Issue Documentation is http://handbook.arctosdb.org/how_to/How-to-Use-Issues-in-Arctos.html
Is your feature request related to a problem? Please describe.
We have been working with organisms for which we have multiple occurrences, specifically Mexican Wolves in the Mexican Wolf recovery program. Throughout their lives, samples of blood are taken from these animals and deposited in the genomic resources collection at MSB. Traditionally, each set of samples (all from the same day) have been given a single catalog number. This results in multiple cataloged items for a single organism, which we can link to each other using the “same individual as” relationship.

These relationships are nice, but they don't allow us to see ALL events for an individual in one place. and they require the addition of a new relationship for ALL related cataloged items every time a new collection of blood is made. Each cataloged item includes the other ID “Mexican Wolf Studbook Number” and we have modified the Other ID url so that clicking this other ID allows us to find all of the samples from any given animal.

This method works, but there is one issue we need to address.
When our data leaves Arctos and is ingested by aggregators such as GBIF and iDigBio, there is no easy way for anyone using the data there to make the connection that the various cataloged items are all from the same animal. Although the Mexican Wolf Studbook numbers are included in the list of related IDs, the connection just isn’t as tight as we would like it to be.


Describe the solution you'd like
Our proposed solution is to make use of the Darwin Core field “Organism ID”. We envision this as a separate and distinct other ID – one which provides a link to all related specimens (the results of that link would look just like the search result you see when you search one of the Mexican Wolf Studbook numbers):

This identifier would be passed to aggregators in the “Organism ID” field – allowing those using the data there to make the appropriate connection between the related cataloged items. Currently it appears that we are just passing the catalog item to that field

which is what led to the solution we have been attempting to make work in https://github.com/ArctosDB/arctos/issues/1545. This has created problems with data entry and maintenance on our end. This new solution will allow us to keep events matched with parts and parts matched with accessions. It will simplify data entry and end the need for the links between events and parts.
We envision a new code table: CTCOLL_ORGANISM_ID set up very much like CTCOLL_OTHER_ID_TYPE where:
IDType = text “Mexican Wolf Studbook Number”
Description = definition of the IDType Studbook number assigned by the Mexican Wolf Recovery Program
BaseURI = http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=
When the Organism ID is used, there would be no need for all of the “same organism as” relationships, but they could be used if a collection so desired. Every cataloged item that included an Organism ID would instead appear like this:

With the text “Mexican Wolf Studbook Number: 1216” being a link taking you to the search results:

We would hope that this link could also be what appears at the aggregators in their “Organism ID” field:

Describe alternatives you've considered
The major challenge we see with this method is how to assign unique Organism IDs for things where there isn’t an obvious one. The Mexican Wolves (and eventually the Red Wolves that are expected to come in from Arkansas) and NEON recaptures are examples of when we would be using this method. These all have obvious unique identifiers (studbook numbers and NEON sample ID numbers). However, when the skin and skeleton of an animal are at DMNS and the tissues for that same animal are at MSB, there is no obvious organism ID type and we would need to come up with one. We are open to suggestions for how best to accomplish this.
What have we missed?
Additional context
See above
Priority
I would like to have this resolved by date: soonish
All 298 comments
I have passed this by John Wieczorek and here is our discussion:
The proposal to use dwc:organismID in Darwin Core resource is right on target. That is exactly what the field is meant for. You are right that Arctos is passing the id for the cataloged item in that field right now. The reasoning was based on the majority of cases, where the cataloged item corresponds to an Organism. Rigorously speaking, I think this is a mistake, because cataloged item does not always correspond to an Organism, and in Arctos, we don't have a fail-proof method of a knowing when it does, and when it doesn't. Given that, I think we should unmap organismID from the cataloged item in all Arctos resources.
I have looked at the proposal for the new code table (CTCOLL_ORGANISM_ID). I think this is unnecessary and unsustainable. I think a sufficient solution, which is also the most scalable, is to add a new type in CTCOLL_OTHER_ID_TYPE, called "organism identifier" or similar. Curators would have the freedom to create a (single) organism identifier, and that should be a persistent resolvable GUID. It could refer to any organism within Arctos, or outside it. Note that in the case of the Mexican Wolf Studbook Number, there would be two entries in the COLL_OTHER_ID table for each cataloged item - one with type "Mexican Wolf Studbook Number", which holds the number, and one with type "organism identifier" with the resolvable GUID to the organism.
There will be issues of "persistence" and of primacy (if two data publishers have distinct organismIDs, which should be used?), but those will exist outside of the scope of the immediate problem anyway. It's something that could conceivably be solved at a level above the publication of primary occurrence data.
Following what I am proposing above, there would be no need to communicate anything to GBIF, iDigBio, or GGBN. We would be following the intended use of dwc:organismID. The misunderstandings from iDigBio and GGBN are around the conflation of Occurrences by Arctos, not about the concept of Organism. The proposed solutions do not save us with respect to GBN either. With them the issue is that they want records of tissue samples, while everyone else in the world expects Occurrences, and these are not always the same thing, especially in Arctos. So, we still have to make distinct resources for GGBN, unfortunately.
My response:
I'm not sure I can wrap my brain around the other ID type solution. I feel like what you describe is what we do with the Mexican Wolves now - how would the GUIDs be created and where would they "live"?
I'm not the most technical person, so without a demo, it's just hard for me to see how two independent Other IDs will resolve to a GUID somewhere...but the idea seems the same as what I proposed just technically more stable? If so, I am on board and I agree that we need to stop sending catalog number as Organism ID AND that MSB needs to stop trying to catalog all collections for a single wolf in a single catalog number - which is why I proposed the solution I did - it is just too messy and information is lost in the process.
This is coming to the forefront for other reasons: https://github.com/tdwg/dwc-qa/issues/131
I'd like to create a simple solution to the organism issue - it really shouldn't be that difficult within Arctos. The problem of everyone agreeing on an ID when you consider stuff outside of Arctos is something we need to tackle as a larger community and is related to unique identifiers in general. Let me know how I can help push a solution forward and I'll do everything I can!
John responds:
I'm not sure I can wrap my brain around the other ID type solution. I feel like what you describe is what we do with the Mexican Wolves now - how would the GUIDs be created and where would they "live"?In Arctos the GUIDs would live in the Coll_Obj_Other_ID_Num table with an OTHER_ID_TYPE of "organism identifier". Curators would be responsible for entering these (read "danger").
I'm not the most technical person, so without a demo, it's just hard for me to see how two independent Other IDs will resolve to a GUID somewhere...but the idea seems the same as what I proposed just technically more stable? If so, I am on board and I agree that we need to stop sending catalog number as Organism ID AND that MSB needs to stop trying to catalog all collections for a single wolf in a single catalog number - which is why I proposed the solution I did - it is just too messy and information is lost in the process.
Two independent Other IDs do not resolve to a GUID somewhere. One of the IDs says "I am this Mexican Wolf Sudbook Number", the other says, "my dwc:orgnismID is this". Hey, maybe that's what to put in the CTOTHER_ID_TYPE table - "dwc:organismID" - it would be quite explicit.
This is coming to the forefront for other reasons: https://github.com/tdwg/dwc-qa/issues/131
I'd like to create a simple solution to the organism issue - it really shouldn't be that difficult within Arctos. The problem of everyone agreeing on an ID when you consider stuff outside of Arctos is something we need to tackle as a larger community and is related to unique identifiers in general. Let me know how I can help push a solution forward and I'll do everything I can!True. It is a community issue. Arctos is a great resource for pushing the limits of what we are able to do. For many outside it is way too far ahead, despite the fact that for some inside it doesn't do all we might want.
From me:
In Arctos the GUIDs would live in the Coll_Obj_Other_ID_Num table with an OTHER_ID_TYPE of "organism identifier". Curators would be responsible for entering these (read "danger").The "danger"is what I was hoping to avoid with the separate table for organism ID - using "Mexican Wolf Studbook Number" as the base of the ID means we don't get "Mexican wolf studbook number 1216", "Mex Wolf Studbook No. 1216", etc.
Two independent Other IDs do not resolve to a GUID somewhere. One of the IDs says "I am this Mexican Wolf Sudbook Number", the other says, "my dwc:orgnismID is this". Hey, maybe that's what to put in the CTOTHER_ID_TYPE table - "dwc:organismID" - it would be quite explicit.
To be clear - I don't propose there be two IDs, but to MOVE those other IDs that are truly Organism IDs to the new table.
In general, I think having some sort of "individual ID" would be very useful. It's not at all clear to me why it would be in a separate table; that invites more denormalization (doing the same thing multiple ways), inevitably leading to even bigger messes.
If the scope of this is Arctos, we could exploit relationships to assemble "individuals" and/or individualID without adding any overhead - there's much more discussion on that in https://github.com/ArctosDB/arctos/issues/1545 - and see below.
I believe that this is implicitly a proposal to recatalog http://arctos.database.museum/guid/MSB:Mamm:292063 as 5 specimens. At least for some use cases that goes against the "catalog the item of scientific interest" mantra; eventually two of the samples from the same wolf will be compared in a publication. I'm not sure that's more evil than the current situation, where 5 samples collected at different times under different conditions are likely seen as equivalent to 5 tubes from the same liver of another specimen, but it should be acknowledged. I think any consistent documented approach is an improvement.
"Occurrences" are occasionally recorded in different collections, both in and out of Arctos, so cataloging Occurrences rather than individuals would make Arctos data more comparable with the rest of the world. I'm not sure how much weight that should carry, but again it is a consideration that should be addressed.
All of that said, I don't think Arctos can or should dictate how material is cataloged. I think the most we can do is to provide documentation/guidance.
This should extend beyond Arctos. A sample of http://arctos.database.museum/guid/MSB:Mamm:292063 stored in another system and shared with GBIF would ideally bear the same "individual ID" as the record(s) in Arctos. If it did, it would be trivial to assemble the individual in GBIF or similar systems.
The "danger" is in assigning the identifiers, and I don't believe there is any technical solution to that - it's a social problem that needs a social solution. It took seconds to find https://arctos.database.museum/guid/MSB:Mamm:317312 and https://arctos.database.museum/guid/MSB:Mamm:324187 which share a NEON ID and probably are not the same organism. I have never encountered a "number series" that didn't have similar issues, and if that exists the NEON ID cannot do what you want. I think this would be best implemented as GUIDs, and for social reasons those should probably not be minted by Arctos. Drawing those from an independent source would let Curators determine what is or is not an Individual on a case-by-case basis independent of any problems with identifiers assigned by other organizations, and at least maintains some possibility that other collections holding material from the same individuals would buy in and assign those IDs to their specimens. Two candidates are UUIDs, which would not be resolvable or actionable, or ARKs which could be resolvable and could point to some shared view (eg, GBIF, which in turn could point to the various bits and pieces of the individual in various systems/collections).
I think that also could be implemented only as guidance; I don't think Arctos can or should prevent someone from using "1" as an IndividualID, but we can help them understand the implications of doing so.
How would this not be denormalization?
organismID = Mexican Wolf Studbook Number 1216
organismID = Mex Wolf Studbook No 1216
organismID = Mexican wolf studbook number 1216
These are all the same organism, but now we have three IDs for it. If we have:
ORGANISM_ID where:
IDType = text “Mexican Wolf Studbook Number” Description = definition of the IDType Studbook number assigned by the Mexican Wolf Recovery Program BaseURI = http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=
At least we eliminate the problem of the many ways "Mexican Wolf Studbook Number" might be spelled.
I think this would be best implemented as GUIDs, and for social reasons those should probably not be minted by Arctos.
I agree with this statement - but no one is stepping up to the plate for biological specimens (at least no one I am aware of). While the solution above does not fix the problems of the world, it would be a start for Arctos collections and maybe we could use that to press the issue with the community.
I looked up ARKs and I'm not clear on how that works - if is a solution, then let's explore, but I need an example because it seems very fuzzy to me and doesn't solve the social problem as far as I can tell.
I believe that this is implicitly a proposal to recatalog http://arctos.database.museum/guid/MSB:Mamm:292063 as 5 specimens. At least for some use cases that goes against the "catalog the item of scientific interest" mantra; eventually two of the samples from the same wolf will be compared in a publication.
Yep - and the cataloging of separate events with one catalog number results in events and parts that are not properly associated with their accessions, their collectors and preparators, nor their attributes. (The event links are OK, but easily broken or incorrectly made).
Should OrganismIDs be a DOI?
I'm still not following. You want another table that's the same structure and does the same thing as OtherIDs??
And yes those data are denormalized - that's a lot easier to deal with that denormalized structure, and one of many reasons a GUID of some sort would be a useful value.
There is no technical solution to social problems. We can make it enticing to assign unifying IDs, but that's about it.
ARKs are functionally much like DOIs, but they're free (and don't come with the buy-in, which I suspect means they also don't come with the persistence).
https://n2t.net/ark:/87299/x6d50k1v
If I a couple million dollars and nothing better to do, everything in Arctos would have a DOI. DOIs would be great "individialIDs" but I don't think I can supply them. And that would lead back into the whole "controlled by Arctos" thing, which I don't think has any chance of being adopted by anyone outside of Arctos. I can provide tools, but the folks who own these specimens should also own the unifying identifiers.
I'm still not following. You want another table that's the same structure and does the same thing as OtherIDs??
EXCEPT - those IDs would be passed to GBIF and other aggregators as "Organism_ID".
I have also considered just using a check box in the Other_ID table "this is an organism ID"....
Thanks - I might actually get it now!
It's Arctos-centric and not very pretty, but at least it's not denormalization: http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none is a perfectly valid value for other_id_type=OrgID (whatever we call it).
That could be generated by a "this is an orgid" button. I could even abstract it to a saved search or ARK, but that gets us back to the "Arctos-centric" thing.
And again, if the scope of this is just "works for Arctos" then I think we'd be better off doing something with relationships. (@tucotuco pointed out that an ID works from a spreadsheet where a relationship may not, so "something" might be generating a URL that finds ID=value as above - IDK, that's details, I'm totally open to ideas).
" That could be generated by a "this is an orgid" button" - you mean in the
code table, correct?
Also, we would not want to see the "messy"
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none
in
the display. We'd want to see "Organism ID: Mexican Wolf Studbook Number:
1216".
possible?
On Wed, Mar 13, 2019 at 3:11 PM dustymc notifications@github.com wrote:
Thanks - I might actually get it now!
It's Arctos-centric and not very pretty, but at least it's not
denormalization:
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none
is a perfectly valid value for other_id_type=OrgID (whatever we call it).That could be generated by a "this is an orgid" button. I could even
abstract it to a saved search or ARK, but that gets us back to the
"Arctos-centric" thing.And again, if the scope of this is just "works for Arctos" then I think
we'd be better off doing something with relationships. (@tucotuco
https://github.com/tucotuco pointed out that an ID works from a
spreadsheet where a relationship may not, so "something" might be
generating a URL that finds ID=value as above - IDK, that's details, I'm
totally open to ideas).—
You are receiving this because you are subscribed to this thread.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472607050,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hCEHhRD5iBe6CGraaQvG4XAq94Duks5vWWl1gaJpZM4buGmY
.
No, in the interface.
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none is a GUID - and an actionable one at that. There's only one of them on the planet and it's easy to tell what it does. (It's not very pretty and may or may not be very persistent, but that's details.)
Mexican Wolf Studbook Number: 1216 is a string. Anyone can use it for any purpose anywhere; it doesn't natively do anything, and trying to do anything with it comes with a big pile of indefensible assumptions.
Edit for completeness: https://n2t.net/ark:/87299/x68g8hqw currently does the same thing as http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none. It's prettier and likely more persistent. If I find another Occurrence of "none" I could re-point the ARK to somewhere mutually agreeable (eg, GBIF) in order to build a more complete picture of the Organism. It's a MUCH better solution than the URL, but also likely to take more investment than clicking a button.
2nd edit: I'm throwing ARKs around only because they're not-Arctos and super easy to create. They're not the only possible GUID, just a convenient and functional example.
...not to mention that the indefensible assumptions would be distinct for
every different id type, ergo not scalable.
On Wed, Mar 13, 2019 at 6:49 PM dustymc notifications@github.com wrote:
No, in the interface.
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none
is a GUID - and an actionable one at that. There's only one of them on the
planet and it's easy to tell what it does. (It's not very pretty and may or
may not be very persistent, but that's details.)Mexican Wolf Studbook Number: 1216 is a string. Anyone can use it for any
purpose anywhere; it doesn't natively do anything, and trying to do
anything with it comes with a big pile of indefensible assumptions.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472619696,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AAcP68SCG5JP36cTtulJpcrF783lkU80ks5vWXJsgaJpZM4buGmY
.
Mexican Wolf Studbook Number: 1216 is a string. Anyone can use it for any purpose anywhere; it doesn't natively do anything, and trying to do anything with it comes with a big pile of indefensible assumptions.
I don't get how what you propose is different from:
IDType = text “Mexican Wolf Studbook Number”
Description = definition of the IDType Studbook number assigned by the Mexican Wolf Recovery Program
base URL = http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=
I had been thinking there would be only one allowed organismID. Maybe that is silly. Maybe it is fine to have as many as you like. That way you could include your own AND those of other collections (in or out of Arctos). That way you could also potentially go directly to GBIF to get the set of Occurrences for all matching organismIDs.
HMMMM..I hadn't considered that.
Maybe it is fine to have as many as you like. That way you could include your own AND those of other collections (in or out of Arctos). That way you could also potentially go directly to GBIF to get the set of Occurrences for all matching organismIDs.
BUT when searching AT GBIF, how would they be related - so that some person who was unaware the two organism IDs were the same organism could make the connection?
We were discussing earlier how we could link specimens at MSB and AMNH and
Collecion Boliviana de Fauna that are all part of the same animal. All
share the same field number, they are all the same organism, but how would
we relate them in GBIF if AMNH assigns one and MSB assigns a different one?
Ideally, we'd use the shared field number as the core ID, or we'd pay for a
doi.
On Wed, Mar 13, 2019 at 5:21 PM John Wieczorek notifications@github.com
wrote:
I had been thinking there would be only one allowed organismID. Maybe that
is silly. Maybe it is fine to have as many as you like. That way you could
include your own AND those of other collections (in or out of Arctos). That
way you could also potentially go directly to GBIF to get the set of
Occurrences for all matching organismIDs.—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472644191,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hEL5hhP7FTXoNDyscDnEluqXKJ11ks5vWYf3gaJpZM4buGmY
.
I don't get how what you propose is different from:
IDType = text “Mexican Wolf Studbook Number”
Description = definition of the IDType Studbook number assigned by the Mexican Wolf Recovery Program
base URL = http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=
It is very different outside the world of Arctos. The organismID would have to be constructed from this, and what would you do to create the organismIDs of the ten collections that have parts of the same plant? Create ten new ID types and base URLS (just to cover that one organism - multiply by all the collections that share any parts of any Organisms in Arctos)?
different
It eliminates data stored in arbitrary places.
only one
Yea, I suspect reality will find a way to stomp all over that, but it would be nice....
link specimens
Arctos can link to anything with a URL, and provides a mechanism for incoming links.
shared field number
Everybody starts at "1." If you want links, you need actionable GUIDs. If you want discoverable, you need shared actionable GUIDs. You might get at "shared" by tracking down the other 40 samples in GBIF and adding their IDs to Arctos, although "here's a nice neutral persistent actionable identifier, would you mind using it so we can talk to each other?" would greatly simplify things.
All share the same field number, they are all the same organism, but how would we relate them in GBIF if AMNH assigns one and MSB assigns a different one?
I think that is what I am getting at in https://github.com/tdwg/dwc-qa/issues/131#issuecomment-472642620
Something akin to IGSNs, but for Organisms instead of for samples.
The organismID would have to be constructed from this, and what would you do to create the organismIDs of the ten collections that have parts of the same plant? Create ten new ID types and base URLS (just to cover that one organism - multiply by all the collections that share any parts of any Organisms in Arctos)?
I don't understand - you would only need one ID type. From any record in Arctos, I can click the link from the Mexican Wolf Studbook Number (no matter what number it is) and I'll get the specimen results page that show all of the wolves that share the same number.
If UTEP or UMNH or any other Arctos collection had a wolf specimen and put the studbook number in the "Mexican Wolf Studbook Number" other ID, then it would show up in the search too, because the link is an actionable guid like Dusty described.
It would be a social issue to decide upon an "ID Type" for the situation that you describe, but we should only need one. The challenge - as I pointed out in the very beginning is assigning the individual organism ID numbers, so that all collections with parts of the same plant would use "Individual Plant ID" = 1, etc.
I guess I am missing something (which doesn't surprise me...) The wolves are easy because they are all here and they have a (somewhat) logical identifier. Everything else will be messy until we have a unique BOI (Biological Organism Identifier).
In all of these situations, there is a shared organism number already that
links specimens. Examples currently in use within Arctos and between Arctos
and outside collections (AMNH, USNM) are Mexican Wolf Studbook Number, NK
number, AF number, Robert L. Rausch collector number, NEON individual ID.
These are used to find and create relationships. The problem with
relationships is that relationships are pairwise - we need a way to
reciprocally link a network, and organism ID would allow us to do that -
like the url link to the above IDs allows us to do that now within Arctos.
Can we mint DOIs or IGSNs?
On Wed, Mar 13, 2019 at 5:28 PM John Wieczorek notifications@github.com
wrote:
I don't get how what you propose is different from:
IDType = text “Mexican Wolf Studbook Number”
Description = definition of the IDType Studbook number assigned by the
Mexican Wolf Recovery Programbase URL =
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=It is very different outside the world of Arctos. The organismID would
have to be constructed from this, and what would you do to create the
organismIDs of the ten collections that have parts of the same plant?
Create ten new ID types and base URLS (just to cover that one organism -
multiply by all the collections that share any parts of any Organisms in
Arctos)?—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472645757,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hPkdJpf-GdEBgmOtXbRz8iLU1X5Bks5vWYmkgaJpZM4buGmY
.
organisms, mint compliant ID
Don't half-bake this! - I want those for events, localities, agents, .... too.
