The hierarchical editor is broken and doesn't export in a way Arctos can understand.
The tiered preference model means that the hierarchical editor can be the primary UI for hierarchical classifications; we no longer need "pull" functionality, only push. (DBAs can help with any initial import requests.)
Export can be directly to the classification loader, potentially with status=autoload==>saves directly to Arctos.
This viewpoint would make https://github.com/ArctosDB/arctos/issues/3121 largely irrelevant; it's possible to muck up a hierarchy in the single-record editor, but that's not the primary data so it'll just be overwritten with the next save.
Adding some urgency; I think @sharpphyl is managing "leftovers" that aren't in WoRMS in the "Arctos" classification, which is of limited utility. This would get a lot more utility out of that effort.
All 17 comments
Adding some urgency; I think @sharpphyl is managing "leftovers" that aren't in WoRMS in the "Arctos" classification, which is of limited utility. This would get a lot more utility out of that effort.
Yes, I still have several hundred names/classifications to work on, but my goal is to have nothing in WoRMS (via Arctos) that doesn't have an aphiaID which ensures that it is 100% externally managed. So I'm moving or adding the classifications to Arctos. I'm sure I could use a hierarchical editor to align the higher classifications on the names I'm editing. I could really use a better understanding of exactly how it would work but I think it sounds like a much simpler process. Even though our primary source is WoRMS (via Arctos), we use enough Arctos classifications for me to be engaged in making sure they are consistent.
Is there a mock-up anywhere to test with?
nothing in WoRMS (via Arctos) that doesn't have an aphiaID
Worthy goal; yay consistency!
moving or adding the classifications to Arctos
... where they have a non-zero chance of being turned into dinosaurs, or at least made inconsistent by individual updates.
exactly how it would work
- Start a new Source, preferred somewhere after WoRMS by your collection
- Manage "new source" data in the hierarchical editor, which would work as before but without all the complexity involved in trying to make bits and pieces of inconsistent things temporarily consistent, and with a bit more permanence in mind. Note that most consistency just isn't possible in a hierarchy, and what is possible - a genus skipping family, for example - is very easy to spot/hard to ignore.
- Periodically "save to core Arctos" from the hierarchical tool
- If something pops up in WoRMS, add the AphiaID to "WoRMS (via Arctos)"
Then...
- Your collection will use "WoRMS (via Arctos)" (even if there's something in your new source), then your new source, then (if necessary) perhaps Arctos or some other "fallback" source
- If (when?!) someone makes something in your new source inconsistent (or turns it into a kelp) with the single-record editor, you'll just overwrite it the next time you use that "save to core Arctos" button.
mock-up
Not really, but there's nothing very new in here either - it's all old tools, doing new things under a slightly different model.
much simpler process.
It should be. Finding all of your stuff should be easy; it's in one Source. Moving a genus to a new family is one drag-drop and all of the species and subspecies can't avoid following. Adding/changing subkingdom involves one edit, not thousands. Etc. There are limitations to a hierarchy - all terms need to also be names, all terms need to have zero or one parents, homonyms just can't exist, etc. - but if you can (or desperately want to!) live within those constraints then this should be a lot less work, and the cleaner data is sort of a side effect.
@sharpphyl if you need help creating the new source, just let me know.
The "seed" link from taxon details is now "download," and there's a new tool which pushes to the classification bulkloader (from which data can be downloaded, and eventually uploaded to the revised hierarchy tool).
@sharpphyl if you need help creating the new source, just let me know.
@Jegelewicz Could we put this on the agenda during or after the next Taxonomy Committee meeting? Thanks.
new source
I'm not sure there's anything to discuss, although I'm happy to do so if it's helpful. If you want to manage a chunk of data hierarchically, then we'll create a source in which to do so.
Using the classification bulkloader to push to the hierarchy editor isn't going to work, so there will be a new bulkloader (which is about half of the classification bulkloader). It shouldn't be too difficult to munge hierarchical data from any source into it.
