Arctos: Locality Attribute - separate biostratigraphy and biochronology
Goal
Biostratigraphic zone and biochronology ended up in the same table and they were supposed to be separate - I missed this while they were in test. They need to be separated.
Context
biochronology is distinct from biostratigraphy - they are defined differently, used differently, and have different units.
Table
https://arctos.database.museum/info/ctDocumentation.cfm?table=ctbiostratigraphic_zone
Value
New Table ctbiochron
Definition
https://en.wikipedia.org/wiki/Biochron
Attribute data type
categorical
edit: fixed the links
All 42 comments
Here are the values that need to be moved over to biochron:
Adamanian
Apachean
Aquilian
Arikareean
Barstovian
Blancan
Bridgerian
Cashenranchian
Chadronian
Clarendonian
Cobrean
Comobluffian
Dawan
Duchesnean
Early Barstovian
Early Hemphillian
Fencelakean
Hemingfordian
Hemphillian
Irdinmanhan
Irvingtonian
Judithian
Kirtlandian
Lancian
Late Barstovian
Lootsbergian
Lysitean
Orellan
Otischalkian
Perovkan
Pu1
Pu2
Pu3
Puercan
Rancholabrean
Revueltian
Sanfordian
Sharamurunian
Tiffanian
To1
To2
To3
Torrejonian
Uintan
Wasatchian
Wassonian
Whitneyan
When we move them over we should make the following alterations to help in searching:
Early Barstovian -> Barstovian, Early
Early Hemphillian -> Hemphillian, Early
Late Barstovian -> Barstovian, Late
Pu1 -> Puercan, Pu1
Pu2 -> Puercan, Pu2
Pu3 -> Puercan, Pu3
To1 -> Torrejonian, To1
To2 -> Torrejonian, To2
To3 -> Torrejonian, To3
Man - I thought I was going crazy! As I was working on the documentation, I was like "What happened to Biochronology?"!!!
BTW - take a look at the updated documentation and let me know what looks crazy.
test
Let's blame https://github.com/ArctosDB/internal/issues/65 - I think there's no data in test so easy to miss.
We should match https://arctos.database.museum/info/ctDocumentation.cfm?table=ctbiostratigraphic_zone as close as possible.
ctbiochronology?? (no zone ???)
ctbiochronology is fine too. The units are called biochrons.
@Nicole-Ridgwell-NMMNHS one more question - does Precambrian belong in informal chronostrat? Seems like is should be in Eon/Eonothem?
The units are called biochrons.
Oh - you pick then!
ctbiochron-->biochron?
Lets do ctbiochronology and the table will be called biochronology.
@Jegelewicz I'm still looking into your question about the Cambrian, but from what I can tell so far, the United States Geological Survey considers it informal, but ICS still uses it and considers it a "supereon" the latter I just got from an un-cited statement in wikipedia though, so I need to track down the actual published statement on it.
Lets do ctbiochronology and the table will be called biochronology.
Agree
CT is built; yay postgres! You'll need to
- add the attribute
- add code-table control to use it
- populate the new CT
I think I can probably script some of that - just give it to me in attention-span-sized (small!) chunks of CSV....
/remind me to create this csv tomorrow
@Jegelewicz set a reminder for Aug 5th 2020
:wave: @Jegelewicz, create this csv
does Precambrian belong in informal chronostrat?
I should have just started my research by taking a look at the fine print. The chart itself says: "Italic fonts indicate informal units" so yes, we should move Precambrian back over to informal.
Precambrian is in informal now.
biochron added to locality attribute type code table
ctbiochronology code table populated
code table control added
@Nicole-Ridgwell-NMMNHS can you review the ctbiochronology code table to make sure the terms are correct?
Once that is done, we can migrate stuff that has these terms under bistratigraphy and remove the terms from that code table.