Seriously, Arctos is built to plug in to something like that. If we have a local identifier for something it's only because nobody else would do it for us.
relationships are pairwise
Not really - there's always an implied second THING out there, but we don't have to be able to find it. "{whatever relationship of} ABC:XYZ:1234" is fine even if ABC:XYZ isn't online, "{whatever relationship of} NK 1" is fine even if 40 specimens (that we can find) wear "NK 1", etc.
reciprocally
I don't think a lack of reciprocity will ever be Arctos' fault.
I know many of your examples are not capable of acting as unique identifiers, and I suspect that's true of all of them.
Can we mint DOIs
Yes, in limited quantities - there are "get a DOI" links scattered all over the place.
IGSNs
Beats me - if they have a service and are willing to provide access we should be able to.
We could also mint ARKs in unlimited quantities if there's a reason to do so.
relationships are pairwise
Not really - there's always an implied second THING out there, but we don't have to be able to find it. "{whatever relationship of} ABC:XYZ:1234" is fine even if ABC:XYZ isn't online, "{whatever relationship of} NK 1" is fine even if 40 specimens (that we can find) wear "NK 1", etc.
But we WANT to find it! 40 fish with "same lot as" requires 39 relationships on all 40 records and then I have no easy way to see them all in one place (or I just don't know how to do it). In the same way - 20 events of blood samples from Mexican wolf studbook number 1216 requires 19 relationships on 20 records (and a relationship needs to be added to ALL of them every time a new set of samples comes in! It is a lot of work....
We have litters of pups that are siblings of each other, offspring of two
parents, and parents of other litters. Each of these individual organisms
in turn may be handled multiple times over their lifetime resulting in
multiple catalog numbers of different accessions of parts, potentially at
different institutions. We need organism IDs to deal with the latter, and
relationships that can deal with the former.
On Wed, Mar 13, 2019 at 5:52 PM dustymc notifications@github.com wrote:
organisms, mint compliant ID
Don't half-bake this! - I want those for events, localities, agents, ....
too.Seriously, Arctos is built to plug in to something like that. If we have a
local identifier for something it's only because nobody else would do it
for us.relationships are pairwise
Not really - there's always an implied second THING out there, but we
don't have to be able to find it. "{whatever relationship of} ABC:XYZ:1234"
is fine even if ABC:XYZ isn't online, "{whatever relationship of} NK 1" is
fine even if 40 specimens (that we can find) wear "NK 1", etc.reciprocally
I don't think a lack of reciprocity will ever be Arctos' fault.
I know many of your examples are not capable of acting as unique
identifiers, and I suspect that's true of all of them.Can we mint DOIs
Yes, in limited quantities - there are "get a DOI" links scattered all
over the place.IGSNs
Beats me - if they have a service and are willing to provide access we
should be able to.We could also mint ARKs in unlimited quantities if there's a reason to do
so.—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472650778,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hM3K93lKilwho96shrOC8Z0_2qe0ks5vWY89gaJpZM4buGmY
.
easy way to see them
That's an interface problem.
a relationship needs to be added to ALL of them every time a new set of samples comes in!
That MAY be an interface problem too - eg, MAYBE I could just magic in reciprocals instead of the email. Not much problem technically, but there are social implications.
40 fish
That does occasionally happen, but more normal is a coyote, a beaver, 3 mice (all because the printer stuck), and all of their parasites (for reasons that don't make much sense to me).
siblings
There's an Issue somewhere about making inferences from relationships - also just a display problem.
organism IDs to deal with the latter, and relationships that can deal with the former.
Yea, there's some overlap that I don't think we can avoid. I think we need both anywhere we can - orgID is useless unless all of the bits are accessible, and relationships can't be used to find all the bits in places like GBIF. I'm not real happy with that, but I think it's reality.
See https://github.com/tdwg/dwc-qa/issues/131#issuecomment-472855544
@campmlc is MSB onboard with this in general? I thought https://github.com/ArctosDB/arctos/issues/1545#issuecomment-398469421 was a flat "no" to cataloging Occurrences.
Do you intend to do with the NEON material whatever you do with the wolves?
I just spent a ridiculous amount of time wandering around GBIF looking for a real-world example. https://www.gbif.org/occurrence/1805830446 could be http://arctos.database.museum/guid/MSB:Mamm:306392 or http://arctos.database.museum/guid/MSB:Mamm:306394, but the data are pretty light in all records so who knows. Bits and pieces of http://arctos.database.museum/guid/MSB:Mamm:306393 work too but the sex doesn't, and who knows how reliable that is in either record.
Surely the zoo issues identifiers? There's just nothing in any record that definitively links all of this stuff together, or rejects any linkage. If I were a researcher I suppose I'd find myself writing to the collections and hoping they have some more information that they're willing to dig out. We can't force other collections to play nice, but maybe we can provide a shining example.
The sex for the MSB records suggests there's more information (http://arctos.database.museum/info/ctDocumentation.cfm?table=CTSEX_CDE&field=not%20recorded - "There is data in the form of a label or field notes, and there is no mention of sex.") Is that accurate - eg, are we entering data accurately, or should that be "unknown" or something?
The NK and the "NK" in the event remarks don't line up on http://arctos.database.museum/guid/MSB:Mamm:306392, which makes me think something important is missing.
I guess my one take-away is that any "organism registry" should aggressively collect anything that might be considered an identifier associated with the individual. It's painful to dig that out of GBIF, it's impossible to know what might have been withheld from GBIF, and it's remotely possible that the zoo or whoever owns the studbook would contribute to (and use) a registry.
Mostly unrelated, USNM is using ARK - http://n2t.net/ark:/65665/359989e34-8719-4907-823e-3f55dc8181e6
If I were a researcher I suppose I'd find myself writing to the collections and hoping they have some more information that they're willing to dig out. We can't force other collections to play nice, but maybe we can provide a shining example.
Assuming researchers even notice any connection. YES to the shining example!!!!
Even if MSB continues to choose to catalog organisms instead of occurrences, we need this solution for other collections and cross collection occurrences, although I am pretty sure the difficulty of cataloging multiple events per organism in a single record has everyone convinced it isn't sustainable over the long term....
The solution will not prevent anyone from cataloging organisms over occurrences if they so desire - but I would never recommend it.
I am mostly on board after a long grieving period for our specimen event
model because of the now obvious difficulties in implementing that model
correctly. NEON seems to be the best of all possible worlds in implementing
the specimen event model, and it seemed to work OK because everything was
cataloged from scratch all according to a single consistent model and
workflow. The NEON ID is equivalent to an organism ID that links all
samples from all occurrences. See
https://arctos.database.museum/guid/MSB:Mamm:299204. And this was a static
collection on our end - we will not be receiving any more samples, samples
are still being collected from the same organisms and being deposited at
NEON's ASU repository. Again, the need for a cross-institutional,
cross-platform organism ID exist regardless of whether or not we personally
catalog organisms or occurrences.
However, Mexican wolves are a very different story, because they were all
originally cataloged as occurrences, We have spent several years and an
entire Master's thesis project on trying to consolidate these occurrences
into single records of a single organism with multiple events. This process
is far from complete. And now we are finding so many errors in the data
entry and conversion process that it appears that effort may have largely
been wasted. In addition, the specimen event model does not permit the
tracking of different accessions, or of maintaining the linkage between a
previous, submerged catalog number and the associated parts. This is a mess
that is going to have to be straightened out, and it may be easier to back
out than go forward.
The zoo specimens do have a global animal number in their database. We will
be using that. But I don't know how global it truly is. Their database does
not provide an associated url. Here is an example: GAN: 22019550
If USNM is minting Arks, then perhaps we can experiment with linking with
them. They have hosts to our parasites and vice versa. We have plenty of
examples within Arctos of different collections housing parts of the same
organism. And NEON has parts of the same organism external to Arctos.
I agree with Teresa that we should not force anyone to use one model or the
other, but we should support both. Regardless, there should be a formal
organism ID for those situations that require one.
If much of this can be resolved via the interface, is it possible to have a
toggle between an organism display and an occurrence display? If there is a
designated organism ID, I'd like to see that above the catalog number,
prominently displayed, with a click to Show in Organism View vs Show As
Separate Occurrences or something. Then we could catalog occurrences
separately, track accessions and separate catalog numbers, but be able to
view the record optionally as we do now with the combined specimen
events/parts linkages etc.?
On Thu, Mar 14, 2019 at 9:54 AM dustymc notifications@github.com wrote:
@campmlc https://github.com/campmlc is MSB onboard with this in
general? I thought #1545 (comment)
https://github.com/ArctosDB/arctos/issues/1545#issuecomment-398469421
was a flat "no" to cataloging Occurrences.Do you intend to do with the NEON material whatever you do with the wolves?
I just spent a ridiculous amount of time wandering around GBIF looking for
a real-world example. https://www.gbif.org/occurrence/1805830446 could be
http://arctos.database.museum/guid/MSB:Mamm:306392 or
http://arctos.database.museum/guid/MSB:Mamm:306394, but the data are
pretty light in all records so who knows. Bits and pieces of
http://arctos.database.museum/guid/MSB:Mamm:306393 work too but the sex
doesn't, and who knows how reliable that is in either record.Surely the zoo issues identifiers? There's just nothing in any record that
definitively links all of this stuff together, or rejects any linkage. If I
were a researcher I suppose I'd find myself writing to the collections and
hoping they have some more information that they're willing to dig out. We
can't force other collections to play nice, but maybe we can provide a
shining example.The sex for the MSB records suggests there's more information (
http://arctos.database.museum/info/ctDocumentation.cfm?table=CTSEX_CDE&field=not%20recorded
- "There is data in the form of a label or field notes, and there is no
mention of sex.") Is that accurate - eg, are we entering data accurately,
or should that be "unknown" or something?The NK and the "NK" in the event remarks don't line up on
http://arctos.database.museum/guid/MSB:Mamm:306392, which makes me think
something important is missing.I guess my one take-away is that any "organism registry" should
aggressively collect anything that might be considered an identifier
associated with the individual. It's painful to dig that out of GBIF, it's
impossible to know what might have been withheld from GBIF, and it's
remotely possible that the zoo or whoever owns the studbook would
contribute to (and use) a registry.Mostly unrelated, USNM is using ARK -
http://n2t.net/ark:/65665/359989e34-8719-4907-823e-3f55dc8181e6—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472931695,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hKm8M4_AMhTCptVm9Dkiwzue5D04ks5vWnCagaJpZM4buGmY
.
will not prevent anyone from cataloging organisms over occurrences
Correct/necessary.
recommend
Wherever this ends up, it should include documentation (and/or a publication).
I think my biggest reservation with cataloging "Occurrences" is citations. Download a bunch of data from GBIF, get some samples (all wearing different primary IDs), eventually publish garbage science because half of your material was from one poor wolf.
https://github.com/ArctosDB/arctos/issues/1130 / http://handbook.arctosdb.org/how_to/cite-specimens.html is still hanging around mostly unresolved. If we use GUIDs for "individual IDs" we could also put them in the "cite this as" column (and force-feed them to GBIF and etc.), which would cause all of those samples from the poor drained wolf to share a 'primary-ish' ID, which I think most researchers (at least those who cite anything) would notice. I'm still not convinced of anything enough to really advocate one way or the other, but this does seem like it gets at the heart of a major problem.
This is absolutely a problem: think my biggest reservation with
cataloging "Occurrences" is citations. Download a bunch of data from GBIF,
get some samples (all wearing different primary IDs), eventually publish
garbage science because half of your material was from one poor wolf."
but it is a problem with all these multiple event specimens outside of
Arctos as well. NEON especially needs to come to grips with this because
they are and will be generating so much of these kinds of data. And they
have not even begun to consider this yet. We have received loan requests
approved by NEON where the researcher did not realize that the 100 samples
being requested included multiple samples from the same individuals over
time. This would absolutely have affected results had we not pointed that
out and insisted on loaning only a single sample per animal. But this
depends on our being able to track this ourselves as well as display it to
the outside world. And yes, citation is an issue, but it is an issue
already. There seems to be no consistent or enforced policy - and while
many people are working on this it will take community discussion as well
as by-in and enforcement by journals and data publishers.
On Thu, Mar 14, 2019 at 10:46 AM dustymc notifications@github.com wrote:
will not prevent anyone from cataloging organisms over occurrences
Correct/necessary.
recommend
Wherever this ends up, it should include documentation (and/or a
publication).I think my biggest reservation with cataloging "Occurrences" is citations.
Download a bunch of data from GBIF, get some samples (all wearing different
primary IDs), eventually publish garbage science because half of your
material was from one poor wolf.1130 https://github.com/ArctosDB/arctos/issues/1130 /
http://handbook.arctosdb.org/how_to/cite-specimens.html is still hanging
around mostly unresolved. If we use GUIDs for "individual IDs" we could
also put them in the "cite this as" column (and force-feed them to GBIF and
etc.), which would cause all of those samples from the poor drained wolf to
share a 'primary-ish' ID, which I think most researchers (at least those
who cite anything) would notice. I'm still not convinced of anything enough
to really advocate one way or the other, but this does seem like it gets at
the heart of a major problem.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472957363,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hCJRclGG-vVWOiL1A9q7KJwb2mp2ks5vWnzTgaJpZM4buGmY
.
specimen event model does not permit the tracking of different accessions,
It's not something that could happen next week, but "The Tuco Model" (everything's an Event) could handle that. The specimen-event<-->part link could be expanded as well - it's the same-ish idea at The Tuco Model, but more duct-tapey.
how global it truly is..GAN: 22019550
That's easy: it isn't.
If USNM is minting Arks, then perhaps we can experiment with linking with them.
Just to be clear, their ARKs are NOT "individual IDs" - they're just proxies to their data. Another ARK could be used to point to some merge-view of their+our data.
should support both
I don't think we have any possibility of avoiding that, at least not one that doesn't break cultural collections and such. And if we ever get a year's worth of GPS tag data or such maybe we'll want to use one catalog number for it, just to avoid minting a million new "records."
interface
Interface is "easy" if we get the data right. I can't quite envision how that might work at the moment, but I think the answer is ultimately "yes."
enforcement by journals and data publishers
I think we can play a part in that as well - eg, did this person asking to borrow specimens do what we ask them to do in the past? There's an "agent rank" option to share those sorts of data internally across Arctos collections.
Can you explain the tuco model?
Yes, all of the current ID's we use to create relationships and link
specimens are not global. Mexican wolf studbook numbers, NK numbers, AF
numbers, collector numbers, NEON numbers, ear tag numbers etc etc. Yet that
is what we have. Short of barcoding every organism and all it's parts at
the moment of collection (which would be nice), I'm not sure I see a way
around that in the current universe. The trick is keeping those
identifiers, acknowledging there will be mistakes caused by duplicates or
mistranscription, but making the data discoverable so that mistakes can be
identified and resolved. So we can use what Teresa originally proposed as
an organism ID, with all it's faults, realizing that that is what human
beings will use and recognize as an ID, and then come up with some other
truly unique id that can some how be assigned to these specimens that are
linked by their relationships across occurrences and collections.
On Thu, Mar 14, 2019 at 10:57 AM dustymc notifications@github.com wrote:
specimen event model does not permit the tracking of different accessions,
It's not something that could happen next week, but "The Tuco Model"
(everything's an Event) could handle that. The specimen-event<-->part link
could be expanded as well - it's the same-ish idea at The Tuco Model, but
more duct-tapey.how global it truly is..GAN: 22019550
That's easy: it isn't.
If USNM is minting Arks, then perhaps we can experiment with linking with
them.Just to be clear, their ARKs are NOT "individual IDs" - they're just
proxies to their data. Another ARK could be used to point to some
merge-view of their+our data.should support both
I don't think we have any possibility of avoiding that, at least not one
that doesn't break cultural collections and such. And if we ever get a
year's worth of GPS tag data or such maybe we'll want to use one catalog
number for it, just to avoid minting a million new "records."interface
Interface is "easy" if we get the data right. I can't quite envision how
that might work at the moment, but I think the answer is ultimately "yes."enforcement by journals and data publishers
I think we can play a part in that as well - eg, did this person asking to
borrow specimens do what we ask them to do in the past? There's an "agent
rank" option to share those sorts of data internally across Arctos
collections.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472962538,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hGBn17F2HPyLurcYhhDqDaJwm1iLks5vWn91gaJpZM4buGmY
.
Is far enough along to throw it at some data and see what sticks?
If so, what should we call it? I still dislike dwc:organismID - I thing we should avoid encouraging this for wolfpacks and such. (We probably can't prevent that.) https://terms.tdwg.org/wiki/dwc:individualID seems closer, but I think maybe it's been deprecated?? https://github.com/tdwg/dwc 404s everything now so ?? @tucotuco
explain the tuco model?
JRW can better, but my understanding is that basically everything's an event. Catch a specimen? Event. Identify something? Event. Assign an ID to something? Event. That would be hugely powerful, but maybe not so friendly to write code to. (Or maybe it is, who knows, as far as I know nobody's ever tried anything even vaguely similar.) The idea has been bouncing around since the first Arctos-in-ABQ Meeting in 20-something.
barcoding every organism and all it's parts at the moment of collection (which would be nice)
There's another good use for ARKs - they'd ensure barcodes are globally-unique and allow them to lead to specimens, making it basically impossible to cite the wrong thing.
some other truly unique id that can some how be assigned to these specimens that are linked by their relationships across occurrences and collections
That's where ARK (or something like them) come in. Give your two specimens (which share an NK or something, which might also be applied to 40 parasites and a duck for some reason) a GUID and point it to something that contains both specimens and which points to the bits-n-pieces of the composite "individual" - http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none works, as could GBIF. If "individualbank" comes to be you'd just redirect your ARK to it. If you find another sample in GBIF you could redirect the ARK to that. Along with the usual GUIDdy stuff, ARK (and DOI and etc.) provides a stable identifier, not a stable resolution - you can change the action without changing the value.
How about the MSB NEON records? They are as consistent and complete as we
can make them given the quality of data we received, which wasn't great.
But it will be a good test case. We converted everything from individual
occurrences to single organisms with multiple events. We had to ask NEON to
create the NEON ID for an organism ID, because they had only sample IDs. If
we can mint official dwc organism ID for these records, and add ARK Ids
etc, we should be able to eventually integrate with the ASU database, which
is under development. We would be ahead of that game.
On Thu, Mar 14, 2019 at 11:28 AM dustymc notifications@github.com wrote:
Is far enough along to throw it at some data and see what sticks?
If so, what should we call it? I still dislike dwc:organismID - I thing we
should avoid encouraging this for wolfpacks and such. (We probably can't
prevent that.) https://terms.tdwg.org/wiki/dwc:individualID seems closer,
but I think maybe it's been deprecated?? https://github.com/tdwg/dwc 404s
everything now so ?? @tucotuco https://github.com/tucotucoexplain the tuco model?
JRW can better, but my understanding is that basically everything's an
event. Catch a specimen? Event. Identify something? Event. Assign an ID to
something? Event. That would be hugely powerful, but maybe not so friendly
to write code to. (Or maybe it is, who knows, as far as I know nobody's
ever tried anything even vaguely similar.) The idea has been bouncing
around since the first Arctos-in-ABQ Meeting in 20-something.barcoding every organism and all it's parts at the moment of collection
(which would be nice)There's another good use for ARKs - they'd ensure barcodes are
globally-unique and allow them to lead to specimens, making it basically
impossible to cite the wrong thing.some other truly unique id that can some how be assigned to these
specimens that are linked by their relationships across occurrences and
collectionsThat's where ARK (or something like them) come in. Give your two specimens
(which share an NK or something, which might also be applied to 40
parasites and a duck for some reason) a GUID and point it to something that
contains both specimens and which points to the bits-n-pieces of the
composite "individual" -
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=none
works, as could GBIF. If "individualbank" comes to be you'd just redirect
your ARK to it. If you find another sample in GBIF you could redirect the
ARK to that. Along with the usual GUIDdy stuff, ARK (and DOI and etc.)
provides a stable identifier, not a stable resolution - you can change the
action without changing the value.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472976833,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hKCFQJxPxVCXcqFdphbD09wlwspFks5vWoadgaJpZM4buGmY
.
I think my biggest reservation with cataloging "Occurrences" is citations. Download a bunch of data from GBIF, get some samples (all wearing different primary IDs), eventually publish garbage science because half of your material was from one poor wolf.
Thus the need for organism ID
more duct-tapey.
No, please no - the duct tape is killing me with the wolves.
There's an "agent rank" option to share those sorts of data internally across Arctos collections.
Wait, what? Needs Documentation (and I gotta go find out what my rank is... :-)
Is far enough along to throw it at some data and see what sticks?
If so, what should we call it? I still dislike dwc:organismID - I thing we should avoid encouraging this for wolfpacks and such. (We probably can't prevent that.) https://terms.tdwg.org/wiki/dwc:individualID seems closer, but I think maybe it's been deprecated?? https://github.com/tdwg/dwc 404s everything now so ??
I think we can test the waters - we aren't going to break anything that isn't already broken as far as I'm concerned. Go here for working definition: https://dwc.tdwg.org/terms/#organismID (I think) although the example is not exactly what I'd hope for (It's an Arctos cataloged item url)
As for the event model. With the wolves, that is what I am arguing we do...essentially. Every time a particular wolf is encountered, we record an event, which in the case of Arctos is we create a catalog item. What we are missing from John's event model is the way to connect the events that are all related to the same organism. The relationship "same individual as" helps, but doesn't provide a way to connect more than two events (at least visually or for anyone outside of Arctos). I would argue that we are doing John's model, just not all the way.
And on a final note, I think I finally processed what John was talking about my solution being Arctos-centric. I'm open to the ARK idea, but I need to go read about it in more detail.
Is this where Arks come from? https://arkids.net/items
I'm not sure that I feel comfortable with the persistence of this.