Pushing to the classification bulkloader works fine, so setting this thing up to autoload back to "core" Arctos should work fine.
I have scripts to pull from Arctos taxonomy into the classification bulkloader, the predictably-messy data means there will be a fair bit of initial cleanup, but the dedicated loader format means it's possible to do that through text files. After that I think managing hierarchically will be pretty easy.
The tool is about 80% resuscitated, but the remainder may take a fair bit of time. It's http://test.arctos.database.museum/tools/hierarchyEditor.cfm if you want a preview.
@sharpphyl @Jegelewicz I need advice.
The core editor is about as functional as I can get it without real data. The general idea for that is to have scripts, which will probably never have a UI, so that by your request and with your guidance I can start hierarchies from Arctos data. The obvious problem there is that Arctos data are absolute garbage....
For example I grabbed "Sorex c%" for some familiar taxonomy. The idea is to get "parent" from names, then insert "parents" as "names" and cycle all the way up the hierarchy. That works for some stuff, but fails for Sorex carolinensis which has no classification data (in test) and Sorex cinereus cincereus which has no "Arctos" classification data (also test) and etc. Those names will end up at the top of the hierarchy, with Kingdom. Not ideal, but maybe manageable....
Then I start working up the hierarchy and hit a hard end at Sorex. http://test.arctos.database.museum/name/Sorex#Arctos claims the next step is http://test.arctos.database.museum/name/Soricini#Arctos, which doesn't exist.
Everything I try ends up something like that. If there's a consistent genus in the Arctos classification, I can't find it.
I could go all the way up the hierarchy from a single name and then fill in the gaps as needed, but that would assign high-level parentage across the hierarchy from a mostly-random term, which could make a great deal of garbage that you might not notice is garbage until you've invested a fair bit of work into it. Doesn't seem like a great idea....
My inclination is that no data is easier to deal with than with garbage data, and therefore eg failing at genus is the least-evil thing I can do. In that case I'd need to manually add and organize all above-genus terms in the hierarchy (or in the template before I import it into the hierarchy), which for Sorex wouldn't take long at all. For "random inverts that aren't in worms" types of situations I don't think it's so trivial.
I think this will be a fairly rare occurrence - you'll want to find all the non-WoRMS taxa you use in IDs, or someone will want all North American shrews, or similar - and then those data will be managed in the hierarchy tool and it doesn't matter than they're a mess in "Arctos." In some cases those data probably won't come from Arctos at all, but from some spreadsheet or checklist or similar. I am therefore hesitant to invest more work in this unless you tell me to, but that also means I'm not testing with very representative data.
I pulled names (again in test) used by DMNS:Inv without an aphiaid as a more realistic test. You can play with it at http://test.arctos.database.museum/tools/hierarchyEditor.cfm?action=manage&hierarchy_id=4, and CSV from the editor is attached.
Is that a useful starting point? Is there a better way I can approach this? Should I not approach it at all, until someone has a well-defined use case?? Help!
loadedCfTempHierarchy(1).csv.zip

I'm messing around with it now and it will probably take me a bit to get around to any of your questions, but as I was editing, I had this happen. I picked taxon status in the term field accidentally, then changed it to aphiaID, but the dropdown from taxon status wouldn't go away, so I chose DELETE and even after a save, it is sitting there. I also had changed the source authority, but after save it still appears as WoRMS.