All terms are correct, but we should adjust the following:
Add Clarkforkian https://en.wikipedia.org/wiki/Clarkforkian
Documentation for Cashenranchian https://econtent.unm.edu/digital/collection/bulletins/id/469
Documentation for Cobrean https://econtent.unm.edu/digital/collection/bulletins/id/765
Documentation for Comobluffian https://vjs.pgi.gov.pl/article/view/26526
Documentation for Dawan https://econtent.unm.edu/digital/collection/bulletins/id/467
Documentation for Fencelakean https://econtent.unm.edu/digital/collection/bulletins/id/4589
Documentation for Kirtlandian https://econtent.unm.edu/digital/collection/bulletins/id/663
Documentation for Lootsbergian https://econtent.unm.edu/digital/collection/bulletins/id/301
Documentation for Perovkan https://doi.org/10.1016/S0031-0182(98)00117-5
Documentation for Sanfordian https://econtent.unm.edu/digital/collection/bulletins/id/7196
Documentation for Wassonian https://econtent.unm.edu/digital/collection/bulletins/id/467
Above completed.
@dustymc here is what we need to do now.
Find all localities with locality attribute type biostratigraphic zone = Adamanian
add to them locality attribute type biochron = Adamanian
delete from them locality attribute type biostratigraphic zone = Adamanian
delete from ctbiostratigraphic zone the term Adamanian
repeat for all terms in ctbiochronology
How do you suggest I proceed to get this done as efficiently as possible?
efficiently
Ain't no such thing in authority-land...
can I just lock down and
update locality_attributes set attribute_type='biochron' where attribute_type='biostratigraphic zone' and attribute_value='Adamanian';
and then clean up what's left?
Heck yeah!
UPDATE 568
@dustymc we still need to
repeat for all terms in ctbiochronology
from https://github.com/ArctosDB/arctos/issues/2987#issuecomment-675741576
Terms are
Aquilian
Arikareean
Barstovian
Barstovian, Early
Barstovian, Late
Blancan
Bridgerian
Cashenranchian
Chadronian
Clarendonian
Clarkforkian
Cobrean
Comobluffian
Dawan
Duchesnean
Fencelakean
Hemingfordian
Hemphillian
Hemphillian, Early
Irdinmanhan
Irvingtonian
Judithian
Kirtlandian
Lancian
Lootsbergian
Lysitean
Orellan
Otischalkian
Perovkan
Puercan
Puercan, Pu1
Puercan, Pu2
Puercan, Pu3
Rancholabrean
Revueltian
Sanfordian
Sharamurunian
Tiffanian
Torrejonian
Torrejonian, To1
Torrejonian, To2
Torrejonian, To3
Uintan
Wasatchian
Wassonian
Whitneyan
Alternatively, you could write me some SQL to get me all locality names with an associated biostratigraphy term so that I can find them and bulkload the new locality attribute. Then I'll need a locality attribute unloader to get rid of the incorrect ones.
Your choice!
repeat for all terms in ctbiochronology
I think I'm lost, and it seems like there's going to be some juggling still to do, so....
SQL
locality.locality_name
from
locality
inner join locality_attributes on locality.locality_id=locality_attributes.locality_id
where
locality_attributes.attribute_type='biostratigraphic zone'
-- just don't add the next line for all
and locality_attributes.attribute_value='Baculites clinolobatus'
group by
locality.locality_name
;
locality attribute unloader
We probably need to talk about how that works, and it'll need prioritized, or if can define the SQL to get locality_attribute_id I could easily use that to unload.
OK - I'll be in touch when I'm ready for the unloads.