What about these guys? https://identifiers.org/
I found it via this: https://www.ebi.ac.uk/miriam/main/datatypes/MIR:00000558
The Research Resource Identification Initiative provides RRIDs to 4 main classes of resources: Antibodies, Cell Lines, Model Organisms, and Databases / Software tools. The initiative works with participating journals to intercept manuscripts in the publication process that use these resources, and allows publication authors to incorporate RRIDs within the methods sections. It also provides resolver services that access curated data from 10 data sources: the antibody registry (a curated catalog of antibodies), the SciCrunch registry (a curated catalog of software tools and databases), and model organism nomenclature authority databases (MGI, FlyBase, WormBase, RGD), as well as various stock centers. These RRIDs are aggregated and can be searched through SciCrunch.
need for organism ID
I don't think I see any silver bullets here. We do, or should, already provide everything needed to do good science, and we still get "3 wolves from Arizona" in publications. This might make it easier to find certain information in certain situations or to understand identifiers, but still relies on accurate entry, conscientious users, good loan instructions, etc.
So the definition of an DWC:organism is what we're trying to avoid, and the example of a DWC:organism is what we're trying to move away from - can we use a different term here pretty please?
missing from John's event model is the way to connect the events that are all related to the same organism
That's just another event.
at least visually
Again, that's just UI. I can eg magic IndividualIDs out of relationships, and there's certainly no quantitative limit on that. If the scope of this is "Arctos" then it's a lot of work to do something that does absolutely nothing new for us. It's not really even useful for other Arctos-like systems - we could easily share relationship-grade data. If symbiota-or-whatever publishes enough information to recognize "duplicates", has a place to store a string, and a Curator is willing to make the world a little more awesome, then this is a useful thing. (Perhaps more useful if some sort of 'individual resolver' was built.) If not, maybe it isn't.
Maybe that's just a matter of who owns/issues/manages the IDs. "Relationships lite" should be fairly trivial, and I'll probably do something like this even if it's just to make GBIF et al. slightly less twitchy. That would be no extra work for ya'll, but you'd have to deal with the possibility that one of "my" IDs will be used in a publication. I could use ARKs-or-similar, UC probably won't redirect them to a "swipe your card to see this resource!" site, but they'd still be managed by machine logic and owned by something other than the collection. If I were a Curator I think I'd probably want to bring my own IDs and use them as I please, although I'd certainly use relationships to find things that could use an ID.
Haha, no ARK isn't the video game thing, but I noticed Google REALLY likes that... Lots of orgs issue ARKs, I get mine from https://ezid.cdlib.org/ just because I get DOIs from them - they're like the Wal-Mart of identifiers! Like all identifiers, ARK is as persistent as a curator is willing to make them. ARKs are also as resolvable as anyone is willing to make them. I think they're technically a good approach, but they also don't have anything remotely like the buy-in of DOI.
@campmlc Do you intend to split the NEON records up or leave them as they are?
If you split, would you use the neon ID as individualID or want to mint something a little more robust? As I mentioned above, the NEON ID has some obvious issues aside from being a random string that anyone could assign to anything for any reason - http://arctos.database.museum/SpecimenResults.cfm?oidtype=NEON%20sample%20ID&oidnum=WOOD.20160928.002A08.V
Either way, if ASU is interested and has the technology to support that interest then it could be a good test of the concept at an above-Arctos scale. I think this is all fairly trivial from Arctos.
I would rather not split our NEON records, but instead mint the organism ID
in such a way that we can eventually link to ASU records, if they can do
the same. I don't really want to involve NEON in the discussion at this
time, honestly, but would prefer to develop something for us as a model
that they can then work with, but which also works for us on a broader
scale. If we do it right, our solution should apply to NEON and other
scenarios.
The NEON ID is what they created for individual organisms at our request.
All their primary data are at the sample level - the NEON sample ID. The
NEON sample ID uses site, date, and ear tag numbers. Their NEON "organism"
ID is domain (=site) and eartag. Not unique, because eartags can be
misread, mistranscribed, and potentially duplicated, and certainly one of
those things happened in the example you provide. There are many others. In
the case of the example, at least we caught the species discrepancy and
there are two separate cataloged organisms. Only one has been given the
organism ID, so in this particular case, there is not a problem. I know for
a fact there are others that are. But that is what they are using, and that
is what we have to go on. We can't fix the errors, but we can make them
discoverable.
On Fri, Mar 15, 2019 at 9:19 AM dustymc notifications@github.com wrote:
@campmlc https://github.com/campmlc Do you intend to split the NEON
records up or leave them as they are?If you split, would you use the neon ID as individualID or want to mint
something a little more robust? As I mentioned above, the NEON ID has some
obvious issues aside from being a random string that anyone could assign to
anything for any reason -
http://arctos.database.museum/SpecimenResults.cfm?oidtype=NEON%20sample%20ID&oidnum=WOOD.20160928.002A08.VEither way, if ASU is interested and has the technology to support that
interest then it could be a good test of the concept at an above-Arctos
scale. I think this is all fairly trivial from Arctos.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-473327051,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hCb1fZ_B0sTznjdpLGKiyDgwqLvMks5vW7nvgaJpZM4buGmY
.
Can you explain the tuco model?
An Event-based model would make Events generic and central to the schema. Events would probably track combinations of action/actor/protocol/place/time/result. Right now I think the closest thing in Arctos to how connected Events would be is Agents - they are everywhere, because they are actors in Events.
We don't have the model modeled, but I imagine that it would consist of the same concepts of focal interest that would translate to tables in the database. Many of those are in Arctos already. Cataloged_Items, Collection_Objects, Agents, Projects, Identifications, Localities, Georeferences... Events would relate pairs of these concepts. Organisms could be one of the concepts. Back in the day, Organisms had a table (Biological_Individual) in Arctos, but that table met its demise at some point.
Go here for working definition: https://dwc.tdwg.org/terms/#organismID (I think) although the example is not exactly what I'd hope for (It's an Arctos cataloged item url)
Though you may not want to include all the things that dwc:Organism includes, everything you want to include is a dwc:Organism. The example in Darwin Core can be easily changed without review. If we have a real one that will persist, I can make that change.
What we are missing from John's event model is the way to connect the events that are all related to the same organism.
If Organisms existed, they could be connected to whatever you want. I don't remember all of the connections Biological_Individual had in the early model. An ER diagram of that probably still exists.
https://terms.tdwg.org/wiki/dwc:individualID seems closer, but I think maybe it's been deprecated?? https://github.com/tdwg/dwc 404s everything now so ?? @tucotuco
dwc:individualID was deprecated on 2014-10-24, replaced by dwc_organismID (http://rs.tdwg.org/dwc/terms/#dwc:organismID with full canonical definition and history currently at https://github.com/tdwg/dwc/blob/master/vocabulary/term_versions.csv#L44 - this line number could change with subsequent changes to Darwin Core). The pattern http://rs.tdwg.org/dwc/terms/organismID was always a way to resolve to the latest version of a Darwin Core term.
@campmlc that sounds reasonable to me - if you do split them, all Occurrences will need to carry the "organism ID" anyway. (Although it would then need to resolve to something different.) And the situation with string-IDs is absolutely typical.
Should I grab ARKs pointed to the Arctos GUID for them? I still think it's better if you "own" the IDs, but that's probably not terribly reasonable here.
@tucotuco I think what we're looking for is "biological individual" (whatever we call it) and I'm happy to ignore the 80% or so of life that most of us probably see as "fringe cases" for now. I think I'd be happy enough if "packs" was removed from the example. And maybe clarify colonies - "a cave full of bats" makes me twitchy, "a Portuguese man o' war" doesn't.
Shall we do this with "organism ID"? "DWC:OrganismID"??
Table biological_individual was a child of cataloged_item, so doesn't do what we need here anyway. It, along with "herp" and "mamm" and such, was replaced by Attributes.
We can go ahead with the ARKs for the NEON IDs as a test. But doesn't GBIF
give doi's to occurrences? Maybe we could ask them for a doi for an
organism ID? Or do both?
On Fri, Mar 15, 2019 at 10:50 AM dustymc notifications@github.com wrote:
@campmlc https://github.com/campmlc that sounds reasonable to me - if
you do split them, all Occurrences will need to carry the "organism ID"
anyway. (Although it would then need to resolve to something different.)
And the situation with string-IDs is absolutely typical.Should I grab ARKs pointed to the Arctos GUID for them? I still think it's
better if you "own" the IDs, but that's probably not terribly reasonable
here.@tucotuco https://github.com/tucotuco I think what we're looking for is
"biological individual" (whatever we call it) and I'm happy to ignore the
80% or so of life that most of us probably see as "fringe cases" for now. I
think I'd be happy enough if "packs" was removed from the example. And
maybe clarify colonies - "a cave full of bats" makes me twitchy, "a
Portuguese man o' war" doesn't.Shall we do this with "organism ID"? "DWC:OrganismID"??
Table biological_individual was a child of cataloged_item, so doesn't do
what we need here anyway. It, along with "herp" and "mamm" and such, was
replaced by Attributes.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-473361654,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hIQ176u5TnVIkYUdjFALW9vSrjBlks5vW89KgaJpZM4buGmY
.
GBIF give doi's to occurrences
Not that I'm aware of - they give DOIs to datasets.
I'm happy to make use of all the DOIs they might be willing to provide!
Yes, I guess you are right - the doi is just for our dataset?. And they
mess it up too. Why is Joe Cook cited? A curator shouldn't be the author.
Cook J (2019). MSB Mammal Collection (Arctos). Version 35.24. Museum of
Southwestern Biology. Occurrence dataset https://doi.org/10.15468/oirgxw
accessed via GBIF.org on 2019-03-15.
https://www.gbif.org/occurrence/1989894063
for
Organism ID http://arctos.database.museum/guid/MSB:Mamm:3061714 occurrences
https://www.gbif.org/occurrence/search?dataset_key=b15d4952-7d20-46f1-8a3e-556a512b04c5&organism_id=http%3A~2F~2Farctos.database.museum~2Fguid~2FMSB%3AMamm%3A306171
On Fri, Mar 15, 2019 at 11:14 AM dustymc notifications@github.com wrote:
GBIF give doi's to occurrences
Not that I'm aware of - they give DOIs to datasets.
I'm happy to make use of all the DOIs they might be willing to provide!
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-473370536,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hJv6qZFZ8HxrRHhq7LE5056YkcW8ks5vW9TrgaJpZM4buGmY
.
Yes, I guess you are right - the doi is just for our dataset?. And they mess it up too. Why is Joe Cook cited? A curator shouldn't be the author.
HAHAHAHAHA! I am the author for UTEP's collections....
As for using NEON - @campmlc do you have one example of something where part of an organism is at MSB and part at some non-Arctos institution that would play nice to see if we could make it all work at the aggregator level?
Oh damn, we can't even agree on the definition of "organism"....
I like the term biological individual for Arctos purposes and I suggest:
Identifier | http://rs.tdwg.org/dwc/terms/Organism
-- | --
Definition | A particular organism identified as a single taxon~or defined group of organisms considered to be taxonomically homogeneous~.
Comments | Instances of the dwc:Organism class are intended to facilitate linking ~one or more dwc:Identification instances to one or more~ dwc:Occurrence instances. Therefore, things that are typically assigned scientific names (such as viruses, hybrids, and lichens) ~and aggregates whose occurrences are typically recorded (such as packs, clones, and colonies)~ are included in the scope of this class.
Examples | A specific bird. ~A specific wolf pack.~ A specific instance of a bacterial culture.
Shouldn't a group of individual, multi-cellular organisms be something else? Colony perhaps? I realize that there will be coral scientists calling a colony of individual polyps an organism, but somehow, we absolutely need to distinguish between a pack of wolves; wolves and their offspring; and a lone wolf...
Congrats on all the publications!
"working at the aggregator level" is going to require a shared whateverwecallitID, so that would need to include finding a mutually-agreeable ID and including it as OrganismID in both datasets.
Yes on the wolf-thing, but your definition (single taxon) still allows it (and presumably disallows anything where one "bit" is identified to a subspecies and another to species/concept/whatever, not that anyone would let that level of detail stop them).
For our purposes, I think a workable definition is "an identifier applied to a biological individual" and we can ignore the corals (and defining taxa and broken sets of teacups and everything else) until they find a way to melt something.
Congrats on all the publications!
I know, right? Damn I am productive! Now if everyone will use my ORCID, I should be able to get hired as faculty somewhere....
"working at the aggregator level" is going to require a shared whateverwecallitID, so that would need to include finding a mutually-agreeable ID and including it as OrganismID in both datasets.
Yeah, that's why I asked about someone who would play nice...
Yes on the wolf-thing, but your definition (single taxon) still allows it (and presumably disallows anything where one "bit" is identified to a subspecies and another to species/concept/whatever, not that anyone would let that level of detail stop them).
Jim Murphy (Curator of herps at the National Zoo) once told me a story about defending his thesis. He was asked how he knew that he wasn't just a collection of individual organisms, his response: "Why are you asking us that? There really is no such thing as a single organism, but if we are going to make sense of stuff we'll have to decide on something eventually! I'll leave it to someone with more brain matter...
For our purposes, I think a workable definition is "an identifier applied to a biological individual" and we can ignore the corals (and defining taxa and broken sets of teacups and everything else) until they find a way to melt something.
I'm good with that. Simple, easy to remember.
I can look for some USNM specimens that have a host/parasite relationship
to UNM, if that would help. Maybe same lot as . . .
Can also find same individual as at AMNH?
On Fri, Mar 15, 2019 at 1:04 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
Congrats on all the publications!
I know, right? Damn I am productive! Now if everyone will use my ORCID, I
should be able to get hired as faculty somewhere...."working at the aggregator level" is going to require a shared
whateverwecallitID, so that would need to include finding a
mutually-agreeable ID and including it as OrganismID in both datasets.Yeah, that's why I asked about someone who would play nice...
Yes on the wolf-thing, but your definition (single taxon) still allows it
(and presumably disallows anything where one "bit" is identified to a
subspecies and another to species/concept/whatever, not that anyone would
let that level of detail stop them).Jim Murphy (Curator of herps at the National Zoo) once told me a story
about defending his thesis. He was asked how he knew that he wasn't just a
collection of individual organisms, his response: "Why are you asking us
that? There really is no such thing as a single organism, but if we are
going to make sense of stuff we'll have to decide on something eventually!
I'll leave it to someone with more brain matter...For our purposes, I think a workable definition is "an identifier applied
to a biological individual" and we can ignore the corals (and defining taxa
and broken sets of teacups and everything else) until they find a way to
melt something.I'm good with that. Simple, easy to remember.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-473407184,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hMvEGzq3Sf3tJpIbyIZz1_E0RpJHks5vW-7LgaJpZM4buGmY
.
@campmlc host/parasite isn't what we are after - needs to be "same individual as", so AMNH?
Here is one that has tissues at MSB, voucher at AMNH, and the voucher is a
symbiotype of parasite holotypes at HWML (not yet cataloged in Arctos).
MSB:Mamm:210449 https://arctos.database.museum/guid/MSB:Mamm:210449
symbiotype
Search of all MSB / AMNH specimens
https://arctos.database.museum/saved/MSB%20AMNH%20shared%20records
http://arctos.database.museum/saved/MSB%20AMNH%20shared%20records
Then there are specimens from the same expeditions that were shared between
MSB and Collecion Boliviana de Fauna (CBF).
Here is a holotype with tissues in the US and voucher in Bolivia:
https://arctos.database.museum/guid/MSB:Mamm:239826
Here is the complete list.
https://arctos.database.museum/saved/MSB%20CBF%20shared%20records
On Fri, Mar 15, 2019 at 3:30 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
@campmlc https://github.com/campmlc host/parasite isn't what we are
after - needs to be "same individual as", so AMNH?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-473447424,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hFRj5IeUamwkKSuPM3m9Ahs7sTLLks5vXBD5gaJpZM4buGmY
.
Note that most of these have not had the relationships created. I added
same individual as for the two examples.
On Sat, Mar 16, 2019 at 5:08 PM Mariel Campbell campbell@carachupa.org
wrote:
Here is one that has tissues at MSB, voucher at AMNH, and the voucher is a
symbiotype of parasite holotypes at HWML (not yet cataloged in Arctos).MSB:Mamm:210449 https://arctos.database.museum/guid/MSB:Mamm:210449
symbiotypeSearch of all MSB / AMNH specimens
https://arctos.database.museum/saved/MSB%20AMNH%20shared%20records
http://arctos.database.museum/saved/MSB%20AMNH%20shared%20recordsThen there are specimens from the same expeditions that were shared
between MSB and Collecion Boliviana de Fauna (CBF).
Here is a holotype with tissues in the US and voucher in Bolivia:
https://arctos.database.museum/guid/MSB:Mamm:239826Here is the complete list.
https://arctos.database.museum/saved/MSB%20CBF%20shared%20records
On Fri, Mar 15, 2019 at 3:30 PM Teresa Mayfield-Meyer <
[email protected]> wrote:@campmlc https://github.com/campmlc host/parasite isn't what we are
after - needs to be "same individual as", so AMNH?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-473447424,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hFRj5IeUamwkKSuPM3m9Ahs7sTLLks5vXBD5gaJpZM4buGmY
.
I had been thinking there would be only one allowed organismID. Maybe that is silly. Maybe it is fine to have as many as you like. That way you could include your own AND those of other collections (in or out of Arctos). That way you could also potentially go directly to GBIF to get the set of Occurrences for all matching organismIDs.
@tucotuco this made me realize that an animal could easily have two organism IDs that mean something different. In the zebra example from tdwg/dwc-qa#131 each zebra might have an organism ID and the herd (or even defined groups within the herd) might also have one, such that each zebra would have multiple Organism IDs with very good reasons, making it all more complicated!
@campmlc do you have a contact at AMNH we can work with? We are going to need them to add the organism ID if we want to demonstrate it working outside of Arctos.
Website lists the CM as Neil Duncan - [email protected] (so I remember)
Well, I'm in NYC right now . . .I'll ask around. Anyone else know?
On Mon, Mar 18, 2019, 3:46 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
@campmlc https://github.com/campmlc do you have a contact at AMNH we
can work with? We are going to need them to add the organism ID if we want
to demonstrate it working outside of Arctos.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474072689,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hJGjf5hWGX6BHugoSTZnO4nuA5dEks5vX-0igaJpZM4buGmY
.
I'll try emailing the CM and we can go from there.
two organism IDs that mean something different
hence my reluctance to tie ourselves too tightly to the DWC concept. We can't stop anyone from doing anything and sharing it via DWC. We can choose good meaningful identifiers, apply them locally, and encourage others to build something bigger than anything we have individually through them.
OK, so can we work on this within Arctos while we wait on a response from AMNH (or have to find a different test partner)?
We still need to agree on the OrganismID term and definition. People are publishing about the problem of "organism", so we aren't alone, but we need something to work with! Anyone with time to spare may want to read:
http://philsci-archive.pitt.edu/13376/
https://link.springer.com/article/10.1007/s10539-016-9551-1
https://plato.stanford.edu/entries/biology-individual/#ProBioInd
After perusing this entertainment, I think we eventually will need several types of other IDs to pass as dwc:OrganismID OR we just need to stick with their definition and let everyone sort out the multiple dwc:OrganismIDs assigned to cataloged stuff. For now we are concerned with "stuff from the same plant/animal" so I propose we create:
OtherID=individual organism
Definition=all cataloged items bearing this ID were obtained from a single, biological representation of a given taxon such as a bird, mouse, lizard, fish, beetle, tree, or worm
UGH - I have been mulling this definition all afternoon and I sincerely hope that someone can make it better.
THEN
Can we acquire an ARK that we can define as Mexican Wolf Studbook Number 1216 so that I can see how this will work before proposing the use of an ARK to AMNH?
I don't think we have to get overly philosophical as long as we're working with wolves!
We may have several kinds of values in our one "individual ID" but I still think having different local types would render this unusable.
I have no serious objections to your term or definition, although I might drop the "such as...." bits. Whatever, definitions are easy to change. Not terribly important either, and may be Department of Redundancy Department country, but "individual organism ID" or "individual organism identifier" somehow feels better to me.
I can get you an ARK or two, I just need a URL to which they point. If you want Arctos to provide lots of them we should probably talk about ownership, but if you're OK with your identifiers being owned by not-you then it's probably not much of a problem to add a 'grab an ARK' button somewhere.
What about including plants?
I had an email from Neil - just trying to find out if I can swing by this
week or not.
On Mon, Mar 18, 2019, 6:23 PM dustymc notifications@github.com wrote:
I don't think we have to get overly philosophical as long as we're working
with wolves!We may have several kinds of values in our one "individual ID" but I still
think having different local types would render this unusable.I have no serious objections to your term or definition, although I might
drop the "such as...." bits. Whatever, definitions are easy to change. Not
terribly important either, and may be Department of Redundancy Department
country, but "individual organism ID" or "individual organism identifier"
somehow feels better to me.I can get you an ARK or two, I just need a URL to which they point. If you
want Arctos to provide lots of them we should probably talk about
ownership, but if you're OK with your identifiers being owned by not-you
then it's probably not much of a problem to add a 'grab an ARK' button
somewhere.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474124531,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hARukKhBg7yOjUCo-M7bDixT47Wlks5vYBHqgaJpZM4buGmY
.
plants
Certainly no objections, and that's a huge use case, but "individual" is really weird (half the aspen in Utah, whatever mushrooms do for fun, etc.) and the things they call "duplicates" (and likely want to apply this to) are not always individuals. The technology is the same - if >1 things are sharing an ID then they can be linked - but the terminology/application/etc. is likely to be something the botanists are going to need some time to sort out.
And now you are coming around to why DwC defines Organism the way it does.
:-)
< @tucotuco https://github.com/tucotuco this made me realize that an
animal could easily have two organism IDs that mean something different. In
the zebra example from tdwg/dwc-qa#131
https://github.com/tdwg/dwc-qa/issues/131 each zebra might have an
organism ID and the herd (or even defined groups within the herd) might
also have one, such that each zebra would have multiple Organism IDs with
very good reasons, making it all more complicated!
There is a subtlety in the semantics world to be aware of. The organismID
identifies the Organism (the organismID identifies the Organism, nothing
more, nothing less). Thus, the identifier for the herd is not apt as an
identifier for the zebra. It may be that the zebra is part of an identified
herd, but it is not the herd. It can not have the organismID of the herd.
It can be related to the herd with a ResourceRelationship zebra id - is a
member of - herd id.
So, what happens when a special fish is found in a lot that was given an
organismID, and pulled out. The fish never had the organismID of the lot.
Only the lot did. The special fish was never given an identity before the
act of separating it from the rest.
