Wall time: 525 ms
CPU times: user 1.1 s, sys: 788 ms, total: 1.89 s
Wall time: 1.89 s
In [2]: from cudf import DataFrame
...: a = DataFrame({'b':[1,2,3,4,5,6,7,8] * 1000000, 'd':range(8000000),'c':range(8000000),'cnt':range(8000000)})
...:
...: %time a1 = a.groupby(['b','d','c'],as_index=False).count()
...:
...: a2 = a1[['b','cnt']]
...: %time a3 = a2.groupby(['b'],as_index=False).count()
CPU times: user 76 ms, sys: 112 ms, total: 188 ms
Wall time: 230 ms
CPU times: user 48.5 s, sys: 35.8 s, total: 1min 24s
Wall time: 1min 24s
@MikeChenfu We've identified a performance issue specifically with the count aggregation (which also impacts mean since it uses count under the hood), that we're working on resolving. Could you try with .sum() just to confirm the performance of that aggregation is as expected?
.sum() may also have a performance issue.
In [3]: from cudf import DataFrame
...: a = DataFrame({'b':[1,2,3,4,5,6,7,8] * 1000000, 'd':range(8000000),'c':range(8000000),'cnt':range(8000000)})
...:
...: %time a1 = a.groupby(['b','d','c'],as_index=False).sum()
...:
...: a2 = a1[['b','cnt']]
...: %time a3 = a2.groupby(['b'],as_index=False).sum()
CPU times: user 176 ms, sys: 160 ms, total: 336 ms
Wall time: 398 ms
CPU times: user 1min 22s, sys: 1min, total: 2min 22s
Wall time: 2min 23s
.sum()may also have a performance issue.In [3]: from cudf import DataFrame ...: a = DataFrame({'b':[1,2,3,4,5,6,7,8] * 1000000, 'd':range(8000000),'c':range(8000000),'cnt':range(8000000)}) ...: ...: %time a1 = a.groupby(['b','d','c'],as_index=False).sum() ...: ...: a2 = a1[['b','cnt']] ...: %time a3 = a2.groupby(['b'],as_index=False).sum() CPU times: user 176 ms, sys: 160 ms, total: 336 ms Wall time: 398 ms CPU times: user 1min 22s, sys: 1min, total: 2min 22s Wall time: 2min 23s
Thanks for the reproducer, I can confirm I can reproduce.
After a fun game of git bisect, I narrowed down the offending commit to https://github.com/rapidsai/cudf/commit/542d960b2b16cbc31f51c2ef86267302e947bf65 which replaced the existing device atomic overloads previously used in the groupby implementation for those developed by @kovaltan. I suspect that a code path that was originally using a native atomicAdd is now instead using atomicCAS and is much slower as a result.
This is the repro I've been using:
import numpy as np
import cudf
size = 5373090
min_int = np.iinfo('int32').min
max_int = np.iinfo('int32').max
keith_gdf = cudf.DataFrame()
keith_gdf['col1'] = np.random.randint(0, 2, size=size, dtype='int32')
keith_gdf['col2'] = np.random.randint(max_int, size=size, dtype='int32')
keith_gdf['col3'] = np.random.randint(max_int, size=size, dtype='int32')
output = keith_gdf.groupby('col1').count()
print(output)
When timing with the Linux time command, a "good" run takes ~1.25s walltime and a "bad" run takes ~44s walltime.
CC @harrism
I think the cause is pretty clear... CUDA doesn't support atomicAdd natively on signed 64-bit ints, only unsigned. The groupby above resolves to building a hash table with key-type int and value type long long int. The latter is what the atomicAdd are performed on.
Before commit 542d960, there was this overload in cuDF, which just casts signed to unsigned to do the atomicAdd:
__inline__ __device__ int64_t atomicAdd(int64_t* address, int64_t val)
{
return (int64_t) atomicAdd((unsigned long long*)address, (unsigned long long)val);
}
But now, we go through this path:
__forceinline__ __device__
int64_t atomicAdd(int64_t* address, int64_t val)
{
// `atomicAdd` supports uint64_t, but not int64_t
return cudf::genericAtomicOperation(address, val, cudf::DeviceSum{});
}
This ultimately calls an atomicCAS-based implementation, which may loop and result in a lot of divergence. It's no wonder the performance is so bad...
What I don't understand is why 64-bit types are used at all here, when the Python code above sets all types to 'int32'... @kkraus14 any idea why that is?
I have a tentative fix, which is to change this:
__forceinline__ __device__
int64_t atomicAdd(int64_t* address, int64_t val)
{
// `atomicAdd` supports uint64_t, but not int64_t
return cudf::genericAtomicOperation(address, val, cudf::DeviceSum{});
}
To this:
__forceinline__ __device__
int64_t atomicAdd(int64_t* address, int64_t val)
{
// `atomicAdd` supports uint64_t, but not int64_t
// return cudf::genericAtomicOperation(address, val, cudf::DeviceSum{});
unsigned long long int result = atomicAdd(reinterpret_cast<unsigned long long int*>(address),
static_cast<unsigned long long int>(val));
return static_cast<int64_t>(result);
}
This change reduces the bottleneck kernel time from 17s to 5.4ms (!!!).
However, I'm concerned about the safety of casting a signed to an unsigned value.
What I don't understand is why 64-bit types are used at all here, when the Python code above sets all types to 'int32'... @kkraus14 any idea why that is?
Because count always uses int64 as output.
Because
countalways usesint64as output.
Can count on a table with gdf_size_type rows ever be larger than can be represented in a gdf_size_type? [Answer: no]
OK, here's a summary:
- CUDA (for some reason) implemented atomicAdd for unsigned int64 but not signed int64
- cuDF used to work around this by casting the inputs to unsigned before atomicAdd, but that is wrong, because if you add a negative value you get wrong results. So we can't work around this with that approach.
- I have re-opened an old internal feature request to get signed int64 atomicAdd added to CUDA.
- Until then, we just have to live with slow signed int64
sum()groupby aggregations. Sorry. Workaround: if you can, use a 32-bit type, or use an unsigned type. - We can improve
count()aggregation performance by changing the Python side to use a 32-bit type for the output of count aggregations. This sounds like an easy fix, but I don't know where in the Python code it is. I'm about to go looking...
https://github.com/rapidsai/cudf/issues/1685#issuecomment-492055527
I feel it is safety for casting a signed to an unsigned value only for integer sumation as long as Two's complement is used for a signed number representation.
https://en.wikipedia.org/wiki/Two%27s_complement
But I'd like someone to confirm it is safe for cuda atomicAdd.
@kovaltan is right, I had forgotten about the equivalence of signed and unsigned arithmetic with two's complement. I'm pretty sure that NVIDIA GPUs use two's complement. Otherwise compatibility of code with the CPU would be quite difficult. :) So this means we can fix the slow sum() aggregations for signed int64 too.
Now, PR #1691 supports signed int64_t native atomicAdd using cast.
Please check it.
@MikeChenfu this issue should now be resolved on https://github.com/rapidsai/cudf/tree/release-0.7.2
Closing the issue. Please reopen if you're still seeing the same performance issues.
Thanks @jrhemstad . Is it available on 0.8?
Thanks @jrhemstad . Is it available on 0.8?
The fix was a hotfix to the 0.7 release. I believe it should already be available.
I see the CHANGLOG.md , this PR may not be ready on 0.8.
Once 0.7.2 is released (should be in the next 24 hours) and merged into master, we'll merge master into branch-0.8 so it's available on stable and development branches.