/remind me to do this tomorrow
@Jegelewicz set a reminder for Aug 25th 2020
:wave: @Jegelewicz, do this
arctosprod@arctos>> \d locality_attributes
Table "public.locality_attributes"
Column | Type | Collation | Nullable | Default
------------------------+-----------------------+-----------+----------+--------------------------------------------------------------------
locality_attribute_id | integer | | not null | nextval('locality_attributes_locality_attribute_id_seq'::regclass)
locality_id | bigint | | not null |
determined_by_agent_id | bigint | | |
attribute_type | character varying(60) | | not null |
attribute_value | character varying | | not null |
attribute_units | character varying(60) | | |
attribute_remark | character varying | | |
determination_method | character varying | | |
determined_date | character varying(22) | | |
Indexes:
"ix_u_locality_attributes_id" UNIQUE, btree (locality_attribute_id)
"ix_locality_attributes_attribute_type" btree (attribute_type)
"ix_locality_attributes_attribute_value" btree (attribute_value)
Check constraints:
"ck_attribute_remark_noprint" CHECK (checkfreetext(attribute_remark))
"ck_attribute_value_noprint" CHECK (checkfreetext(attribute_value))
"ck_determination_method_noprint" CHECK (checkfreetext(determination_method))
"ck_determined_date_iso8601" CHECK (ck_iso8601(determined_date::text))
Foreign-key constraints:
"locality_attributes_attribute_type_fkey" FOREIGN KEY (attribute_type) REFERENCES ctlocality_attribute_type(attribute_type)
"locality_attributes_determined_by_agent_id_fkey" FOREIGN KEY (determined_by_agent_id) REFERENCES agent(agent_id)
"locality_attributes_locality_id_fkey" FOREIGN KEY (locality_id) REFERENCES locality(locality_id)
Triggers:
trg_locality_attribute_archive AFTER INSERT OR DELETE OR UPDATE ON locality_attributes FOR EACH ROW EXECUTE FUNCTION trigger_fct_trg_locality_attribute_archive()
https://github.com/ArctosDB/PG_DDL/blob/master/function/getpreferredagentname.sql will be handy if you want names.
In case I need it, used this to get current, incorrect biostratigraphy
select
locality.locality_name,
locality_attributes.attribute_value,
locality_attributes.determined_by_agent_id,
locality_attributes.determined_date,
locality_attributes.attribute_remark
from
locality
inner join locality_attributes on locality.locality_id=locality_attributes.locality_id
where
locality_attributes.attribute_type='biostratigraphic zone'
-- just don't add the next line for all
and locality_attributes.attribute_value='To3'
or locality_attributes.attribute_value='To1'
or locality_attributes.attribute_value='To2'
or locality_attributes.attribute_value='Aquilian'
or locality_attributes.attribute_value='Arikareean'
or locality_attributes.attribute_value='Barstovian'
or locality_attributes.attribute_value='Early Barstovian'
or locality_attributes.attribute_value='Late Barstovian'
or locality_attributes.attribute_value='Blancan'
or locality_attributes.attribute_value='Bridgerian'
or locality_attributes.attribute_value='Cashenranchian'
or locality_attributes.attribute_value='Chadronian'
or locality_attributes.attribute_value='Clarendonian'
or locality_attributes.attribute_value='Clarkforkian'
or locality_attributes.attribute_value='Cobrean'
or locality_attributes.attribute_value='Comobluffian'
or locality_attributes.attribute_value='Dawan'
or locality_attributes.attribute_value='Duchesnean'
or locality_attributes.attribute_value='Fencelakean'
or locality_attributes.attribute_value='Hemingfordian'
or locality_attributes.attribute_value='Hemphillian'
or locality_attributes.attribute_value='Early Hemphillian'
or locality_attributes.attribute_value='Irdinmanhan'
or locality_attributes.attribute_value='Irvingtonian'
or locality_attributes.attribute_value='Judithian'
or locality_attributes.attribute_value='Kirtlandian'
or locality_attributes.attribute_value='Lancian'
or locality_attributes.attribute_value='Lootsbergian'
or locality_attributes.attribute_value='Lysitean'
or locality_attributes.attribute_value='Orellan'
or locality_attributes.attribute_value='Otischalkian'
or locality_attributes.attribute_value='Perovkan'
or locality_attributes.attribute_value='Puercan'
or locality_attributes.attribute_value='Pu1'
or locality_attributes.attribute_value='Pu2'
or locality_attributes.attribute_value='Pu3'
or locality_attributes.attribute_value='Rancholabrean'
or locality_attributes.attribute_value='Revueltian'
or locality_attributes.attribute_value='Sanfordian'
or locality_attributes.attribute_value='Sharamurunian'
or locality_attributes.attribute_value='Tiffanian'
or locality_attributes.attribute_value='Torrejonian'
or locality_attributes.attribute_value='Uintan'
or locality_attributes.attribute_value='Wasatchian'
or locality_attributes.attribute_value='Wassonian'
or locality_attributes.attribute_value='Whitneyan'
FYI check out "in"
and locality_attributes.attribute_value in (
'To3',
'To1',
...