Once these semantics are understood, I think it isn't complicated anymore.
On Mon, Mar 18, 2019 at 11:33 PM dustymc notifications@github.com wrote:
plants
Certainly no objections, and that's a huge use case, but "individual" is
really weird (half the aspen in Utah, whatever mushrooms do for fun, etc.)
and the things they call "duplicates" (and likely want to apply this to)
are not always individuals. The technology is the same - if >1 things are
sharing an ID then they can be linked - but the
terminology/application/etc. is likely to be something the botanists are
going to need some time to sort out.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474174597,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AAcP684Mf8As8j5qOJ-F93JkXa1X2PJEks5vYEyBgaJpZM4buGmY
.
it isn't complicated anymore
You find 10 things that share an orgid on GBIF. Are they 10 hunks of the same critter, or 10 individual critters that happened to be in the general vicinity of each other?
I more or less see why an exchange standard would need broad definitions, but I don't think it's appropriate for Arctos. We may well not be able to find a definition of a single ID that works for wolves and "same meadow or better" 'duplicate' plants, in which case we can just fire up another ID (and possibly share it with DWC via OrgID, if it still fits under that umbrella).
We may well not be able to find a definition of a single ID that works for wolves and "same meadow or better" 'duplicate' plants, in which case we can just fire up another ID (and possibly share it with DWC via OrgID, if it still fits under that umbrella).
So we are back to somehow identifying different types "OrganismIDs"? Can we do this WITHIN the OTHER_ID table we have now or do they need to be somewhere else so that we can easily tell what is being sent as a dwc:OrganismID?
somehow identifying different types "OrganismIDs"?
No!
We may have various things under strict definitions in Arctos that get mapped to a single thing with a broader definition in DWC. We don't (and shouldn't, unless it somehow satisfactorily answers the little GBIF though experiment above) have to find a single identifier that covers everything under the scope of DWC:OrganismID. We may end up with "individual ID," "polyp ID," "whatever the zebra-people are trying to do ID," "these plants all got stuffed into the same ziplock for some reason ID" and 50 more things that all get mapped to DWC:OrganismID. Only one of them will make sense for a wolf (or polyp or whatever), and all would face the exact same considerations regarding values and all that jazz.
So, what happens when a special fish is found in a lot that was given an organismID, and pulled out. The fish never had the organismID of the lot. Only the lot did. The special fish was never given an identity before the act of separating it from the rest.
Well, we currently have two ways this could go in Arctos. If the "specialness" of the fish DOES NOT include a new Identification Event, Locality, or Other_ID (maybe it was measured), then we could just change the Part structure so that rather than one Part Name with a count of 45, we now have one Part Name with count of 44 and one Part Name with count of 1 which has some Part Attribute associated with it, because it is Special. In this case, the organismID will still be associated with all the original Parts of the lot.
However, if the "specialness" is due to a separate Identification Event, the assignment of a new LOCALITY, or a unique Other_ID; the organismID will remain with the cataloged lot, which should also retain the original count of Part Name. Hopefully, an Event will be recorded that records the Use of the special fish, then the special fish will be given a catalog number with all the appropriate Events AND a Relationship Event will be recorded for both the Lot and the Special Fish using "same lot as".
How does anyone outside of Arctos know that the special fish is part of that lot AND how can I see all of the "same lot as" cataloged items in one place? In Arctos, if we make the relationship between the catalog numbers, then I can toggle between them, but if a second special fish is removed from the lot, I now have to toggle between three and so on with no easy way to see them all in one place. However, if the lot has an OrganismID and I place that same OrganismID on all "special" members of the lot with the "same lot as" relationship, I could then see them all in one place - the same way that the Mexican Wolf Studbook Number now works. Can we use the relationship of the Other_ID to decide whether we are sending an OrganismID (same individual as) or a ResourceRelationship (same lot as)?
organismID will still be associated with all the original Parts of the lot.
In DWC, sure, if you're into that sort of thing. In Arctos, a lot of fish (or any other aggregate of the items of scientific interest) should not fall under the definition of the identifiers we intend to give wolves, so this situation can never happen.
outside of Arctos know that the special fish is part of that lot AND
We could give them all a 'found in the same general area ID' that would/could do the same thing as the 'individual ID' and might end up mapped to the same DWC concept.
relationship of the Other_ID
For Arctos, I could magic new numbers out of relationships but they can't possibly do anything that we couldn't also do with the data from which the ID is derived (except force you to change two things instead of one when you get more info). For >Arctos, this works just like the wolves: assigned meaningful shared identifiers can do stuff, other things probably can't.
I just need a URL to which they point.
Here is where I am stumped. How do we define that if we don't own it?
How do we define that if we don't own it?
Pick something that works for everyone who owns the bits-n-pieces. GBIF is the obvious option.
One of the benefits of ARK/DOI is that you can change the action without changing the identifier. If the zoo-people do something awesome you could repoint the IDs from GBIF to them, for example.
OtherID=individual organism
Definition=all cataloged items bearing this ID were obtained from a single, biological representation of a given taxon
not sure how that definition is different from
A particular organism or defined group of organisms considered to be taxonomically homogeneous.
I mean, a "biological representation" could be a school of fish....
And after @tucotuco 's comment, I don't think it matters because the RELATIONSHIP that we assign to the identifier determines whether we are talking about a wolf (same individual as) or a member of a wolf pack (same lot as)
How do we define that if we don't own it?Pick something that works for everyone who owns the bits-n-pieces. GBIF is the obvious option.
One of the benefits of ARK/DOI is that you can change the action without changing the identifier. If the zoo-people do something awesome you could repoint the IDs from GBIF to them, for example.
WHAT at GBIF? ALL the information from GBIF comes from US. If we change something, stuff at GBIF changes. Which occurrence at GBIF will be stable enough to point at? This seems like creation of a circular reference....
definition
However we say it, "a single wolf" should fit and "more than one wolf" should not.
RELATIONSHIP that we assign to the identifier
These are separate things.
WHAT at GBIF?
"Things with this OccurrenceID" - eg, https://www.gbif.org/occurrence/search?organism_id=http:~2F~2Farctos.database.museum~2Fguid~2FMSB:Mamm:306171&advanced=1
Which occurrence at GBIF
The query finds Occurrences, it does not rely on them being stable.
circular reference
CollectionA feeds GBIF OrgID1. CollectionB feeds GBIF OrgID1. organism_id=OrgID1 finds them both in GBIF. It might be circular if all involved parties have the capability to point from GBIF to themselves and from themselves to each other, but if they had that we wouldn't much need this ID.
That makes my head hurt. Can we just own it.
Can we make the ARK point to
You can point ARKs to anything with a URL, but that URL probably isn't terribly stable and it won't be very enticing to anyone who's not already using Arctos (and therefore doesn't much need this).
Yeah - so what we really need is a url somewhere that has this:
Mexican Wolf
Studbook Number 1216
Managed under the blah blah blah
Born: yyyy-mm-dd
plus other "metadata" about the individual
This has been my hangup all through the conversation. We need more than a random pointer (at least that is how it feels to me).
The NMNH ARK tells me nada about the wolf that I can connect to anything that we have. In fact, I'm not sure what the heck it does. I need to stop thinking about this for a while.
This is why the studbook number itself should be part of the URL, because
it is the only linking field.
On Tue, Mar 19, 2019, 3:11 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
Yeah - so what we really need is a url somewhere that has this:
Mexican Wolf
Studbook Number 1216
Managed under the blah blah blah
Born: yyyy-mm-dd
plus other "metadata" about the individualThis has been my hangup all through the conversation. We need more than a
random pointer (at least that is how it feels to me).The NMNH ARK
http://n2t.net/ark:/65665/359989e34-8719-4907-823e-3f55dc8181e6 tells
me nada about the wolf that I can connect to anything that we have. In
fact, I'm not sure what the heck it does. I need to stop thinking about
this for a while.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474532230,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hCLrYn32Bal7tSqSnQAAI6AzrQJRks5vYTZegaJpZM4buGmY
.
what we really need
Yes, that would be ideal. The zoo people could easily pull it off for zoo animals (and add in relationships to parents and other cool stuff that they likely have)."individualBank" could probably pull it off at a much broader scope, except....
https://collections.nmnh.si.edu/search/mammals/?ark=ark:/65665/359989e3487194907823e3f55dc8181e6 seems to be "their catalog record." I don't know if that's all the information they have, or all the information they're willing to share, or a limitation of their system, or something else. I found it by matching taxonomy and locality in GBIF. Who knows if it's the same wolf, and if that's all the information they have it might not be possible to know. As always, GIGO applies....
At least the NMNH record is online. Bits of your wolf may also be in places that aren't.
Lacking a better solution, GBIF exists and is more likely than anything else to have information on the bits-n-pieces. It's far from perfect, but it may be functional-enough to serve as a neutral meeting place.
@campmlc I don't know what you mean by that. "1216" is very unlikely to be a useful identifier at any useful scale. https://www.gbif.org/occurrence/search?organism_id=1&advanced=1
So, what happens when a special fish is found in a lot that was given an organismID, and pulled out. The fish never had the organismID of the lot. Only the lot did. The special fish was never given an identity before the act of separating it from the rest.
Well, we currently have two ways this could go in Arctos. If the "specialness" of the fish DOES NOT include a new Identification Event, Locality, or Other_ID (maybe it was measured), then we could just change the Part structure so that rather than one Part Name with a count of 45, we now have one Part Name with count of 44 and one Part Name with count of 1 which has some Part Attribute associated with it, because it is Special. In this case, the organismID will still be associated with all the original Parts of the lot.
If you go the first route, the fish is represented as a part. One cannot say that parts are organisms. They are only in special cases. So, tracking them as parts gives you no capabilities of tracking them as individuals. I think that is a kludge, not a best practice use of Arctos. What was given the organismID in the first place? A Cataloged Item? Well, one cannot say that cataloged items are organisms either. They too are organisms only in special cases.
I think that if you want to fit organisms (biological individuals) into Arctos, you should do it rigorously, otherwise you are putting masking tape on top of duct tape. In the absence of an external OrganismBank, you could make one inside of Arctos as a proof of concept. It might be that you only need to add one table, Organism, and do the rest with relationships. Two tables if you want to control the types of relationships.
table
Why do we need a table/what would we put in said table?
@tucotuco I agree, but I think we need more than a table.
If we want to be rigorous, we should create a registry, even if it is pretty sparse. It seems like we need something akin to a collection in Arctos, only with some different data than we are used to so that we could generate stable urls for the individuals cataloged there, which would then become the "organismID", which anyone could use.
ARCTOS:OrgID:1 = http://arctos.database.museum/guid/ARCTOS:OrgID:1
Which holds:
Identification (taxon name) - _Canis lupus baileyi_
Description (how I tell this individual from any other of the same taxon) - "The individual in the Mexican Wolf Recovery Studbook numbered 1216"
And maybe we add metadata as needed.
Then we add "Individual Organism" to the other ID code table with the base url http://arctos.database.museum/guid/ARCTOS:OrgID:
Use only that ID as the dwc:OrganismID when sending stuff out and there you have it.
Can we accomplish the same idea with a table? My brain hurts....
Ahhhh....
Yes, we could catalog them (or otherwise give them Arctos URL/GUIDs) but that sort of seems like massive overkill. Everything we'd put in the record should already be at GBIF (and friends). And given 100 hunks of a wolf in 100 collections I think you're likely to find at least dozens of variations in most everything, which should probably be accurately reflected in the "registry."
That would also involve getting everyone else to adopt an "Arctos GUID." That sounds great to me personally, but I doubt the folks using KE-or-whatever share the sentiment.
What, other than a prettier GUID (which we can abstract through ARK-or-similar), would an ideal registry contain/do/whatever that GBIF does not?
Everything we'd put in the record should already be at GBIF (and friends)
I disagree. Take a Mexican Wolf - we have dates that blood was collected, but we don't have birth and death dates or locations (except when a final collection is made...), "names" (they all have some sort of friendly name like Caramel, etc.), parentage may be in Arctos, but if no material from a parent is in the collection, then that is missing - same with offspring. There is a bunch of metadata about the individual wolf that isn't in Arctos and would help others ensure they are using the appropriate organism ID. The registry is just a place to give the individual an ID NOT to record every physical piece of it, that happens at a bunch of disparate places and this ID allows them all to be connected.
What, other than a prettier GUID (which we can abstract through ARK-or-similar), would an ideal registry contain/do/whatever that GBIF does not?
Provide an identifier that people could comprehend and use to decide if whatever they have is part of that organism and then be able to connect whatever part of the organism they have with parts that others may have.
"Provide an identifier that people could comprehend and use to decide if
whatever they have is part of that organism"
So back to
Organism ID = "
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%20wolf%20studbook%20number&oidnum=
"
?
On Tue, Mar 19, 2019 at 7:19 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
Everything we'd put in the record should already be at GBIF (and friends)
I disagree. Take a Mexican Wolf - we have dates that blood was collected,
but we don't have birth and death dates or locations (except when a final
collection is made...), "names" (they all have some sort of friendly name
like Caramel, etc.), parentage may be in Arctos, but if no material from a
parent is in the collection, then that is missing - same with offspring.
There is a bunch of metadata about the individual wolf that isn't in Arctos
and would help others ensure they are using the appropriate organism ID.
The registry is just a place to give the individual an ID NOT to record
every physical piece of it, that happens at a bunch of disparate places and
this ID allows them all to be connected.What, other than a prettier GUID (which we can abstract through
ARK-or-similar), would an ideal registry contain/do/whatever that GBIF does
not?Provide an identifier that people could comprehend and use to decide if
whatever they have is part of that organism and then be able to connect
whatever part of the organism they have with parts that others may have.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474621641,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hLOKe4lVJ95ILTjLJP1NjwWtute9ks5vYXB8gaJpZM4buGmY
.
I'm too tired to think right now, but I think "everything" was intended to mean " everyone's identifiers."
I suppose it's ultimately up to whoever builds the registry, or whatever we might use in the absence of such a thing, but I think it would be exceedingly handy to find all the stuff you mention (which may be scattered across many records owned by many institutions and "natively" presented in many formats) in one place, and AFAIK GBIF does that better than anyone at the moment (just because of their scope).
ID allows them all to be connected
Yep, that's the critical component. Get that right and the rest is details.
identifier that people could comprehend and use to decide if whatever they have is part of that organism
That sounds a lot like printing "smart" barcodes. Those ALWAYS find a way to cause problems; information changes and you can't change the information without either changing the identifier or creating a misleading identifier or etc. "Dumb" or opaque identifiers (like barcodes) are just identifiers. ARK and DOI are both capable of carrying metadata - more information, which can be changed without messing with the identifier itself, is less than a click away.
curl https://ezid.cdlib.org/id/ark:/87299/x68g8hqw
success: ark:/87299/x68g8hqw
_updated: 1553038988
_target: http://arctos.database.museum/SpecimenResults.cfm?oidtype=Mexican%2520wolf%2520studbook%2520number&oidnum=none
erc.who: This could be some useful text.
erc.when: This could be some useful text.
_export: yes
_owner: ucb-mvz
_ownergroup: ucblibrary
_profile: erc
_created: 1552513887
_status: public
erc.what: This could be some useful text.
@campmlc that does work, but I think it will come with social problems, it may not survive the next code table cleanup or Arctos revamp, etc. It would not be my first choice. Still worlds better than "1"!
I wasn't expecting the ID to provide any meaningful information, just the definition.
all the stuff you mention (which may be scattered across many records owned by many institutions and "natively" presented in many formats) in one place, and AFAIK GBIF does that better than anyone at the moment (just because of their scope).
Again, I disagree. GBIF holds Darwin Core Fields, pretty sure birth and death dates aren't in there along with other stuff that might help someone decide if the cataloged item they have is from the same animal as that described in the registry ID.
I think brains are melting from trying to think about too much at once -
specifically, how this would work in Arctos and how it would work globally.
I'll try to approach this from a different angle and see if it helps. Here
are some principles to consider for dwc:Organisms on the global scale. Is
there anything you disagree with? Are there other things you can think of
to add?
1) It would be useful to organize information around dwc:Organisms (the
Arctos biological individual concept is a subset of the dwc:Organism
concept, so I feel OK framing it this way for its maximal scope and formal
definitions - you don't have to put anything more than biological
individuals in Arctos and they will still be dwc:Organisms).
2) In order to organize information around dwc:Organisms, there need to be
instances of dwc:Organisms (i.e., data about real-world dwc:Organisms).
3) Identifiers are needed to track instances of dwc:Organisms
4) Best practices around identifiers suggest that, to the extent possible,
they should be persistent, resolvable (something happens when you use a
service to resolve them), globally unique identifiers
5) Uniqueness of dwc:Organism instance identifiers can be enforced in a
registry.
6) A useful registry would support and allow users to posit relationships
for dwc:Organisms. Question: Should a dwc:Organism be registered to
participate in a relationship?
Examples of potential dwc:Organism relationship types:
sameAs dwc:Organism
hasSample dwc:MaterialSample
(siblingOf, offspringOf, parentOf, ...) dwc:Organism
memberOf dwc:Organism (where the object dwc:Organism is a group)
presentIn dwc:Occurrence
7) Identifiers for dwc:Organisms are only actually needed when a
dwc:Organism enters into a relationship, though they could be minted and
applied to anything that is a dwc:Organism in anticipation of there being
relationships one day.
8) dwc:Organisms can have attributes, both mutable and immutable.
9) The list of attributes of dwc:Organisms available to populate in the
registry should be limited, stable, and carefully chosen.
The idea of registries for Biodiversity Data Classes, including Organisms,
are on GBIFs radar, though no work plan for them currently exists.
GBIF is a DataCite member and can issue dois, but is already too big for
DataCite to handle using dois as Occurrence identifiers. GBIF would like to
use IGSNs for biodiversity identifiers of all kinds, but that conversation
is still to be had.
On Tue, Mar 19, 2019 at 8:19 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
Everything we'd put in the record should already be at GBIF (and friends)
I disagree. Take a Mexican Wolf - we have dates that blood was collected,
but we don't have birth and death dates or locations (except when a final
collection is made...), "names" (they all have some sort of friendly name
like Caramel, etc.), parentage may be in Arctos, but if no material from a
parent is in the collection, then that is missing - same with offspring.
There is a bunch of metadata about the individual wolf that isn't in Arctos
and would help others ensure they are using the appropriate organism ID.
The registry is just a place to give the individual an ID NOT to record
every physical piece of it, that happens at a bunch of disparate places and
this ID allows them all to be connected.What, other than a prettier GUID (which we can abstract through
ARK-or-similar), would an ideal registry contain/do/whatever that GBIF does
not?Provide an identifier that people could comprehend and use to decide if
whatever they have is part of that organism and then be able to connect
whatever part of the organism they have with parts that others may have.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474621641,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AAcP6_HVIk2S4a5C20t6Pn_OFQuCjJebks5vYXB9gaJpZM4buGmY
.
Speaking of brain melting, I spoke with the AMNH Mammal CM today, and it
was a surprise to him to hear that there are tissues in other collections
for mammals they hold from 1990s Bolivian expeditions. They have nothing
recorded about tissues or about their having been a division of the
specimens between three institutions.To give an idea of the challenge here,
the AMNH KeMU database has room for only one identifier beyond the catalog
number, a primary collector number, and this is not the number which links
the tissues. The identifier that links the tissue and all other data is the
NK field number, which is present in their scanned handwritten catalogs,
but it has never been transcribed, and there is no place for a second
identifier currently in their database. They did not know what the NK
number represented. We will need to talk with their data administrator
about the possibility of adding any fields - and this may cost them some
money, I'm afraid. However, he was very interested in working with us to
sort out their records and try to make these linkages.
So, if this is representative, the challenge we face is in making sure the
primary linking identifiers can be recognized by the outside world,can be
identified clearly as an organism ID, and can be used to join records, even
before we start adding additional urls or dois or ARKS etc with metadata.
I'm sure I'm butchering the SQL and not understanding the details, but what
about at least in the short term:
http://arctos.database.museum/OrganismID/oidtype=NK&oidnum=
On Wed, Mar 20, 2019 at 8:58 PM John Wieczorek notifications@github.com
wrote:
I think brains are melting from trying to think about too much at once -
specifically, how this would work in Arctos and how it would work globally.
I'll try to approach this from a different angle and see if it helps. Here
are some principles to consider for dwc:Organisms on the global scale. Is
there anything you disagree with? Are there other things you can think of
to add?1) It would be useful to organize information around dwc:Organisms (the
Arctos biological individual concept is a subset of the dwc:Organism
concept, so I feel OK framing it this way for its maximal scope and formal
definitions - you don't have to put anything more than biological
individuals in Arctos and they will still be dwc:Organisms).
2) In order to organize information around dwc:Organisms, there need to be
instances of dwc:Organisms (i.e., data about real-world dwc:Organisms).
3) Identifiers are needed to track instances of dwc:Organisms
4) Best practices around identifiers suggest that, to the extent possible,
they should be persistent, resolvable (something happens when you use a
service to resolve them), globally unique identifiers
5) Uniqueness of dwc:Organism instance identifiers can be enforced in a
registry.
6) A useful registry would support and allow users to posit relationships
for dwc:Organisms. Question: Should a dwc:Organism be registered to
participate in a relationship?
Examples of potential dwc:Organism relationship types:
sameAs dwc:Organism
hasSample dwc:MaterialSample
(siblingOf, offspringOf, parentOf, ...) dwc:Organism
memberOf dwc:Organism (where the object dwc:Organism is a group)
presentIn dwc:Occurrence
7) Identifiers for dwc:Organisms are only actually needed when a
dwc:Organism enters into a relationship, though they could be minted and
applied to anything that is a dwc:Organism in anticipation of there being
relationships one day.
8) dwc:Organisms can have attributes, both mutable and immutable.