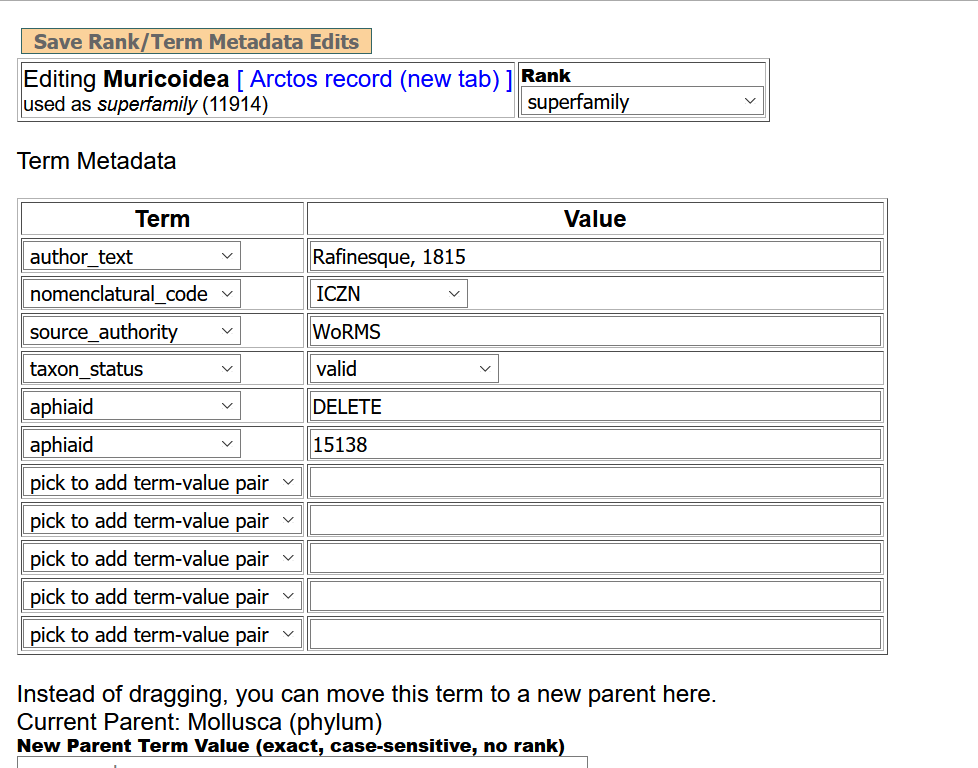
OK, so I played with the little 20 term thing from Arctos Plants, which was fairly easy to organize.
My inclination is that no data is easier to deal with than with garbage data, and therefore eg failing at genus is the least-evil thing I can do. In that case I'd need to manually add and organize all above-genus terms in the hierarchy (or in the template before I import it into the hierarchy), which for Sorex wouldn't take long at all. For "random inverts that aren't in worms" types of situations I don't think it's so trivial.
I think I understand what you are getting at here, but I am not 100% sure. I sort-of did this by adding the terms Arcida and Neogastropoda to fill in the hierarchy.
I need to understand how these test cases were created and why they appear in the order they do and what the numbers here mean

to help me figure out what is going on and what will happen when I import this hierarchy to Arctos Plants.
One observation - it would be nice to have a button to expand the whole thing rather than clicking a million times to see what is nested under stuff (if anything - frustrating to double-click and find there aren't any children...).
I'll keep messing around....
If it isn't already at the top, there is no way to move kingdom to the top?
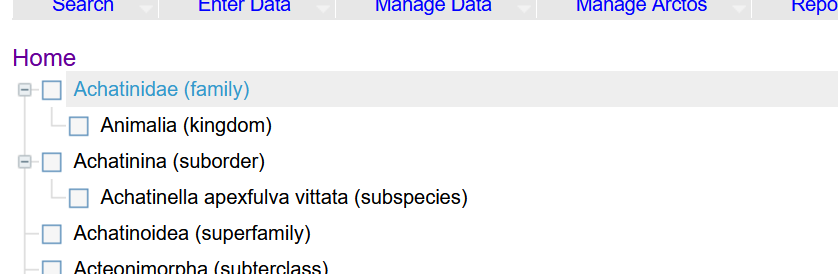
I don't know - this is super hard to work with. Dragging and dropping when the distance is more than a page is not possible without a two-handed click and scroll. Not being able to get the "top" of the hierarchy at the top makes it harder. Stuff collapses at certain saves and has to be clicked a million times to expand. It might be useful for a single genus or a small family, but more than that and it is clunky. In the end, there is going to be some term stuck at the top that I can't move to its appropriate location in the tree. I can delete and re-create, but that will probably lead to errors.