)
https://arctos.database.museum/tools/bulkloadLocalityAttributes.cfm seems to be loading super slow - of course I made the mistake of not ISO-ing my dates so I'm going to have a few errors, but still I set 2500 to autoload earlier this morning and so far only about 20 have processed.
super slow
It's slightly less slow now.
Biochrons have been added.
@dustymc all locality attributes in this file need to be DELETED. Let me know if you have questions.
Let's go to #2967
Have unloaded all incorrect biostratigraphy, need to remove values from the code table.
- [x] Adamanian
- [x] Apachean
- [x] Aquilian
- [x] Arikareean
- [x] Barstovian
- [x] Blancan
- [x] Bridgerian
- [x] Cashenranchian
- [x] Chadronian
- [x] Clarendonian
- [x] Cobrean
- [x] Comobluffian
- [x] Dawan
- [x] Duchesnean
- [x] Early Barstovian
- [x] Early Hemphillian
- [x] Fencelakean
- [x] Hemingfordian
- [x] Hemphillian
- [x] Irdinmanhan
- [x] Irvingtonian
- [x] Judithian
- [x] Kirtlandian
- [x] Lancian
- [x] Late Barstovian
- [x] Lootsbergian
- [x] Lysitean
- [x] Orellan
- [x] Otischalkian
- [x] Perovkan
- [x] Pu1
- [x] Pu2
- [x] Pu3
- [x] Puercan
- [x] Rancholabrean
- [x] Revueltian
- [x] Sanfordian
- [x] Sharamurunian
- [x] Tiffanian
- [x] To1
- [x] To2
- [x] To3
- [x] Torrejonian
- [x] Uintan
- [x] Wasatchian
- [x] Wassonian
- [x] Whitneyan
UGH - need to re-run for remaining bad usage.
UGH - need to re-run for remaining bad usage.
What is left?
@dustymc something is strange here.
Ran this SQL
locality
inner join locality_attributes on locality.locality_id=locality_attributes.locality_id
where
locality_attributes.attribute_type='biostratigraphic zone'
-- just don't add the next line for all
and locality_attributes.attribute_value='Blancan'
or locality_attributes.attribute_value='Puercan'
or locality_attributes.attribute_value='Rancholabrean'
or locality_attributes.attribute_value='Revueltian'
And got a bunch of results like this
LOCALITY_NAME | ATTRIBUTE_VALUE | DETERMINED_BY_AGENT_ID | DETERMINED_DATE | ATTRIBUTE_REMARK
-- | -- | -- | -- | --
NMMNH:Paleo:L-317 | Puercan | 21301807 | 1977-04-16 |
NMMNH:Paleo:L-12346 | Puercan | 21319446 | |
NMMNH:Paleo:L-12346_public | Puercan | 21319446 | |
NMMNH:Paleo:L-72_public | Revueltian | 21319446 | 1987-01-01 |
NMMNH:Paleo:L-1 | Revueltian | | |
NMMNH:Paleo:L-1_public | Revueltian | | |
BUT when I go look at locality NMMNH:Paleo:L-317, it DOES NOT have that attribute.