9) The list of attributes of dwc:Organisms available to populate in the
registry should be limited, stable, and carefully chosen.The idea of registries for Biodiversity Data Classes, including Organisms,
are on GBIFs radar, though no work plan for them currently exists.GBIF is a DataCite member and can issue dois, but is already too big for
DataCite to handle using dois as Occurrence identifiers. GBIF would like to
use IGSNs for biodiversity identifiers of all kinds, but that conversation
is still to be had.On Tue, Mar 19, 2019 at 8:19 PM Teresa Mayfield-Meyer <
[email protected]> wrote:Everything we'd put in the record should already be at GBIF (and friends)
I disagree. Take a Mexican Wolf - we have dates that blood was collected,
but we don't have birth and death dates or locations (except when a final
collection is made...), "names" (they all have some sort of friendly name
like Caramel, etc.), parentage may be in Arctos, but if no material from
a
parent is in the collection, then that is missing - same with offspring.
There is a bunch of metadata about the individual wolf that isn't in
Arctos
and would help others ensure they are using the appropriate organism ID.
The registry is just a place to give the individual an ID NOT to record
every physical piece of it, that happens at a bunch of disparate places
and
this ID allows them all to be connected.What, other than a prettier GUID (which we can abstract through
ARK-or-similar), would an ideal registry contain/do/whatever that GBIF
does
not?Provide an identifier that people could comprehend and use to decide if
whatever they have is part of that organism and then be able to connect
whatever part of the organism they have with parts that others may have.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-474621641,
or mute the thread
<
https://github.com/notifications/unsubscribe-auth/AAcP6_HVIk2S4a5C20t6Pn_OFQuCjJebks5vYXB9gaJpZM4buGmY.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-475082651,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hDddi4dcK2zV9Fl3lkU08zDwn1Ghks5vYtkrgaJpZM4buGmY
.
My brain melted hours ago, lemme jump in here!
maximal scope
Yes, agreed, I think this is where we should focus. This is trivial and/or redundant locally.
subset
Right - and if we get "pack identifiers" we might also treat them as a subset of OrganismID (and then we'd probably have to whine about structure vs. strings to yon DWC crowd).
Question: Should a dwc:Organism be registered to participate in a relationship?
I think no, although that's probably an ideal. We still want to play nice with someone who doesn't have the resources/technology/whatever to do more than map a string to DWC, or just doesn't like the IDs issued by the registry for whatever random reason.
anticipation
Yea, that's been driving me crazy/ier. Eartagged squirrels, identifiable whales, anything that comes through an agency that owns a freezer, etc. could have more data/material floating around out there. A significant part of every collection, probably, not that we're likely to actually make those connections.
though they could be minted and applied to anything that is a dwc:Organism in anticipation
- Someone sees a whale, registers an OrgID.
- Someone else sees the same whale, registers an OrgID
- Whale dies, F&G lops off a hunk, registers an OrgID
- NMFS lops off a hunk, more OrgIDs
- MSB gets a hunk of meat, you know the drill
- MVZ gets the bones, here we go again
Eventually someone figures all that out, now what? Some sort of merge-magic from the registry would be a huge benefit. That'd even fix AMNH's problem - just synonymize their ARK-url with our whatever-we-do with....
I'm starting to wonder if there's much point in doing anything without a registry. AMNH has technical limitations, I think the NEON stuff is going to come up in a system with different limitations, the zoo-people seem to still be working in desktop systems - can anyone talk to us without an interpreter in the middle, and is there any point of us throwing something up if they can't? Build it and they will come?
attributes ...registry...stable
I'm not sure we're on the same page, but if we're talking about someone accumulating data I'd like to see the other way - adding birth date to the registry in such a way that we can access it (and copy it to Arctos so we can query it) would add value to all of the bits-n-pieces, so if that (or anything else of potential value) pops up I'd like to see them grab it so we can grab it.
room for only one identifier
Oh my.
/OrganismID/
I can make up URLs, although I'm not sure where you're going with that.
/oidtype=NK&oidnum=
That seems overly complicated and brings up back to having data arbitrarily in two places and all that jazz.
I need beer....
I need beer....
Ditto
I agree with everything @tucotuco outlined. I guess the real question is WHO is going to do it? We can wait around, or we can take the (individual) bull by the horns.
if we're talking about someone accumulating data I'd like to see the other way - adding birth date to the registry in such a way that we can access it (and copy it to Arctos so we can query it) would add value to all of the bits-n-pieces, so if that (or anything else of potential value) pops up I'd like to see them grab it so we can grab it.
It should be a two-way street and yes, we will need some way to merge stuff when it is determined that two registered organisms are actually the same thing.
BUT
Before we jump into creating a registry, maybe we can do a not-so permanent version as proof of concept?
And as for AMNH and Emu...sheesh
two-way street
Arctos has all sorts of ways to get at those data, and we could send them to DWC via dynamicProperties or WHATEVER. If we're a bottleneck it should only be because we don't know we're a bottleneck.
not-so permanent
Evil...
I could build something that's minimally-functional and that has no reason (other than the 'ownership' of the IDs) to not be permanent/transferable fairly quickly. Could GBIF (or someone) be convinced to use it to find "OrganismID synonyms"? I'm not sure that's necessary functionality, but it would make it a lot shinier if we're eg seeking funding to make it less-minimal, and it would make it work from the AMNH specimen without them doing anything.
I could build something that's minimally-functional and that has no reason (other than the 'ownership' of the IDs) to not be permanent/transferable fairly quickly.
I'm down
I was actually just thinking that if we had "OrganismID" as an other ID and passed the AMNH catalog number there that GBIF should be able to figure it out (maaayyyyybeee). It would be interesting to see what happens with that since it seems like we are currently the only people capable of passing that field to anyone.
But I would prefer to set it up right and make everyone else realize they need to do better.
Maybe in the case if tissues one place and voucher elsewhere, the organism
ID should reflect the voucher?
In the other hand, I still think it should be the one number that links
them all, eg NK etc
On Thu, Mar 21, 2019, 6:33 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
I could build something that's minimally-functional and that has no reason
(other than the 'ownership' of the IDs) to not be permanent/transferable
fairly quickly.I'm down
I was actually just thinking that if we had "OrganismID" as an other ID
and passed the AMNH catalog number there that GBIF should be able to figure
it out (maaayyyyybeee). It would be interesting to see what happens with
that since it seems like we are currently the only people capable of
passing that field to anyone.But I would prefer to set it up right and make everyone else realize they
need to do better.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-475428353,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hCZxq8erqDL6qMdey_w123TB3kVcks5vZAi1gaJpZM4buGmY
.
I was actually just thinking that if we had "OrganismID" as an other ID
and passed the AMNH catalog number there that GBIF should be able to figure
it out (maaayyyyybeee).
For every Occurrence record you will be able to have exactly one
dwc:organismID. Not because the that organismID identifies the record, but
because every Occurrence can only have one participation dwc:Organism. You
are talking about multiple dwc:OrganismIDs for an Organism. That's fine to
do, but you would have to share that information in a different way, and
no, GBIF would not "figure it out" automatically. The only thing they can
"figure out" is that two dwc:organismIDs are equal.
The different way would be one or both of a) populate
dwc:associatedOrganisms with the list of dwc:organismIDs, and b) create
ResourceRelationship records showing the actual relationships (including
sameAs) between the Organisms identified in the ResourceRelationship
records.
On Thu, Mar 21, 2019 at 7:33 PM Teresa Mayfield-Meyer <
[email protected]> wrote:
I could build something that's minimally-functional and that has no reason
(other than the 'ownership' of the IDs) to not be permanent/transferable
fairly quickly.I'm down
I was actually just thinking that if we had "OrganismID" as an other ID
and passed the AMNH catalog number there that GBIF should be able to figure
it out (maaayyyyybeee). It would be interesting to see what happens with
that since it seems like we are currently the only people capable of
passing that field to anyone.But I would prefer to set it up right and make everyone else realize they
need to do better.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-475428353,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AAcP6_e7RloBiYO-TEY91KJiLaNduwSGks5vZAi2gaJpZM4buGmY
.
Maybe in the case if tissues one place and voucher elsewhere, the
organism ID should reflect the voucher? In the other hand, I still think it
should be the one number that links them all, eg NK etc
One ID to rule them all didn't work for Sauron. What hope does biodiversity
have?
Seriously though. You are trying to solve a social problem here. It won't
work. The best we can hope for is to be able to say, "These are [derived
from] the same Organism", where it might be reasonable to elaborate on
[derived from].
Yea all agreed - "their" catalog number (if you mean the ARK) will do something, but not what we want and only in very limited circumstances.
NK caused massive problems in a closed and controlled environment, as DGR so painfully demonstrated. I doubt that gets better at scale!
Agreed re: this is a social problem, and it can't be just OUR social problem - if nobody else cares or has the technical capacity to care, then this starts to look like more work to solve a problem that already has a local solution.
So here's an idea:
We build a resolver/synonimizer-er, or sucker someone else into building a resolver/synonomizer-er. It provides an identifier, which we (and hopefully others) use as OrgID. That identifier, which anyone can use, resolves to a service, which provides a list of IDs that are hopefully in GBIF et al. GBIF uses the IDs to round everything up.
If GBIF can't do that, the OrganismID would still lead to a list of links to "components" and could still be used by anybody. (And it could pull everything from GBIF-er-whatever, with development, if we have to.)
Can we already approximate the rounding-up through some sort of DWC-magic? If not, anyone want to run it by GBIF/iDigBio/whatever and see if it's a possibility? Maybe just get them to build it....
The different way would be one or both of a) populate dwc:associatedOrganisms with the list of dwc:organismIDs, and b) create ResourceRelationship records showing the actual relationships (including sameAs) between the Organisms identified in the ResourceRelationship records.
We do this already (sort of)! For those pesky AMNH specimens, we have the Other_ID_Type = AMNH and any time it is used, we assign a relationship (same organism as) and an identifier (the AMNH catalog number) which as GBIF does things now is also the organismID (if AMNH will just pass it that way).
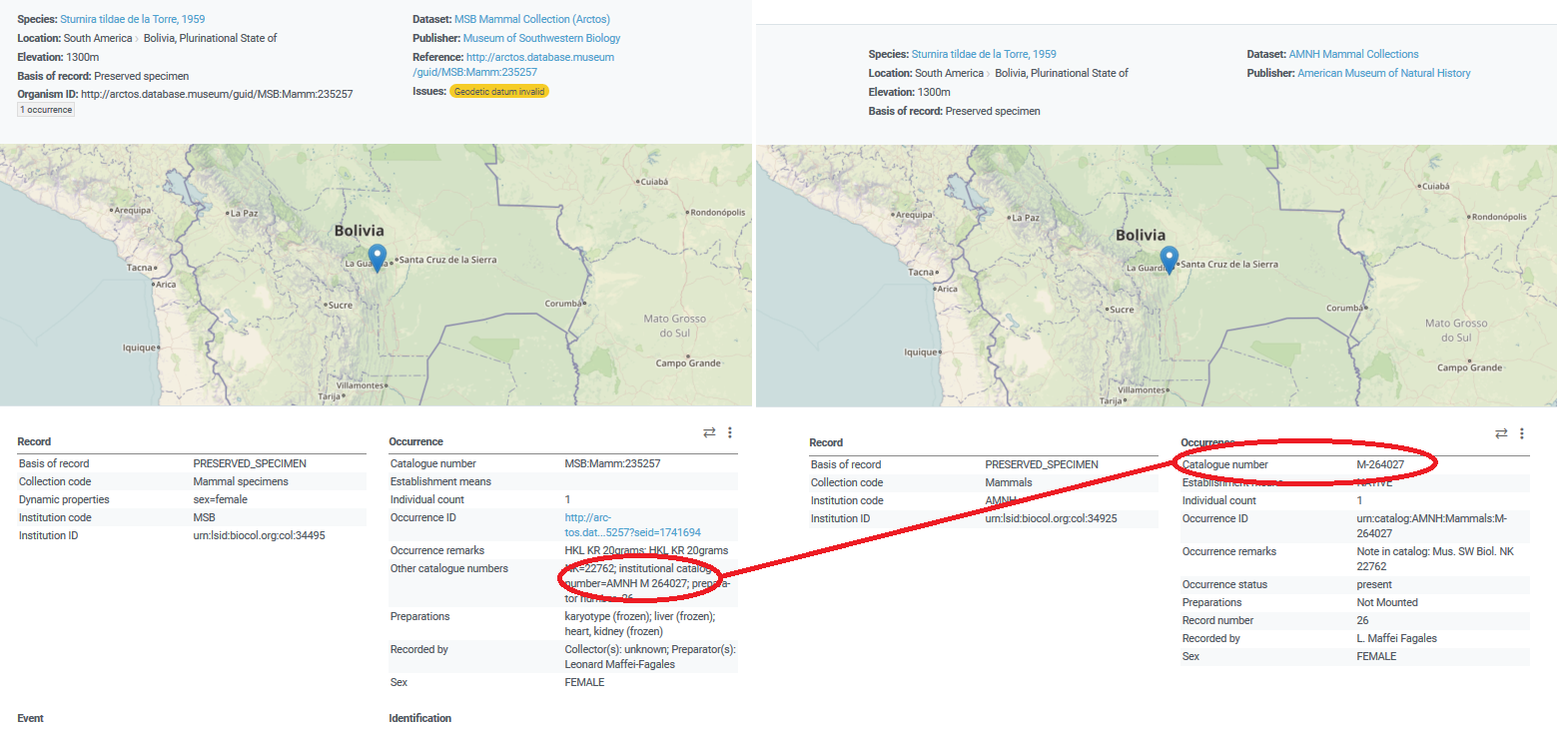
(also note that they DO know there are tissues at MSB....BUT they have the NK, which isn't very helpful)
If we would pass the Other_IDs as dwc:associated Organisms and include the relationship as Resource Relationship, then we would at least be telling everyone "this is the same organism". We would need to clean up the AMNH catalog numbers in Arctos so that they are in the proper format "AMNH:Mammals:M-264027"? as opposed to "institutional catalog number: AMNH M 264027" AND we would need to get AMNH to pass their catalog number to OrganismID (or work with GBIF so that it's there?)
This seems like a potential start at getting the collections world to think about how the data that is being aggregated needs to be better assimilated?
This sounds like a plan.
On Fri, Mar 22, 2019, 11:50 AM Teresa Mayfield-Meyer <
[email protected]> wrote:
The different way would be one or both of a) populate
dwc:associatedOrganisms with the list of dwc:organismIDs, and b) create
ResourceRelationship records showing the actual relationships (including
sameAs) between the Organisms identified in the ResourceRelationship
records.We do this already (sort of)! For those pesky AMNH specimens, we have the
Other_ID_Type = AMNH and any time it is used, we assign a relationship
(same organism as) and an identifier (the AMNH catalog number) which as
GBIF does things now is also the organismID (if AMNH will just pass it that
way).[image: image]
https://user-images.githubusercontent.com/5725767/54834729-5f514100-4c86-11e9-8dce-e7746b0efe7d.png(also note that they DO know there are tissues at MSB....BUT they have the
NK, which isn't very helpful)If we would pass the Other_IDs as dwc:associated Organisms and include the
relationship as Resource Relationship, then we would at least be telling
everyone "this is the same organism". We would need to clean up the AMNH
catalog numbers in Arctos so that they are in the proper format
"AMNH:Mammals:M-264027"? as opposed to "institutional catalog number: AMNH
M 264027" AND we would need to get AMNH to pass their catalog number to
OrganismID (or work with GBIF so that it's there?)This seems like a potential start at getting the collections world to
think about how the data that is being aggregated needs to be better
assimilated?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-475673421,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hLhqy39ZEdkbdkBJS_Da5JPTs0Cvks5vZPu2gaJpZM4buGmY
.
"These are [derived from] the same Organism", where it might be reasonable to elaborate on [derived from].
I think we can just use "same individual as" (as opposed to self) and use the GBIF search to connect the two? See http://arctos.database.museum/guid/MSB:Mamm:235257
Would (or could) that create:
associatedOrganisms = "same individual as", AMNH:American Museum of Natural History https://www.gbif.org/occurrence/search?institution_code=amnh&catalog_number=M-264027"
OR could we use the same idea to create:
associatedOccurrences = https://www.gbif.org/occurrence/search?institution_code=amnh&catalog_number=M-264027
I don't know how these particular fields are transmitted to the ipt, but it seems like we could make this work if we structure the data correctly...
AND based upon OUR link, GBIF could create the reciprocal?
use "same individual as" (as opposed to self)
Yes, that's what should have been happening - http://arctos.database.museum/info/ctDocumentation.cfm?table=CTID_REFERENCES
self--> current cataloged item
everything else--> a cataloged item
I'm mapping relationships to RELATEDCATALOGEDITEMS - I'm not sure what happens downstream from that.
To back way up, I think we have three options here:
- do something 'local' and, at best, redundant with existing IDs (eg, try to make links with human-assigned IDs)
- do something duct-tapey (eg, try to trick GBIF into doing something)
- do something capable of solving a major issue in bioinformatics (eg, build a resolver)
Can we do the first two and work towards the third?
On Fri, Mar 29, 2019, 9:18 AM dustymc notifications@github.com wrote:
use "same individual as" (as opposed to self)
Yes, that's what should have been happening -
http://arctos.database.museum/info/ctDocumentation.cfm?table=CTID_REFERENCESself--> current cataloged item
everything else--> a cataloged itemI'm mapping relationships to RELATEDCATALOGEDITEMS - I'm not sure what
happens downstream from that.To back way up, I think we have three options here:
- do something 'local' and, at best, redundant with existing IDs (eg,
try to make links with human-assigned IDs)- do something duct-tapey (eg, try to trick GBIF into doing something)
- do something capable of solving a major issue in bioinformatics (eg,
build a resolver)—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-478036447,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hD2PsU7CET-uLZvKBKIwy8Jikhi1ks5vbi6rgaJpZM4buGmY
.
I think you already have!
You can add an organismID to http://arctos.database.museum/info/ctDocumentation.cfm?table=CTCOLL_OTHER_ID_TYPE if you want, but it's not clear to me what it can do that existing IDs cannot.
I'm mapping relationships to RELATEDCATALOGEDITEMS - I'm not sure what happens downstream from that. @dbloom ?
do something 'local' and, at best, redundant with existing IDs (eg, try to make links with human-assigned IDs) do something duct-tapey (eg, try to trick GBIF into doing something) do something capable of solving a major issue in bioinformatics (eg, build a resolver)
I think what I am proposing is solution number 2, but I am hopeful that it will start a conversation that could move us on to number 3.
I loaded new IDs for all of the MSB Mamm records that had an original identifier that included AMNH - I did not remove those original IDs, but we probably should eventually as many of them are fomatted incorrectly. When is the next update to GBIF? I'd like to see what happens.
Is it possible to have a code table for the prefix for organism IDs and
create a dropdown? For example, the ID type would be Organism ID, the
prefix could be "Mexican Wolf Studbook Number" or "AMNH (American Museum of
Natural History)" and the integer = 1216 etc. This would help standardize
and eliminate formatting problems.
On Fri, Mar 29, 2019 at 9:29 AM dustymc notifications@github.com wrote:
I think you already have!
You can add an organismID to
http://arctos.database.museum/info/ctDocumentation.cfm?table=CTCOLL_OTHER_ID_TYPE
if you want, but it's not clear to me what it can do that existing IDs
cannot.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-478041109,
or mute the thread
https://github.com/notifications/unsubscribe-auth/AOH0hKGdIQFyFICkDYhjIvgU5sDdGYwLks5vbjFxgaJpZM4buGmY
.
New Idea:
I was thinking about this in the context of Mexican Wolves last night. Could we create a project for each wolf and get a doi for the project, then use the doi as the organism ID?
doi for the project
Check out Archives. http://handbook.arctosdb.org/documentation/archive.html
HMMMM. Sounds appropriate, but we may need to discuss locked vs.unlocked...
OK @dustymc I experimented. I created a locked archive for Mexican Wolf Studbook number 1294 and obtained a DOI. Now, can I put that DOI in an other ID called "organism ID" or is there some other way I should indicate that? The DOI is http://dx.doi.org/10.7299/X7WD40WN and it takes me right to the Arctos archive, but shouldn't each record now have that doi added?
Also, next week when we get a new batch of blood, can I add it to the archive? That is very important....
Oh wait, I see that is not an option. So the archive doesn't seem to be a good way to do this....
Also, can you undo everything I just did?
locked vs.unlocked
Locked are immutable (ish...). They're designed for "looked at these things for this publication, don't want them to wander off or the IDs to be recycled or something."
That's not this. You should expect to find other components of the individual scattered all over the place; these data are KNOWN to be dynamic, that blood you'll get next week requires changing the Archive. Unlocked Archives are appropriate here.
Yes you can add (to unlocked) - it's in the popup.
'myarchive' will create a new archive (or fail); '+myarchive' will append to your existing archive 'myarchive' (or fail).
I suppose the DOI could be added to the specimen record, but it's not an identifier for the specimen record either - it's an identifier for something that contains the specimen record. We could show archives which contain the specimen on the specimen detail page - that's certainly structurally more appropriate (eg, it's normalized).
DOIs are nice here in that if someone builds a real resovler we can just redirect the DOI to it. I think my only hesitation in allowing them on unlocked Archives is in the idea that DOIs should be permanent; they should come with some sort of maintenance commitment. Maybe that's just a matter of disallowing deletion of unlocked DOI-bearing Archives? In any case I think we can find something that both allows DOIs on unlocked Archives and provides some sort of stability to the identifier.
Maybe we should plug that into ARK (="DOI light") instead??
I took a fairly serious approach to it when I designed locked archives; I can't readily unlock them any more than anyone else can. I don't think there's any harm in just abandoning this and creating a new unlocked archive; if you disagree I can dig out the scary password, lock everything down, disable all of my security stuff, and unlock the archive sometime when Arctos isn't busy.
I don't think there's any harm in just abandoning this and creating a new unlocked archive; if you disagree I can dig out the scary password, lock everything down, disable all of my security stuff, and unlock the archive sometime when Arctos isn't busy.
The way things go, there is probably going to be someone who will want to delete or encumber one of these records. UGH I'm sorry!!!! But can you do that for me? I'll owe you big time and I will just give up on this until someone wants to do it right.
encumber
Yea, good reason. You're unlocked, don't think anyone noticed, no problem.
The DOI should still work. Here's an ARK too, because: https://n2t.net/ark:/87299/x67h1gmh / https://ezid.cdlib.org/id/ark:/87299/x67h1gmh.