I'm done for the day, but I'll continue to think on it. I know this was a lot of work and it will probably help when moving a bunch of species from one family to another, otherwise, I am not sure about what it will be helpful for.
Is there anything different in the tool from the way it was before? It looks pretty similar. I can't really test it without having a complete group so that I have the higher taxonomy as well. I tried to create one just to see how it worked, and it doesn't, so feel free to delete my Chondrothyra (that should ultimately be an option). The names that don't have an aphiaID tend to be isolated ones. @dustymc could you create another set - perhaps all the Annulariidae (and its higher classification) just to play with.
A few ideas: It would be great to be able to add a taxon name from within the hierarchical tool even if it just started a csv with the data of where it's positioned and then you completed it later or even within the tool?
Will it be possible to download a csv without going through the tool? For a lot of what I've done, it was just as easy to work within the csv as within the tool, but you had to go through the tool to get the csv rather than a direct download from the taxonomic data.
I see in the download of the DMNSinv that many of the terms still didn't have ICZN in test (although I thought you added that at one time but maybe only in the tool), so just being able to update that from a csv is a big help.
I'll play more with it over the weekend and try to address your questions.
overlay is weird
On my list
what the numbers here mean
How the term is ranked in Arctos - eg, the core data are a huge inconsistent mess.
what will happen when I import this hierarchy to Arctos Plants.
It will replace all classifications for the names in your hierarchy. (Which will almost certainly increase overall entropy and get everybody yelling at everybody else, which is where https://github.com/ArctosDB/arctos/issues/3121 came from. Editing the giant messy "mega classifications" in this tool just isn't safe; it's aimed at smaller, cleaner, more consistent chunks of data.)
expand the whole thing
"Impossible" isn't the right word, but that would make eating your browser and servers something I'd have to be concerned about. I'm sure we can figure out something workable if we get that far.
Dragging and dropping when the distance is more than a page

get the "top" of the hierarchy a
I think I can let that accept NULL.
it is clunky.
Perhaps, but it's also structurally consistent.
The original idea here was to use external tools, but those never materialized. Maybe one of the many taxonomy projects has built something useful; find that and it should plug in....
useful for a single genus or a small family
Once the original import is dealt with, if it can be, that should be about what you see.
different in the tool from the way it was before?
Very little - I had to rebuild some under-the-hood stuff to deal with PG, I cleaned some minor things up, changed some structure to deal with things that have been strange in the past, but the UI should basically be the same.
delete ... should ultimately be an option).
Vague plan is that it won't be, for the same reasons you can't delete an entire Source from the main UI. This should be a primary tool holding primary data. There should be some sort of one-time perhaps-painful process to make a hierarchy, from Arctos or elsewhere, then all edits to that Source should happen in this tool. We'll see how that works out - lots of details I haven't quite resolved yes - and I can of course delete stuff along the way.
csv
Using Excel as your primary UI is a realistic option. Two obvious pathways:
- structure your data for the classification loader, use it when you want to
- structure your data for the hierarchical editor, use it to transform into the classification loader. I could even pull that part out as a stand-alone tool pretty easily.
ICZN
The tool writes to the classification loader. You can export, download, change, re-upload, which probably isn't entirely unrealistic for something that should happen about once.
@dustymc As I recall, I was able to create a csv from multiple points in the previous editor. For example, if I wanted to just update a genus, I could open the genus entry, and click on export. I don't see that feature in here perhaps because it's an incomplete list but it was a helpful tool.