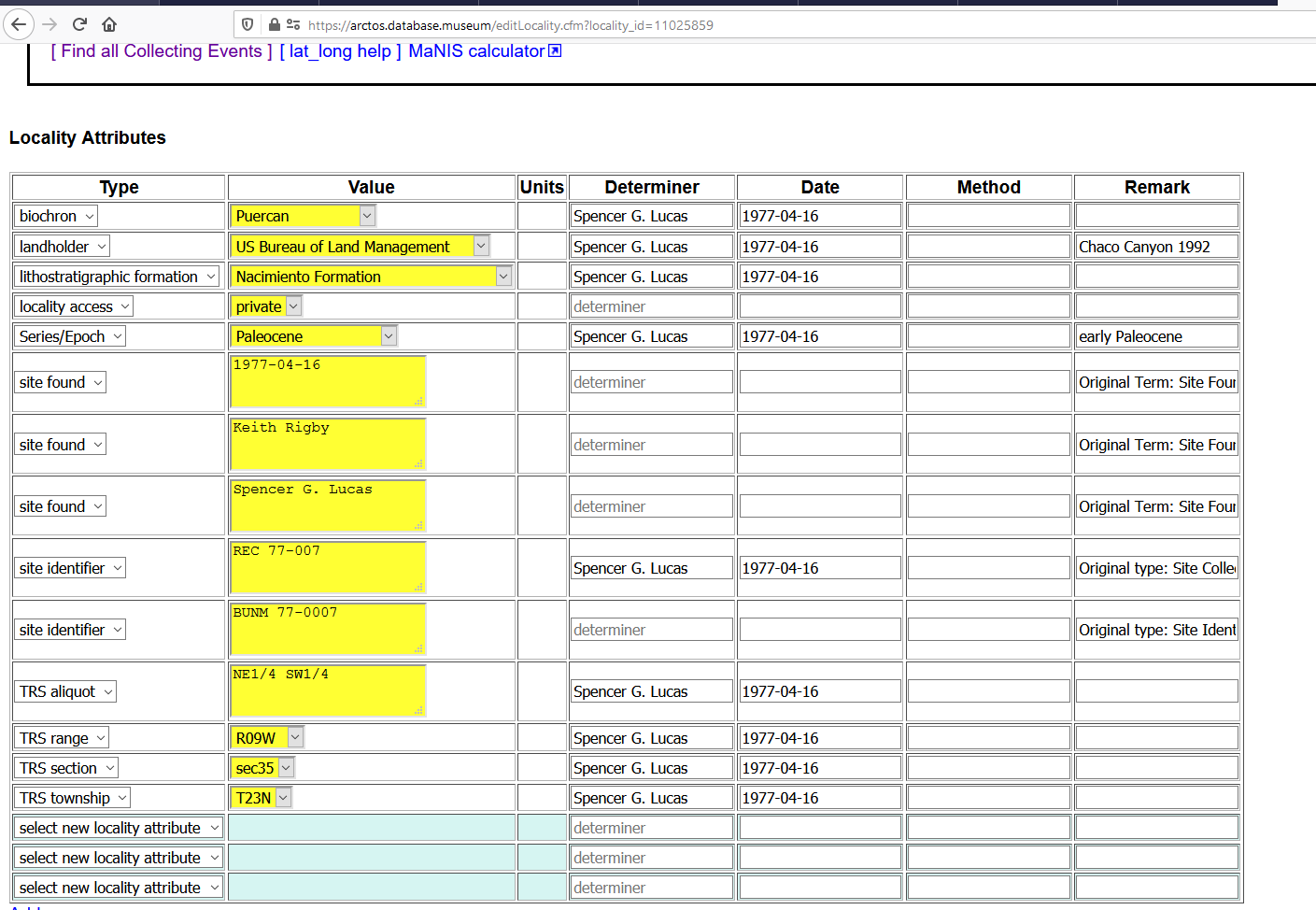
I am unable to remove the unchecked items in https://github.com/ArctosDB/arctos/issues/2987#issuecomment-693517925 because the are "in use", but I don't think they actually are.....
and ( locality_attributes.attribute_value='Blancan'
or locality_attributes.attribute_value='Puercan'
or locality_attributes.attribute_value='Rancholabrean'
or locality_attributes.attribute_value='Revueltian'
)
maybe...
locality.locality_name,
locality_attributes.attribute_value,
locality_attributes.determined_by_agent_id,
locality_attributes.determined_date,
locality_attributes.attribute_remark
from
locality
inner join locality_attributes on locality.locality_id=locality_attributes.locality_id
where
locality_attributes.attribute_type='biostratigraphic zone'
-- just don't add the next line for all
and locality_attributes.attribute_value IN ('Blancan',
'Puercan',
'Rancholabrean',
'Revueltian')
Did the trick.
Gonna take me a bit to clean these up - my bad as they are ALMNH stuff.
All cleaned up!