Oh man! That's a load off my mind - I'm buying the next time we are in the same bar.
And now for what I woke up thinking about....
Isn't what we need an Agent? If this wolf were an agent, we could do all the appropriate relationships with other wolves, add identifiers from other institutions (like ORCids), provide birth and death dates, etc. And add the agent to all of the records for that particular wolf.
If we added the comment/report bad data to the Agent pages, then someone outside of Arctos with information about some part of a particular organism could let us know and we could add the "address".
So, would something similar to an agent work? Would we need a separate agent-like table or could we just add a new type of agent (organism)? Could this agent name appear on the specimen pages just below the catalog number?
I'm sorry that I am obsessing over this, but it is bugging the heck out of me right now for various reasons....
Agents
That seems to be stretching our model in strange ways for very limited utility. If you can't get AMNH to enter "Mexican wolf studbook number 1294" in a field meant for that sort of thing, I'm not sure why they'd enter it (or some agentish proxy to it) in a field intended for something else?
Were I them, I wouldn't enter "Mexican wolf studbook number 1294" either - it has incredibly limited utility even if it wasn't used on 20 wolves (and 3 aardvarks for some reason) after The Great Printer Jam Of '04. I might agree to add http://dx.doi.org/10.7299/X7WD40WN, which I know to be unique and resolvable, if it would do something for me.
What I proposed above is something like the agents model, except there are no preferred names. https://n2t.net/ark:/87299/x67h1gmh, http://dx.doi.org/10.7299/X7WD40WN, https://arctos.database.museum/guid/MSB:Mamm:267935, Mexican wolf studbook number: 1294, NK:226608, NK226724, NK=226735, NK: 226893, etc., are all "names" for one entity. They have various intent, capability, trustworthiness, and functionality. Anyone could add another "name," anyone could use existing names for any purpose, and everyone should be encouraged to use the resolvable names intended to identify the composite organism to do just that wherever they can. (OK, maybe there is a "semi-preferred name:" one would allow assembling the Organism from eg, GBIF downloads without passing through a service. Problem is, I don't think there's a way to get everyone to use it, and it would be misleading to pretend it's something it can't be without that.)
ORCids
I think that's the useful analogy, in its relationship to "names," its political position, its functionality, etc. It doesn't quite hold up because while there's only one entity allowed to ask for a unique way of identifying THAT John Doe, there are any number of entities which might ask for a unique way of identifying THAT Mexican wolf studbook number 1294. (And maybe I suddenly understand why ORCID won't help us with deceased collectors...) While being able to say "this is THE one true OrganismID for THAT wolf" is nice in theory, I just don't think it's remotely practical due to the way identifiers are acquired.
I'm still pretty confident that I know how to build this and to make it DO STUFF. I'm much, much less confident I know how to get anyone to use it, especially if it's associated with Arctos (which a huge percentage of "the collections community" seems to view as complicated Excel....). I think realistically this needs built as it's own entity (like ORCID), or on top of some existing broad-scope "neutral" entity (perhaps GBIF).
While being able to say "this is THE one true OrganismID for THAT wolf" is nice in theory, I just don't think it's remotely practical due to the way identifiers are acquired.
So we need the WikiData model, like Bloodhound uses for deceased people.
I'm still pretty confident that I know how to build this and to make it DO STUFF. I'm much, much less confident I know how to get anyone to use it, especially if it's associated with Arctos (which a huge percentage of "the collections community" seems to view as complicated Excel....). I think realistically this needs built as it's own entity (like ORCID), or on top of some existing broad-scope "neutral" entity (perhaps GBIF).
OK, so let's talk about cost, platform and hosting. Seems like we could get TACC on board for the hosting. Give me some numbers and then I'll see what I can do about scraping up funding.
WikiData
Yea that's why I added it to http://arctos.database.museum/info/ctDocumentation.cfm?table=CTADDRESS_TYPE
funding
Let's chat. AWG meeting? Maybe TACC could be persuaded to play a more central role?
https://scicrunch.org/resources might be a way to do this?
Big-picture, I still think that there are two critical components:
- Some sort of synonomizer, to unify the inevitable "2 users get an ID for their bits of the same rat" situation, and
- Community buy-in. The idea can do little/nothing within Arctos, it's only useful if eg AMNH starts applying (and sharing in a useful way) shared/synonomized identifiers to their bits of "our" rats.
RRID seems to be aimed in the right direction and actually doing stuff.
iSamples seems a bit theoretical and I can't tell what they're thinking, but there are some familiar names and they've got a lot of $$. Convincing the larger community that this is useful/critical may be the big hurdle, and perhaps they're better positioned to do that.
See also https://github.com/tdwg/dwc/issues/314#issuecomment-824890294
Thanks, @Jegelewicz, for reviving this. This issue is becoming critical. We can't wait around to build some external feature. We need something that will work within Arctos now, something that we can designate as a dwc organism ID and export to GBIF.
Currently just at MSB in Arctos, we have the Mexican Wolf Recovery Program, which has more a decade now generated a continuous flow of blood, serum, and carcass samples for hundreds of wolves all with repeat sampling, using the Mexican Wolf Studbook number ID. This dataset has some cataloged items as single wolves with multiple events, and some that are have separate catalog numbers for the same wolf. It is this dataset that both led to and then revealed the extreme shortcomings of the multiple event model that led to the start of this discussion.
We have tissue samples at MSB and vouchers at AMNH (and in Bolivia) linked to each other by an admittedly imperfect field number, the NK number.
We have all legacy NEON mammals linked by a NEON: National Ecological Observatory Network: ID which MSB minted (NEON didn't have any such thing, because they were thinking of and only had IDs for samples, not organisms, like GGBN) to act as a defacto organism ID. And yes, there are many examples of problems with this dataset, going back to shortcomings in the original data management and field collection at NEON, which should have used scannable barcodes/ uuids/guids rather than things like eartags.
As a result of a recent MOU, MSB now also has thousands of zoo animal blood/serum/carcasses collected over many years, many of which are multiple occurrence records for the same animal. Same problem here too with lack of an a priori unique "organism ID" - there are instead organism identifiers assigned by the zoo community, some of them shared across multiple institutions, for example a "GAN = global animal number". This is tracked in their ZIMS database, where we are downloading data from in order to import to Arctos. Some zoos use other databases with other identifiers, so there is no centralized repository or authority, and of course duplication and transcription error occurs.
Regardless of the known limitations of these existing versions of "organism IDs" - e.g Mex wolf studbook numbers, NKs, NEON numbers, GANs, etc. - and the desirability of something more reliable like a doi, Ark, etc - these are WHAT WE HAVE. Yes, there are going to be duplicates, problems, parts linked to the wrong specimens, etc etc. But we cannot discover these problems until we make the attempt to start linking them, and make these linkages discoverable both in Arctos and in GBIF.
Can we please move forward with a means of designating an ID in Arctos as an "organism ID" which each collection can decide to use or not use, that will be published to GBIF? I need these immediately at MSB. I don't want to load another several thousand samples for yet another project without some means of linking related records via an organism ID, because I am cataloging each occurrence separately, abandoning the multiple occurrence/single catalog number method as a result of the unresolved limitations discussed above.
@mkoo
I think the issue at TDWG makes an important point about distinguishing between an occurrence, a material sample and an organism and we should think carefully about that before we leap. The Mexican wolves are one kind of problem (multiple occurrences of a known individual sometimes with multiple material samples from each occurrence) and the MSB/AMNH is another (multiple material samples from a single occurrence scattered around in various collections). Can a single solution work for both of these scenarios?
THIS - https://github.com/tdwg/dwc/issues/314#issuecomment-825088348
Per our discussion today, I've set up some examples in test.
Organism Profile example - http://test.arctos.database.museum/info/agentActivity.cfm?agent_id=21327831
Advantage to this is all you have to do to connect the records is add the organism profile name. Relationships can also be made via the organism profile so that you don't have to add "offspring of" to EVERY record. Media associated with catalog records is automatically associated with the profile. Other identifiers associated with the organism are easily added to the organism profile (just like we add OrcID, Github name etc.) Passing the profile url as "OrganismID provides a way for anyone outside of Arctos to determine if they have the same organism. Disadvantage is denormalization of relationships, but we could eliminate that if we create organism "profiles" for all of these kinds of things or other ideas?
Project example - http://test.arctos.database.museum/project/10003362
Advantage - connects parts to usage. Disadvantage - every new sample has to be added to the project data loan. All relationships have to be created for every new sample as well as the old samples when a new one comes in. Project name is not connected to the records in such a way that allows us to pass the project url as "organismID".
What else could we do with these scenarios? What is just really dumb?
@campmlc @ebraker
"OrganismID" is clearly an identifier, not an agent or geography or .... If someone wants an identifier, just make and use one.
We don't need to break random things to create new views of the data. Use some resolvable (eg unique) value for the identifier (ARKs are still convenient) in the expected place and we can build whatever we want around it.
Disagree. The identifiers concept is extremely limited when you are trying
to related 100 fish in a lot, or 100 parts of 1 wolf collected over 10
different collecting events. The neat thing about using something like
"agents' (and we don't have to call it that - we just need a similar
structure) is the richness of the info that it provides, including but not
limited to relationships.
On Thu, Apr 22, 2021 at 9:56 PM dustymc @.*> wrote:
- [EXTERNAL]*
"OrganismID" is clearly an identifier, not an agent or geography or ....
If someone wants an identifier, just make and use one.We don't need to break random things to create new views of the data. Use
some resolvable (eg unique) value for the identifier (ARKs are still
convenient) in the expected place and we can build whatever we want around
it.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-825367860,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBDIG7CDAWQEZWXARYTTKDVXDANCNFSM4G5YNGMA
.
I don't think there's anything subjective about this; it's just the wrong tool for the job.
"Similar structure" is another matter entirely - everything in Arctos is always subject to better ideas, we can certainly talk about rebuilding OtherIDs. (But those ARE relationships, so I'll need some elaboration.)
I think one huge problem we are sticking a bandaid on is related to the material sample discussion at TDWG. Are the 5 vials of blood in one record the "parent of" the 6 vials in another one? NO, they are EVIDENCE of that relationship. The wolves the blood came from have the parent/child relationship and we are NOT cataloging the wolves. It seems to me that the parent/child relationship is in the wrong place.
It may or may not be different for "this flea is parasite of that mouse", but I kinda don't think it is. We probably have the flea in it's entirety, but it is far less likely that we have the mouse, we only have EVIDENCE (parts) of the mouse. At some point this may also be true for the flea. We are mixing up organisms and parts in the way we catalog things and that causes these problems.
Somewhere along the way, what we used to think of as "specimens" now became "occurrences", as if they were congruent concepts. But of course, specimens are physical entities with all sorts of properties important to the people who care for them (such as preparations, disposition, etc.), whereas (as @dshorthouse already noted) occurrences are ephemeral things, capturing the abstract idea of an Organism being present in the context of an Event.
Thank you @camwebb but why haven't we all read this paper?
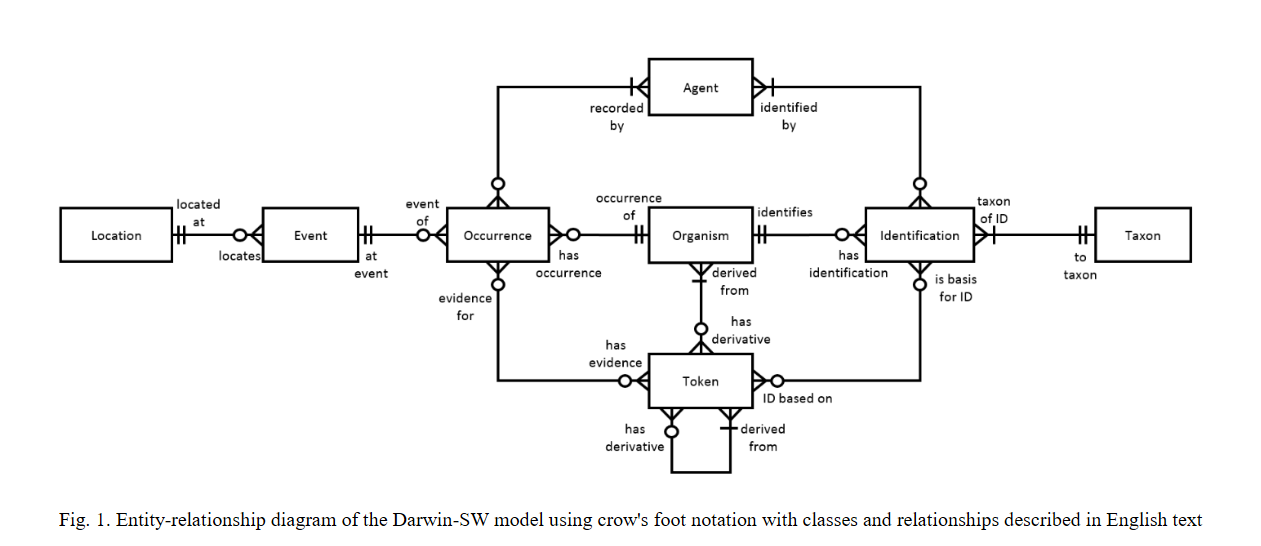
I think we are conflating organism and token.....
Are the 5 vials of blood in one record the "parent of" the 6 vials in another one?
We do not assert that. They are "children" (database term, nothing else implied) of things that may be in relationships (bio included) with each other. Seeing a vial of blood with wolf DNA in it as evidence for the catalog record representing a wolf in some fashion is entirely consistent with the Arctos model.
we are NOT cataloging the wolves
We are very explicitly avoiding that question - https://handbook.arctosdb.org/documentation/catalog.html#understanding-cataloged-items. From the philosophical perspective, we are cataloging whatever someone wants to think we're cataloging. From the data design perspective, we are cataloging ephemeral THINGS representing WHATEVER. Anything "real" (parts) is dealt with elsewhere.
We probably have the flea in it's entirety,
That is not something we could or would assert - "entirety" could mean anything (we probably don't have it's exuvia or offspring or ...), we can only assert what we do have (which can include nothing physical).
We are mixing up organisms and parts in the way we catalog things and that causes these problems.
See above - we are not, and that's not possible in our model. Elaborate please - I am obviously missing something.
what we used to think of as "specimens" now became "occurrences",
We do not. We map "specimens" to Occurrences, but not very accurately.
And I won't argue that we should be doing otherwise - Occurrences are useful metrics, "things someone felt like slapping a catalog number on" are less so. DWC, like any good exchange standard, standardizes. Arctos, like any good CMS, is much more general. (Probably - whatever you think of Arctos and/or DWC, they are unequivocally DIFFERENT.)
specimens are physical entities
See above - this simply isn't true in Arctos; we (stretching all the way back to - beyond, probably - Barb Stein) have very carefully avoided mixing these concepts.
occurrences are ephemeral things
Most catalog records in Arctos represent one Occurrence, then a few represent either a slice of an Occurrence (think co-cataloged stuff) or multiple Occurrences (thing eggs and nest parasite). However, zero is entirely possible - Arctos does not require "Event" (or taxa, sorta...) information, and Occurrences can't exist without that.
A third feature of an Organism is that it serves as an anchor point for resources derived from it, such as specimens, images, and samples. So although an Organism can be described in conceptual terms by comparison with biological organisms, from the standpoint of the DSW model, an Organism is a node that connects Occurrences, Identifications, and de-rived resources
A third feature of an Organism is that it serves as an anchor point for resources derived from it, such as specimens, images, and samples. So although an Organism can be described in conceptual terms by comparison with biological organisms, from the standpoint of the DSW model, an Organism is a node that connects Occurrences, Identifications, and de-rived resources
I have no argument with that, but it's also mostly irrelevant for Arctos: Our model explicitly allows cataloging the "Organism" or cataloging "samples" (or both, or something else, or whatever). For some of those options, it's useful (for some users, probably) to reassemble the bits into "Organisms" - the subject of this discussion.
http://test.arctos.database.museum/organism/ark:/99999/fk4c26bq3x is a quick-n-dirty demo of the "shared identifier" method of linking bits together. It's just a shared ID (one that's known to be globally unique, is capable of carrying metadata, and has an external resolver in this case, but that's not necessary at all) with a simple handler, should return something like...
and those two catalog records are (or should be, I'm just grabbing random stuff right now) the "components" of the organism. There's no metadata of the identifier itself, but that could be changed to a certain extent by revising the otherID model.
http://test.arctos.database.museum/guid/UAM:Mamm:12 is a QnD demo of the "just catalog everything" approach. That catalog record represents (poorly, because I just picked something random) the Organism, these related catalog records....

(if they exist, I didn't bother checking) represent "Occurrences" (or something like it).
The first approach is super easy and there are tools for everything NOW. It's entirely limited to Arctos; there's no way to fetch an "Occurrence" stored elsewhere.
The second approach is more complicated (but we can always build more UI) but is capable of carrying extensive metadata, of acting as an independent record in its own right, and is not limited by the boundaries of Arctos - it can "include" (by linking, minimally) Occurrences in other systems. It's also probably likely to end up representing 16 of the "we looked at these 20 wolves and...." in some paper, but maybe that ain't our problem.
My ideal solution would look a bit like the second option, but exist somewhere outside of Arctos and be accessible to anyone who has an "Occurrence" (or other organism derivative). We'd probably fire up an "Organism Collection" if we went this route, so that's probably sorta-possible within Arctos as well, but I doubt the politics are enticing to some KE user.
I'm sure there are lots of other ways to accomplish this (none involving things that aren't identifiers!), those two are just what I could do in a few minutes without making any significant changes to anything.
An occurrence (e.g., each instance of the capture of an individual)
This situation inevitably leads to confusing citations and bad science when an individual sampled multiple times at multiple locations is assumed by users to be multiple distinct individuals. Arctos supports cataloging encounters as specimen events under one cataloged item.
This statement has all kinds of problems. Our bandaid system for connecting parts with events is not functional and it introduces confusion. I will never approve of this method until a catalog record can have multiple accession numbers and parts are associated directly with the events that created them.
Did we know about this? http://bioguid.org/about
connecting parts with events
Oh, that. It is functional (if limited), and it's potentially a nice first babystep on the path to an event-based model. We need to keep stepping, one way or the other. I'm not convinced (anymore, maybe) that there's a "correct" way to go from here, but backwards (and then towards "organisms" in some way, probably) has a much lower bar.
Oh, that. It is functional (if limited), and it's potentially a nice first babystep on the path to an event-based model.
I don't think museums are interested in an event based model. They are interested in caring for the physical objects entrusted to them. This is where Arctos is not serving the user community well and instead striving to satisfy the aggregators who want an "occurrence". I don't believe that tracking organisms separately from cataloged items is "going backwards" - I actually think it is making a decision about who Arctos is serving, the collections who use it or the aggregators. I also think that in the long run, we can please both the collections and the aggregators if we separate (or at least allow separation of) "organisms" from "cataloged objects".
I agree with Teresa. We also need to make this model functional for
collection staff. The current event based model is not, for multiple
reasons repeatedly described, but mostly because of its complexity. Making
it even more complex with an even bigger learning curve in order to deal
with its shortcomings is not the answer. Cataloging collection objects
separately as we always do, but providing a means to tie them to an
organism, and then see the "organism" as we see agents, would be a huge
improvement.
On Sat, Apr 24, 2021, 6:51 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
Oh, that. It is functional (if limited), and it's potentially a nice first
babystep on the path to an event-based model.I don't think museums are interested in an event based model. They are
interested in caring for the physical objects entrusted to them. This is
where Arctos is not serving the user community well and instead striving to
satisfy the aggregators who want an "occurrence". I don't believe that
tracking organisms separately from cataloged items is "going backwards" - I
actually think it is making a decision about who Arctos is serving, the
collections who use it or the aggregators. I also think that in the long
run, we can please both the collections and the aggregators if we separate
(or at least allow separation of) "organisms" from "cataloged objects".—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-826088361,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBCNE2XEIQHV5UQVGPDTKK5F3ANCNFSM4G5YNGMA
.
From https://github.com/ArctosDB/arctos/issues/2085#issuecomment-829719822
Yes, so this "NK 12345"in Arctos is probably that "NK12345" at GBIF
which is supposedly AMNH:Mamm:678910.
In this case, the NK number, or the GBIF occurrence, are the best proxies
we have for organism, as inaccurate as that could potentially be. I think
the NK needs to be designated as the organism identifier, because that is
what ultimately will be traced back to specimen tags and field notes - not
the derivative assertions at GBIF or even AMNH.
We need to be able to designate any type of identifier - NK, AF, collector
number, zoo number - as the organism ID. If this is also treated as an
assertion, not a fact, then it can be open to interpretation and
correction, and documenting who and when said what and why.
I'm not sure we can gain anything that would justify development if it all comes back to nonresolvable identifiers.
I do not think there can be a solution involving records controlled by someone who won't or can't talk to us. Have you talked to anyone at AMNH? Would they use Arctos-issued IDs, or provide something we can use, or go halfsies on building something, or help coerce GBIF into building something, or ???????
We need an organism ID first to link everything in Arctos. We've all
already discussed that it would require an external service to link to
other collections, and we don't have that. We already have Lot ID, which
can be shared among individuals from the same lot and allow discovery. We
need the same for organism ID, something we can designate to link and track
network vs binary relationships and enable discovery The tdwg commy is
still grappling with the distinction between organism, occurrence, and
material sample. There is no external solution happening soon.
On Thu, Apr 29, 2021, 7:28 PM dustymc @.*> wrote:
- [EXTERNAL]*
From #2085 (comment)
https://github.com/ArctosDB/arctos/issues/2085#issuecomment-829719822Yes, so this "NK 12345"in Arctos is probably that "NK12345" at GBIF
which is supposedly AMNH:Mamm:678910.
In this case, the NK number, or the GBIF occurrence, are the best proxies
we have for organism, as inaccurate as that could potentially be. I think
the NK needs to be designated as the organism identifier, because that is
what ultimately will be traced back to specimen tags and field notes - not
the derivative assertions at GBIF or even AMNH.