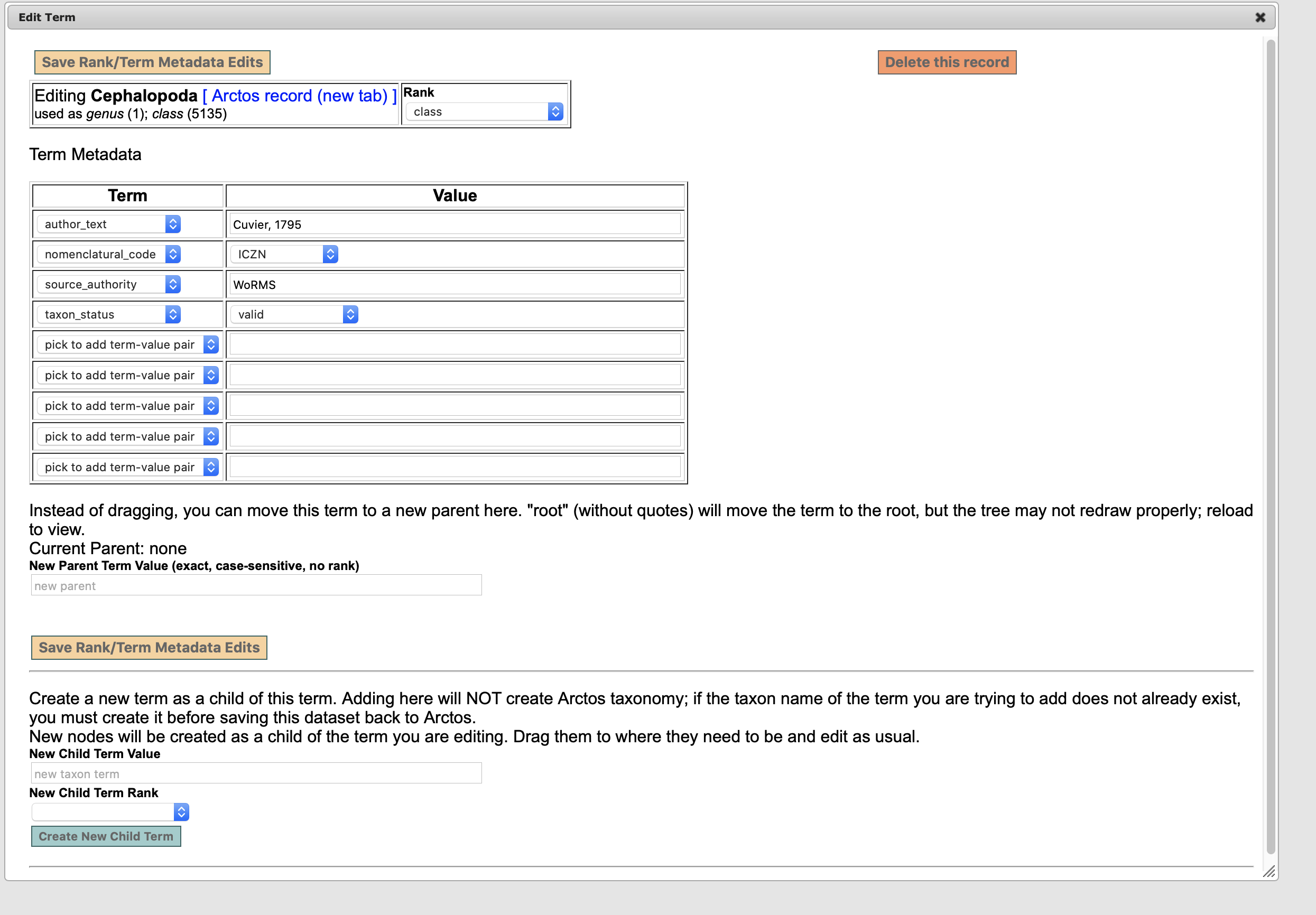
Since we have a Taxonomy Committee meeting tomorrow, I'd like to have @Jegelewicz or you walk me through the changes from the old tool and how this would work. Overall, I think it's somewhat more streamlined but would love to tweak it a bit more.