We need to be able to designate any type of identifier - NK, AF, collector
number, zoo number - as the organism ID. If this is also treated as an
assertion, not a fact, then it can be open to interpretation and
correction, and documenting who and when said what and why.I'm not sure we can gain anything that would justify development if it all
comes back to nonresolvable identifiers.I do not think there can be a solution involving records controlled by
someone who won't or can't talk to us. Have you talked to anyone at AMNH?
Would they use Arctos-issued IDs, or provide something we can use, or go
halfsies on building something, or help coerce GBIF into building
something, or ???????—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-829734470,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBASVC7GYEHULFPSVADTLIBULANCNFSM4G5YNGMA
.
The two plausible things I can think of are https://github.com/ArctosDB/arctos/issues/1966#issuecomment-825784676.
As always, a functional requirements document (doesn't have to be fancy) before we discuss the mechanism would be useful.
functional requirements document = http://test.arctos.database.museum/info/agentActivity.cfm?agent_id=21327831
All you have to do to connect the records is add the organism profile name. Relationships can also be made via the organism profile so that you don't have to add "offspring of" to EVERY record. Media associated with catalog records is automatically associated with the profile. Other identifiers associated with the organism are easily added to the organism profile (just like we add OrcID, Github name etc.) Passing the profile url as "OrganismID provides a way for anyone outside of Arctos to determine if they have the same organism.
Disadvantage is denormalization of relationships, but we could eliminate that if we create organism "profiles" for all of these kinds of things or other ideas?
But maybe it isn't denormalization. Any record that includes this agent as "organism" inherits the relationships which can be mapped to other records with the appropriate related "organism". (We auto generate the identifier relationships).
This seems like the most functional answer so far. It immediately allowed
me to identify problems with records associated with this Studbook number,
so I can go in and fix them. Ideally, we'd have "Organism profile" in a
separate pane on the catalog record page, displayed separately - using this
model for an ID, rather than an agent? It shouldn't be "subject" under
agents. If it stay in agents, perhaps" Organism Name"?
On Fri, Apr 30, 2021 at 11:16 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
functional requirements document =
http://test.arctos.database.museum/info/agentActivity.cfm?agent_id=21327831All you have to do to connect the records is add the organism profile
name. Relationships can also be made via the organism profile so that you
don't have to add "offspring of" to EVERY record. Media associated with
catalog records is automatically associated with the profile. Other
identifiers associated with the organism are easily added to the organism
profile (just like we add OrcID, Github name etc.) Passing the profile url
as "OrganismID provides a way for anyone outside of Arctos to determine if
they have the same organism.Disadvantage is denormalization of relationships, but we could eliminate
that if we create organism "profiles" for all of these kinds of things or
other ideas?But maybe it isn't denormalization. Any record that includes this agent as
"organism" inherits the relationships which can be mapped to other records
with the appropriate related "organism". (We auto generate the identifier
relationships).—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-830239733,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBAJBXQETVXN73LRMQLTLLQV3ANCNFSM4G5YNGMA
.
And there would be nothing stopping us from adding an ARK ID or DOI to
this, correct?
On Fri, Apr 30, 2021 at 11:21 AM Mariel Campbell @.*>
wrote:
This seems like the most functional answer so far. It immediately allowed
me to identify problems with records associated with this Studbook number,
so I can go in and fix them. Ideally, we'd have "Organism profile" in a
separate pane on the catalog record page, displayed separately - using this
model for an ID, rather than an agent? It shouldn't be "subject" under
agents. If it stay in agents, perhaps" Organism Name"?On Fri, Apr 30, 2021 at 11:16 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
functional requirements document =
http://test.arctos.database.museum/info/agentActivity.cfm?agent_id=21327831All you have to do to connect the records is add the organism profile
name. Relationships can also be made via the organism profile so that you
don't have to add "offspring of" to EVERY record. Media associated with
catalog records is automatically associated with the profile. Other
identifiers associated with the organism are easily added to the organism
profile (just like we add OrcID, Github name etc.) Passing the profile url
as "OrganismID provides a way for anyone outside of Arctos to determine if
they have the same organism.Disadvantage is denormalization of relationships, but we could eliminate
that if we create organism "profiles" for all of these kinds of things or
other ideas?But maybe it isn't denormalization. Any record that includes this agent
as "organism" inherits the relationships which can be mapped to other
records with the appropriate related "organism". (We auto generate the
identifier relationships).—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-830239733,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBAJBXQETVXN73LRMQLTLLQV3ANCNFSM4G5YNGMA
.
https://en.wikipedia.org/wiki/Product_requirements_document
I suppose at some level Agents are just my "common identifier" approach, and at some level identifiers are just strings.
I'm not entirely certain that every rat that ever lived isn't an Agent; that's probably a standalone question for The Community. Agents and other data objects (like catalog records) do have different "shapes" - having functional requirements would help determine what sort of data object might be capable of meeting your requirements and what isn't. Agents would not easily map to https://dwc.tdwg.org/terms/#organism, if that's part of the requirements.
Agents would not easily map to https://dwc.tdwg.org/terms/#organism, if that's part of the requirements.
@deepreef Might have something to add on this front :)~
I'm not entirely certain that every rat that ever lived isn't an Agent
I'm certain they were/are....
Agents would not easily map to https://dwc.tdwg.org/terms/#organism, if that's part of the requirements.
I am not saying we should add anything but _Homo sapiens_ to the agent table. I am saying the functionality of the agent table is what we need for an organism table.
@deepreef Might have something to add on this front :)~
Do you really want me to "go" there?
For the record: Yes, we track Occurrence records for Homo sapiens as Organisms. It started as a joke, then we realized it might actually work, and ever since we started the practice, it's proven to be extremely useful (and increasingly so).
Where it gets a bit squirrely in the DwC context is how to manage Organizations in this context. For us it's easy, because we regard "Organism" as a subclass of "Individual", which encompasses non-biological instances. But that may be a bridge too far for the DwC community.
But in any case, it's rather amazing how much more powerful a data model becomes when you track objects according to their properties, as opposed to our legacy biases in how we imagine them to be.
Oh, and yes, humans are not the only species of Organism for which it's useful to assign unique names to individuals, so we also solved the "Stumpy" the great white shark issue at the same time.
humans are not the only species of Organism for which it's useful to assign unique names to individuals
That is what this entire issue is about! We are perfectly content giving humans an identifier and adding a bunch of information to it, why not other individuals? The skin, skeleton and blood of Mexican Wolf 200 are not all of Mexican Wolf 200 over all of the years it was alive. Even if we contend that we cataloged everything for Mexican Wolf 200 in MSB:MAMM:9999999, we haven't. Mexican Wolf 200 had a life outside of the pieces we have. It was an agent in its sphere of influence. We have only collected and cataloged part of it no matter what.
I am not saying we should add anything but Homo sapiens to the agent table. I am saying the functionality of the agent table is what we need for an organism table.
AHA!, maybe.
adding a bunch of information
Can you quantify that? Might be a decent approximation of functional requirements.
Same info as currently can be captured by agents?
On Fri, Apr 30, 2021, 1:24 PM dustymc @.*> wrote:
- [EXTERNAL]*
I am not saying we should add anything but Homo sapiens to the agent
table. I am saying the functionality of the agent table is what we need for
an organism table.AHA!, maybe.
adding a bunch of information
Can you quantify that? Might be a decent approximation of functional
requirements.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-830324665,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBH7OV7CYCAEUVMIO23TLL7VNANCNFSM4G5YNGMA
.
See above - people are animals, why wouldn't we need ALMOST all the same info?
No need for most of the name types we use for people. Keep aka.
Rethink "preferred name" and maybe even drop it.
Agent types - doe we need organism types? Wild animal, zoo animal, etc?
agent status - these are all applicable as organism status
relationship - these should be the other ID relationships we have now
address - change this to "identifiers" I think we will need the wikidata identifier but also, this is where we can add the other identifiers given to the organism like "Mexican Wolf Recovery Program Wolf 200". Hopefully we can eventually link up with others when they get on board with digital info.
Same info as currently can be captured by agents?
Aha, again, maybe.
- [preferred name] - unique primary identifier (why: something resolvable is needed, other "names" could all be "collector number one" equivalents)
- [ type] - https://dwc.tdwg.org/list/#dwc_organismScope
- [names] - any number of alternate identifiers (including wikidata IDs, I don't think an address analog for "special names" is needed)
- [name type] - occurrence,organism, parent organism, whatever
- simple "events" (or not, just pull them via the "name" IDs when possible, whatever)
"Names" are inherently potential relationships, I don't see a need for an additional way of doing the same thing
I think that's more or less back to https://github.com/ArctosDB/arctos/issues/1966#issuecomment-475648521, and I still think such a thing should be separate from/bigger than Arctos.
- [preferred name] - https://dwc.tdwg.org/terms/#dwc:organismName unique primary identifier (why: something resolvable is needed, other "names" could all be "collector number one" equivalents)
- [ type] - https://dwc.tdwg.org/terms/#dwc:organismScope
- [names] - https://dwc.tdwg.org/terms/#dwc:associatedOrganisms any number of alternate identifiers (including wikidata IDs, I don't think an address analog for "special names" is needed) or do we want to distinguish between "other names for same organism" and "names of things related to this organism"?
- [name type] - occurrence,organism, parent organism, whatever - I like it, lets us use this for lots and stuff... but no to "parent organism" - that is a relationship created by name....
- simple "events" (or not, just pull them via the "name" IDs when possible, whatever) - https://dwc.tdwg.org/terms/#dwc:associatedOccurrences
I think that's more or less back to #1966 (comment), and I still think such a thing should be separate from/bigger than Arctos.
I could say the same about Agents, but we didn't wait around for someone to create the world agent tool....
That is what this entire issue is about!
Apologies to all: I have actually not read the context of this thread prior to the comment from @dshorthouse , and my understanding of Arctos is limited.. I still haven't had time (sorry again), but it seems like a really interesting discussion related to what we've been dealing with for several years now in our own data management system.
We are perfectly content giving humans an identifier and adding a bunch of information to it, why not other individuals?
I actually look at it from the other direction: we are perfectly content tracking Organisms via Occurrences, so why exclude one particular species of mammal from that infrastructure?. What you describe as "Individuals" is exactly what the dwc:Organism class is intended for. When that new DwC class was being discussed, we went back and forth between Individual (at the time already enshrined within DwC in the form of the individualID term), and Organism. We ended up with Organism, but as I said I see that as a subclass of Individual, which includes non-biological things (some of which also have names, like organizations, ships, etc.)
I am not saying we should add anything but Homo sapiens to the agent table. I am saying the functionality of the agent table is what we need for an organism table.
Exactly! My working data model has two core tables: Agent (representing the abstract entity) and AgentName (representing the 1-to-many names applied to Agents). But after the snickering about our "joke" of treating humans present at Events as simply additional Occurrence instances of Organisms identified as Homo sapiens (BTW, we tag our Identifications to TNUs, so we actually standardize on Homo sapiens L. sec. Wilson & Reeder), suddenly it became clear that our concept of "Agent" was nearly indistinguishable from Organism. MANY different "bonus" functions emerged from this. For example, the code for tracking the path of a tagged whale (from satellite tags) over time is functionally identical to tracking the career of a naturalist as they travelled the world over time. In other words, all kinds of cool questions we hadn't previously thought to ask suddenly had easy answers.
Rethink "preferred name" and maybe even drop it.
Yup! With you there! For years I tracked this, until it became clear that it was a high cost/low reward situation.
@dusty I can't access either of your examples above eg http://test.arctos.database.museum/organism/ark:/99999/fk4c26bq3x i, getting a 404 error on all of my login profiles.
Any way we can move forward with this? I just cataloged over 1000 zoo records representing multiple occurrences of the same organism. I need a way to quickly see all catalog records associated with the same individual GAN number. Same with Mexican Wolf Studbook number. The virtually invisible "Find all" tool needs at least to show all catalog records of an "organism" sharing the same ID - can we at least have that link include the record it was linked from as well as all the records it links to?
This is not semantics, we need actionable solutions, even if imperfect. I don't want this discussion to bog down.
That's not me, I'm @dustymc
The first example is just a special handler for a specific type of otherID; http://test.arctos.database.museum/SpecimenResults.cfm?anyid=ark%3A%2F99999%2Ffk4c26bq3x goes to the same place.
The second is just a catalog record representing the organism, with relationships to the "occurrences" (or "components" of whatever type).
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-475648521 / https://github.com/ArctosDB/arctos/issues/1966#issuecomment-830429632 / https://github.com/ArctosDB/arctos/issues/1966#issuecomment-830450110 / etc is precisely the same functionality as the first example, but
- there's just an "authority" issuing the identifiers, and
- that authority might have some data about the "Organism" (or whatever the composite might be).
Some examples would be useful - I'm not sure what invisible tool you're referring to, and I don't see anything like that entered by you recently.
Apologies @dustymc - made the mistake of replying from email.
The first example shows the Ark ID as an identifier, but this does not link to anything. Ideally, it would be a live link to all the associated records in Arctos and perhaps GBIF - some way of visualizing all records associated with this organism in one place. This is the reason for requesting to use the "agent" model for other identifiers.
Currently we have:
Here is a screenshot of the tool we currently have to find related catalog items. It only works if we use relationships, which I have done here. But the resulting link to specimens does not include a link to the specimen record that the list linked from:
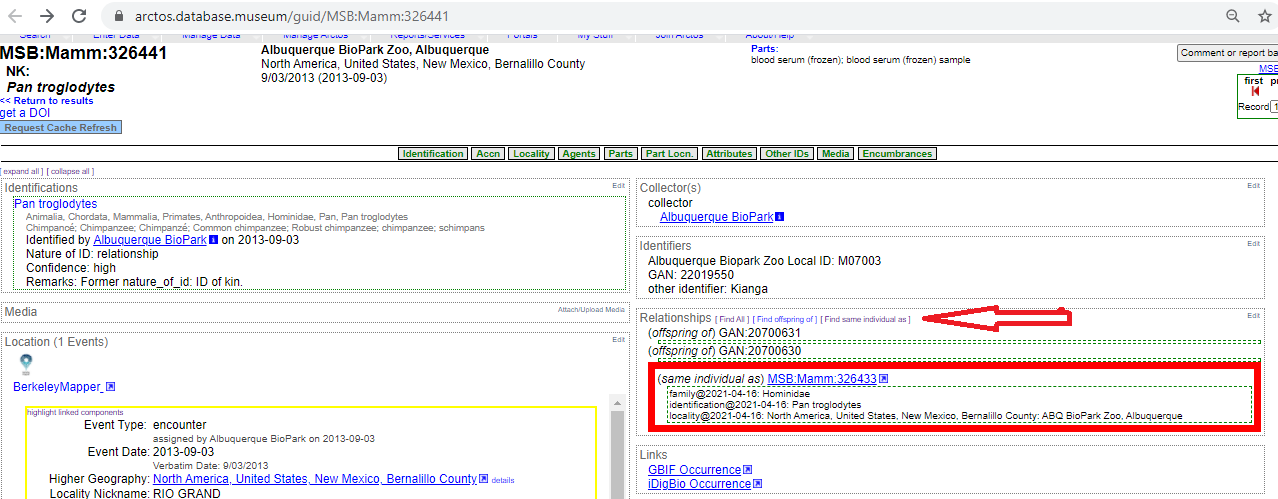
That "Find all" link goes to:
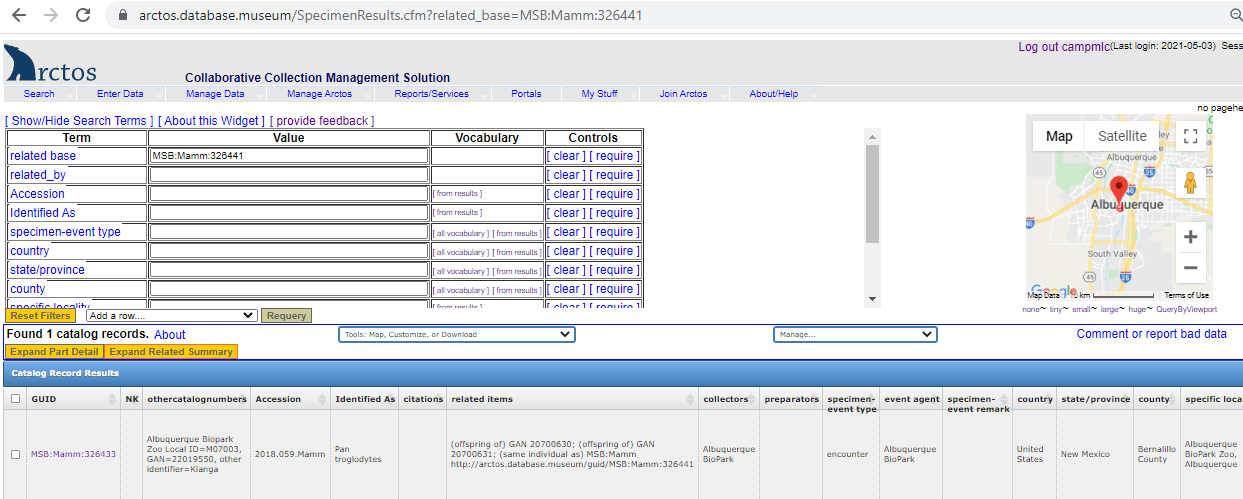
I would prefer the search results show ALL the related specimens, including MSB:Mamm:326441, the related object. And I would like to have this same search be link through a "dwcOrganismID" with the same metadata that we capture for agents. This chimp is really just another agent . . .
Note that the "offspring of" relationships can't show up in the current model, because they have no url. Our original discussion centered around creating a url based on a dynamic search table linked to an ID.
shows the Ark ID as an identifier
The benefit of that is it's unique - it should resolve to the stuff to which you assign it and nothing else. You can use anything for the identifier (only the type is important), just know that there's a whole bunch of stuff out there wearing "1" in some capacity.
live link to all the associated records in Arctos
It is.
and perhaps GBIF
That requires something bigger than Arctos, it's why I keep asking if we can get someone else to do this.
the resulting link to specimens does not include a link to the specimen record that the list linked from:
You haven't used a resolvable identifier. In everything I've proposed, OrganismID would be resolvable (although it'll resolve to a list of semi-random things if you use "1" for the value).
But the resulting link to specimens does not include a link to the specimen record that the list linked from:
It'll resolve to things that bear the identifier. (If not, I need to know about it.)
link through a "dwcOrganismID"
I think the only question is what's at the other end of that link - three things have been proposed so far.
"offspring of" relationships can't show up in the current model
I really need an example; I don't think what you said is how it works.
This chimp is really just another agent
Could be, but that's significant development. (FWIW I don't think they are usefully the same kinds of data objects, although individuals can certainly serve in both roles. We don't much record ear length - not to mention more personal attributes! - for one of those things, nor send loans to the other, for example.)
Can we move forward with this model? In the meantime, perhaps we can get by with and ARK ID or DOI for a saved search on a particular identifier shared between related catalog items of the same organism.
- [preferred name] - https://dwc.tdwg.org/terms/#dwc:organismName unique primary identifier (why: something resolvable is needed, other "names" could all be "collector number one" equivalents)
- [ type] - https://dwc.tdwg.org/terms/#dwc:organismScope
- [names] - https://dwc.tdwg.org/terms/#dwc:associatedOrganisms any number of alternate identifiers (including wikidata IDs, I don't think an address analog for "special names" is needed) or do we want to distinguish between "other names for same organism" and "names of things related to this organism"?
- [name type] - occurrence,organism, parent organism, whatever - I like it, lets us use this for lots and stuff... but no to "parent organism" - that is a relationship created by name....
- simple "events" (or not, just pull them via the "name" IDs when possible, whatever) - https://dwc.tdwg.org/terms/#dwc:associatedOccurrences
I think that's more or less back to #1966 (comment), and I still think such a thing should be separate from/bigger than Arctos.
I could say the same about Agents, but we didn't wait around for someone to create the world agent tool....
move forward
Needs added to AWG agenda, I can work up a better model if you're sure that's the direction you want to go.
get by with and ARK ID or DOI for a saved search
I don't know what you consider getting by. That can do some stuff, can't do some other stuff.
I added this to the agenda for Thursday's issues meeting.
Thanks, all. FYI this example is being presented at an iDigBio Zoos and Museums talk tomorrow by @mkoo.
https://arctos.database.museum/guid/MSB:Mamm:326441
See https://arctos.database.museum/guid/MSB:Mamm:326441 and https://arctos.database.museum/guid/MSB:Mamm:326433 as cataloged records for same individual, but, with only two of the 16 relationships created for all of the following:
https://arctos.database.museum/saved/Kianga%20Organism%20Profile
Actually, this works: https://arctos.database.museum/saved/Kianga%20Organism%20Profile
because it just allowed me to find errors- two of the records are duplicates - cataloged by different people. This is why it is so important to have this kind of overarching view of all interrelated records.
Actually, this works:
That's what I've been trying to say for a while! Maybe we just need a better demo, so....
I slipped my "organism handler" into production, created a new ID type, and changed the type of your 4 records.
https://arctos.database.museum/organism/Kianga
- I got there by asserting IDs, no extra hoopjumping required
- "There" will maintain itself based on identifiers - get a new blood sample, enter it correctly, the link will magically include it
- It's a specific type which exists for a specific purpose, I could ship it ("it" being
https://arctos.database.museum/organism/Kianganot justKianga, I think) to GBIF as OrganismID
The "agent-like model" could/would be implemented exactly like this, but you'd have to jump through some hoops to acquire the identifier and that process would leave some metadata of the identifier that one could access in some way. I'm still pretty baffled as to how that's preventing you from just using this, but perhaps that will fall out of the little demo as well.
so important to have this kind of overarching view
We'd have to discuss how the "Organism Node" gets those metadata, but it's potentially a place where a human could add confusing or conflicting information. What I've done is all live - there can still be conflicts, but they can't take the shape of independent assertions.
If this still works, then you might want to open a new Issue regarding DWC mapping and we can revisit doing something more complicated via proposal.
If it doesn't I'll fix your 4 records, delete the new ID, and apologize for my sins.