Again, is there any reason to include WoRMS (via Arctos) and any other externally managed source in the Export Targets since we're trying to not have any taxon names in that source that don't have an aphiaID, so they can't be permanently managed internally.

I do have to add taxon names that came into WoRMS within the past year and for whatever reason haven't made it to WoRMS (via Arctos) yet, but when I do the aphiaID creates everything. I think, even if I bulkloaded the name and aphiaID, I would still have to open the entry to refresh it, so I just do each one manually.
create a csv from multiple points
I could revive that, but it looks like 'complexity worth avoiding' to me - I think a simple "save all" can work now, so why not? (Maybe lots of good reasons, I don't see them yet!)
is there any reason to include WoRMS (via Arctos) and any other externally managed source in the Export Targets
There are compelling reasons to disallow that, which is what https://github.com/ArctosDB/arctos/issues/3121 would do.
have an aphiaID
Those could easily be bulkloaded via spreadsheet. I would advocate for allowing only bulkloading if I were managing those - it would be a few more clicks to add one or two, but I'd sleep better knowing that nobody could delete (or change to kelp) my aphiaids for no particular reason....
I would still have to open the entry to refresh it,
Nope, they should catch up by themselves.
tomorrow
I'll try to show up. Short version as I imagine it - but note that my imagination doesn't always line up with reality! - below.
- import some data of particular importance - cricetid rodents because that's what you work on, or North American woodpeckers because your Curator can't stand what's been done in some other otherwise-nice Source, or possibly 'stuff that isn't marine-y enough for WoRMS' (although that might be by nature inconsistent enough to make a compelling case for Excel as a UI...) "Birds" should be workable, but that's not really what this tool is aimed at. The bulkloader should now handle any checklist you might find, and the tiered preference model lets you use the bits of it you want, so there's not much reason to "manage" (beyond downloading/munging/uploading) that at all, much less in a hierarchy. I fully expect this step to be a giant pain in the butt from about any source, call it the price of admission.
- Drag/drop, get your friendly local DBA to add a "nomenclatural_code" tag to everything, maybe occasionally add a new name and have to separately bulkload it to Arctos. In general, easily do consistent things to this data you've brought into a consistent form.
- Occasionally click the 'export' tool (which after some testing/usage could be wired to 'autoload'), find your nice consistent data in the core DB after a couple days, suddenly find yourself getting much better answers to taxonomic questions from your specimens.
Stuff collapses at certain saves
That's a relic of having multiple ways of manipulating the hierarchy; I've got the 'move to parent' build into the 'tags' form, so I have to assume the tree's been manipulated and refresh it on save. I'll split that out, which will make the code a bit easier to deal with and only require refreshing the tree when the 'reparent' option is used.
This is functional and will be in next release. We can deal with any remaining issues as necessary, if anyone chooses to use this tool.
See also https://handbook.arctosdb.org/documentation/taxonomy.html#taxon-classification-sources - this development has lead to the idea (and tools) that managing hierarchically (or otherwise) does not necessarily involve using Arctos proper.
The "download" option (with every "local" classification term on every /name/... page) is not (yet, perhaps) directly capable of feeding the hierarchical editor, but does seem useful in pulling data in a generic flat format that could be fed into about any tool with which one might wish to manage taxonomy.
Most helpful comment
Worthy goal; yay consistency!
... where they have a non-zero chance of being turned into dinosaurs, or at least made inconsistent by individual updates.
Then...
Not really, but there's nothing very new in here either - it's all old tools, doing new things under a slightly different model.
It should be. Finding all of your stuff should be easy; it's in one Source. Moving a genus to a new family is one drag-drop and all of the species and subspecies can't avoid following. Adding/changing subkingdom involves one edit, not thousands. Etc. There are limitations to a hierarchy - all terms need to also be names, all terms need to have zero or one parents, homonyms just can't exist, etc. - but if you can (or desperately want to!) live within those constraints then this should be a lot less work, and the cleaner data is sort of a side effect.