When I am here
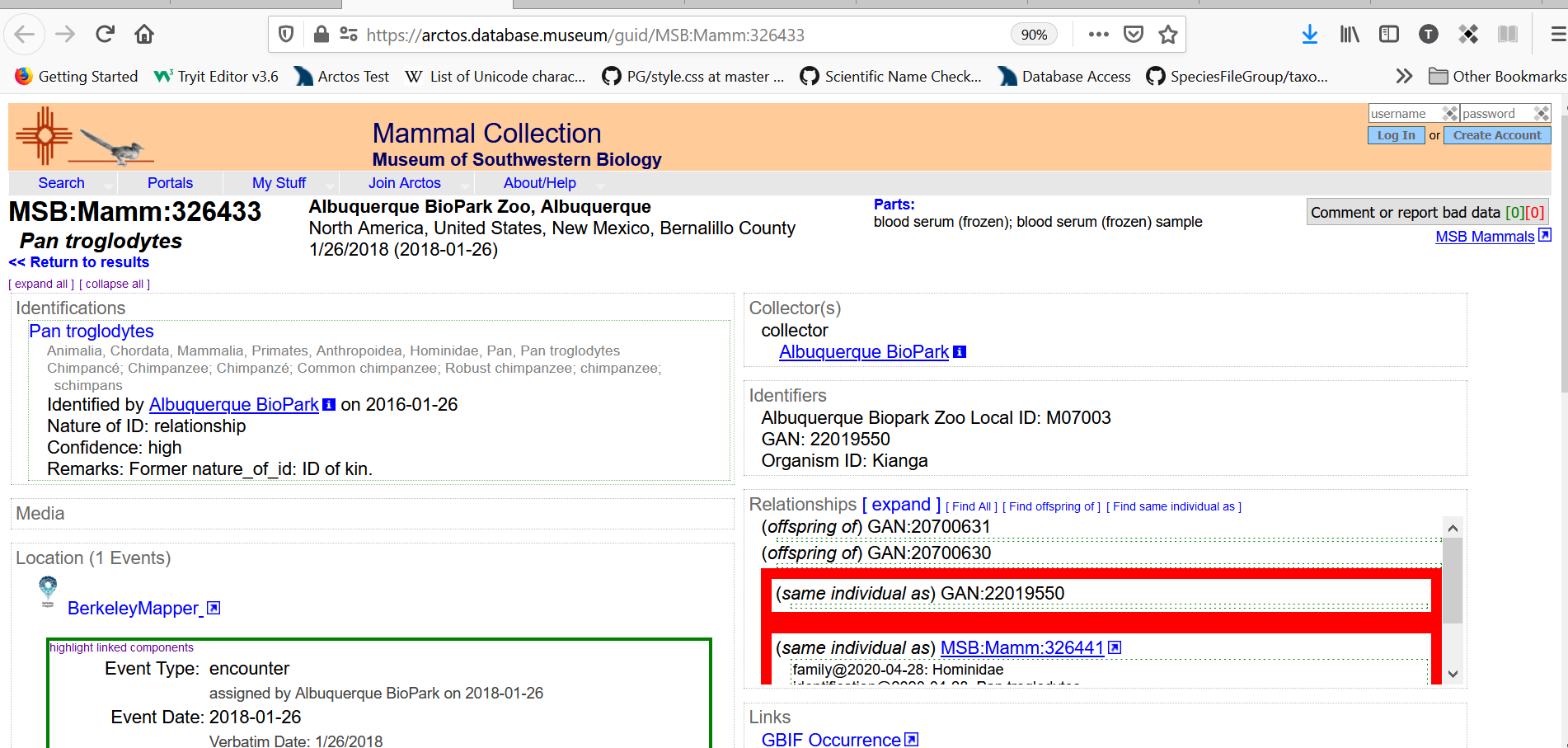
How do I get here?
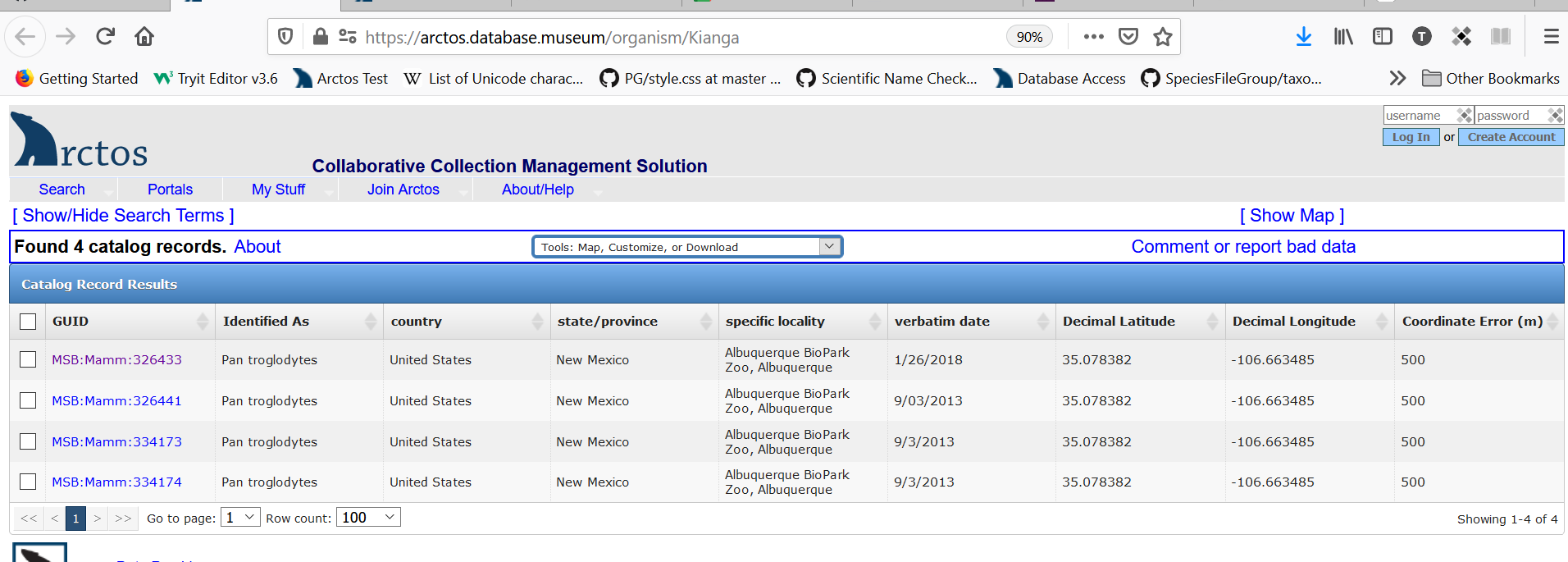
and WHERE do I record basic information about Kianga so that I might tell that Kianga apart from any other Kianga?
I think, if we are all on the same page, through adding a dwcOrganism field
that is actually a link to the search? That would be for "any ID we have
that could allow to find all these records, ideally a uuid if one exists",
and the search itself it a url = uuid. In this case, ideally we'd designate
the GAN number as the search term, although we could use "Kianga" as well,
or perhaps both?
Note that at MSB we have loaded over 1000 records in this project in the
last couple of days, and we had to load the "same individual as"
relationship to the GAN number, which is not a live link. We need then to
turn the GAN number relationship to a relationship to the search for the
GAN number, rather than the number itself. We could create those "organism"
searches in advance, or after bulkload. ?
On Wed, May 5, 2021 at 9:54 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
When I am here
[image: image]
https://user-images.githubusercontent.com/5725767/117170888-a24bd100-ad87-11eb-89de-fb39c77aa990.pngHow do I get here?
[image: image]
https://user-images.githubusercontent.com/5725767/117170958-b1328380-ad87-11eb-9af6-ad6379341d62.pngand WHERE do I record basic information about Kianga so that I might tell
that Kianga apart from any other Kianga?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-832807777,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBFU5HEYVDJACICZT4DTMFS3LANCNFSM4G5YNGMA
.
How do I get here?
Add a base_url of https://arctos.database.museum/organism/ to the identifier to get a clicky link.
WHERE do I record basic information
In the "Occurrences." I'm not arguing that we don't need a maybe-agent-like Occurrence node, just saying it's not very functional - we have the tools to address the core problem without major development, even if some of the icing is elusive for the time being. (Or maybe the alleged icing is evil - see above.)
tell that Kianga apart from any other Kianga?
How do you tell that NK1 apart from the other 27 printer-jam-caused NK1s? I think the choice here is between minting some known-unique identifier in some way (grab a UUID, grab an ARK, let OrganismBase give you something, WHATEVER) and then trying to find all of THAT NK1 and no other NK1s and giving them that identifier, or using something "natural" and knowing that from time to time it's going to involve some disambiguation. I don't think that's a technical problem - technology can certainly help with various aspects, but it all comes back to assigning identifiers - and it's certainly nothing unique to this particular usage of identifiers. However you approach it, it's ultimately going to require digging around in the Occurrences and probably trying to guess if the data or identifiers or something else has been mangled in some way.
ideally
See above, social problem, probably one heavily reliant upon your local procedures.
Note that the "agent-like Organism node" would remove the choice - you'd need to use identifiers issued by it. (Perhaps there could be some magic to find/create those, but that's back to 'significant development' mode.)
both
Arctos doesn't care, you have have any number of identifiers of any type. There is no mechanism to pass multiples to GBIF, though.
We need then to
turn the GAN number relationship to a relationship to the search for the
GAN number, rather than the number itself.
I really need an example, I don't know what this means.
We could create those "organism"
searches in advance, or after bulkload. ?
If you mean https://arctos.database.museum/organism/Kianga, they create themselves. If you mean something else, I need clarification.
WHERE do I record basic information about Kianga
Alternately: catalog the Organism, use the GUID produced by that as OrganismID in the Occurrence catalog records. As above that probably requires some minor change to indicate that the record is an Organism (maybe something in https://arctos.database.museum/info/ctDocumentation.cfm?table=ctcataloged_item_type), but I think that's minor details. That approach provides access to much more than an agent-like object could. (And some of the synthesis could be automated, with development.)
I am sticking with an agent-like table as a go to. All of the above looks like a LOT of extra work for little return. I may be dense and eventually I'll get there, but I just don't understand the opposition to Kianga being a THING with metadata.
No opposition from me - I think the agent concept would be excellent. But
in the shorter term, without that, all I can see doing is somehow hanging a
search url onto an identifier that we call OrganismID.
On Wed, May 5, 2021 at 11:03 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
I am sticking with an agent-like table as a go to. All of the above looks
like a LOT of extra work for little return. I may be dense and eventually
I'll get there, but I just don't understand the opposition to Kianga being
a THING with metadata.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-832858450,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBBA5OXG6WWJCBIWKIDTMF26VANCNFSM4G5YNGMA
.
extra work
If I have any "opposition" to the proposed new node, it comes back to that - but exactly opposite of what I think you're implying. They're both shared identifiers, the difference is in how the identifier is acquired.
To use "Kianga" as an identifier, you type it in under the correct IDtype. That's it.
To use whatever the agent-like thingee would do, you have to somehow negotiation with it to be issued an identifier, then you have to add that to all of the records (or somehow develop a way to abstract existing IDs to "the real organism ID", or SOMETHING.) Whatever the details turn out to be, it will almost certainly turn out to be MORE work - perhaps significantly more - than typing "Kianga." I'm not sure that would be used.
somehow hanging a
search url onto an identifier that we call OrganismID.
I don't think we're on the same page yet. All you need to do it this:

If that's not what you're talking about or doesn't clarify anything, maybe we should set up a zoom?
hanging a search url onto an identifier that we call OrganismID.
I feel like I have been chastised when I suggested/tried it https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472642753
Plus it doesn't fix https://github.com/ArctosDB/arctos/issues/1966#issuecomment-472565489
I don't understand the "agent-like" table opposition. Why create a whole new collection for "organisms" when we have a working model for that in Agents?
base URL
Why are you always about 10 seconds ahead of me?! Yea agreed, that's going to be hard on external things, there's now a handler for Organism ID.
Agents
????????????
Are you saying that "Mexican Wolf Studbook number 1216" should just be an agent? Because that would work.
No, I don't think that's an Agent - I think it's a different kind of data object. (I thought you were saying that, then I thought you weren't, now I just don't know what you're saying.)
Assuming that is intended to represent an individual (or something similar), it is a THING. That THING could be
- just the composite of the other things that wear the identifier, which is what https://arctos.database.museum/organism/Kianga is
- a catalog record, which provides an identifier and has the capability of carrying extensive data (including more identifiers) of its own
- something entirely new, which might look a bit like Agents (or not, whatever, functional requirements would shape it, without those we're just tossing stuff at the wall hoping something sticks)
This is what Teresa proposed, and I support, as the best idea - but then I
thought that got shot down, so was trying other suggestions:
- something entirely new, which might look a bit like Agents (or not,
whatever, functional requirements would shape it, without those we're just
tossing stuff at the wall hoping something sticks)
On Wed, May 5, 2021 at 11:56 AM dustymc @.*> wrote:
- [EXTERNAL]*
No, I don't think that's an Agent - I think it's a different kind of data
object. (I thought you were saying that, then I thought you weren't, now I
just don't know what you're saying.)Assuming that is intended to represent an individual (or something
similar), it is a THING. That THING could be
- just the composite of the other things that wear the identifier,
which is what https://arctos.database.museum/organism/Kianga is- a catalog record, which provides an identifier and has the
capability of carrying extensive data (including more identifiers) of its
own- something entirely new, which might look a bit like Agents (or not,
whatever, functional requirements would shape it, without those we're just
tossing stuff at the wall hoping something sticks)—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-832891943,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBA6NGVOEMDNSUZB363TMGBFPANCNFSM4G5YNGMA
.
I don't think anything is currently shot down. There are things we can do without development, and things that would require some sort of development. (And some - maybe all - of the things that we can do now could be migrated to some future tool.)
Nothing yet looks "best" to me - I think everything has some sort of cost and some sort of benefit. Functional requirements should change that view.
Let's discuss at tomorrow's meeting? Or set up a separate call?
On Wed, May 5, 2021 at 2:23 PM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
Kianga - https://www.wikidata.org/wiki/Q106716237
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-832982254,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBBUF7HMQSWH4JIM3TDTMGSM5ANCNFSM4G5YNGMA
.
There is also a giraffe named Kianga - https://www.courier-journal.com/story/news/2018/04/04/one-year-old-giraffe-kianga-joins-louisville-zoo-sunshine/485859002/
a rhino - https://he-il.facebook.com/sdzsafaripark/videos/baby-kianga-rocking-out/1322623687754304/
maybe another rhino - https://www.facebook.com/racinezoo/videos/meet-kianga-and-timu-our-gentle-giants/266705597661072/
That's why we need to use the GAN number which is the formal identifier
used by the multi-zoo ZIMS database.
On Wed, May 5, 2021 at 3:09 PM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
There is also a giraffe named Kianga -
https://www.courier-journal.com/story/news/2018/04/04/one-year-old-giraffe-kianga-joins-louisville-zoo-sunshine/485859002/a rhino -
https://he-il.facebook.com/sdzsafaripark/videos/baby-kianga-rocking-out/1322623687754304/maybe another rhino -
https://www.facebook.com/racinezoo/videos/meet-kianga-and-timu-our-gentle-giants/266705597661072/—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-833008515,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBDX5ZSTTTGZJMHCLFTTMGXXHANCNFSM4G5YNGMA
.
The great thing about Wikidata is we can make it whatever we want. All we need is an other ID of "Wikidata ID" - The Q identifier in Wikidata associated with this cataloged item. base url = https://www.wikidata.org/wiki/
How about that Bus? https://www.wikidata.org/wiki/Q97275473
Anyone can contribute to the knowledge about the things. I can add as much info as I want or can find.
If those GAN numbers were available somewhere, they could be added to wikidata.
Other advantages:
Provides a GUID for whatever organism you want
The whole world can understand it and see what you mean (it is bigger than Arctos)
No need to pay for storage, DOI, or whatever
The PROBLEM is that Wikidata probably won't let me create and keep the item "mouse 12345"
They are in the ZIMS database, currently a non-public database used by
multiple AZA zoos. There is likely content in there that should not be
public, but we are adding the GAN numbers to Arctos. We need to include zoo
folks in the discussion prior to going through Wikidata.
On Wed, May 5, 2021 at 3:14 PM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
If those GAN numbers were available somewhere, they could be added to
wikidata.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-833011582,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBDVSZSJD4RVR4TLXHTTMGYLPANCNFSM4G5YNGMA
.
Everything in Wikidata I found either in the news article or Arctos. It is already published somewhere. If it is already available on line, there isn't a lot to be done.
Fine, whatever, that's essentially the "build something around the ID itself" model, but it's already built. I don't and can't care what the value of the ID is!
Great, strings are strings, go for it.
(But do you really want to make a facebook page for each of the bajillion mark-recapture Peros? Because that's what using some sort of "issued" ID requires, although perhaps much of it could be done via webservices.)
use the GAN number
Awesome - use it!
GAN numbers...added to wikidata.
Sure, whatever, maybe there's some procedural reason to get all of the identifiers from the same place, but it doesn't matter from here.
All we need is an other ID of "Wikidata ID"
If you want Organism functionality (eg if we're handing this off to GBIF) then you need to use Organism ID (or whatever we decide on). The TYPE of identifier is important.
I added another OrgID (which will break GBIF if we get there, see above): https://arctos.database.museum/organism/https://www.wikidata.org/wiki/Q106716237
discuss
Sure, I'm up for either or both.
Just this week there has been a meeting to discuss zoo and museum
collaborations, so we need to get with that community to move forward with
data sharing. We can use the MSB /ABQ Biopark project as a prototype of
what can be done -but best if we start with integration between Arctos and
ZIMS.
On Wed, May 5, 2021 at 3:37 PM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
Other advantages:
Provides a GUID for whatever organism you want
The whole world can understand it and see what you mean (it is bigger than
Arctos)No need to pay for storage, DOI, or whatever
The PROBLEM is that Wikidata probably won't let me create and keep the
item "mouse 12345"—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-833023452,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBC46VUBGLBGDTO3D5DTMG3AZANCNFSM4G5YNGMA
.
We need this to work also for many thousands of AMNH and MSB specimens that
are "same individual as" - and I don't see using Wikidata for that. Let's
back up and look at other options.
On Wed, May 5, 2021 at 3:41 PM dustymc @.*> wrote:
- [EXTERNAL]*
https://www.wikidata.org/wiki/Q106716237
Fine, whatever, that's essentially the "build something around the ID
itself" model, but it's already built. I don't and can't care what the
value of the ID is!Great, strings are strings, go for it.
(But do you really want to make a facebook page for each of the bajillion
mark-recapture Peros? Because that's what using some sort of "issued" ID
requires, although perhaps much of it could be done via webservices.)use the GAN number
Awesome - use it!
GAN numbers...added to wikidata.
Sure, whatever, maybe there's some procedural reason to get all of the
identifiers from the same place, but it doesn't matter from here.All we need is an other ID of "Wikidata ID"
If you want Organism functionality (eg if we're handing this off to GBIF)
then you need to use Organism ID (or whatever we decide on). The TYPE of
identifier is important.I added another OrgID (which will break GBIF if we get there, see above):
https://arctos.database.museum/organism/https://www.wikidata.org/wiki/Q106716237discuss
Sure, I'm up for either or both.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-833027791,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBBJXJJ62WELL5DOT4LTMG3RHANCNFSM4G5YNGMA
.
And I added my solution to this set with subject = Kianga
I'd like everyone to imagine that instead of "subject" this could be "organism".
The Kianga "organism" page would need some tweaking. Instead of aka those would be identifiers (from a code table) with relationships. Any catalog records associated with the Kianga organism would automagically get these identifiers as well. There would need to be some additional fields that would also propagate to any catalog records. These would be attributes, some of which already exist in agents like date of birth and date of death, but we would need others - like sex. The "associate of" relationships could stay the same or maybe would be more appropriate as "housed at". And note that Kianga already has her wikidata ID associated.
And I say that rather than relying on "preferred name" these organism profiles should rely on some identifier. Only some organisms will have names, the majority could just use "12345" or the like.
Add new "node" for Arctos users to create entities.
Entity table that works like agents but with different attributes.
AWG discussed.
Priority: just do it, now, continue to use Organism ID
- works like any other identifier, suitable for any situation, requires ~no extra work for "normal" data entry (eg banded bird), allows any amount of data for URLs that carry it
Development
Create "entity node" which will...
Immediately
- issue IDs suitable for use as Organism ID
- provide a place for human-supplied metadata
Eventually
- gather data (including identifiers) from bits-n-pieces (=build a dynamic picture of the Entity)
- provide data to display on bits-n-pieces (=dynamically enhance Occurrences)
- auto-create (suggest? not sure, need some real data to play with) relationships based on Organism ID.
- find and suggest Organism ID (from identifiers, collector+location, etc.)
- Other Cool Stuff, maybe
Entity Model
Table Entity
- Entity_ID varchar primary key
- Entity_Type varchar not null IN (organism, other stuff later)
- created_by fkey(agent.agent_id)
- created_date timestamp
Table Entity_Assertion
- Entity_Assertion_ID pkey
- Entity_ID FKEY(entity.Entity_ID)
- assertion_type varchar not null IN (preferred entity ID, alternate ID, organism identification, birth date, etc., etc., etc.)
- assertion_value varchar not null
- assertion_url varchar
- assertion_remark varchar
- asserted_by fkey(agent.agent_id)
- asserted_date timestamp
Entity_ID is eg https://arctos.database.museum/entity/Kianga
Entity_Type allows using this to stitch together arbitrary - well, anything, I think - certainly anything represented by catalog records, like dinnerware sets, botanical duplicates, wolfpacks, WHATEVER
Assertions should accommodate about anything people want to assert, plus (eventually) things pulled from eg GBIF (potentially using an ID asserted by a person).
Assertion type preferred entity ID could serve as a redirect when someone inevitably creates two entity records for the same entity, including if some external source becomes available/preferred.
alternate ID (or maybe component ID IDK it's just data, we can make these up as we go) with values like https://arctos.database.museum/guid/MSB:Mamm:302171 could serve as belt-and-suspenders, and would provide some redundancy in case someone deletes the Entity ID from a catalog record.
Example
from https://arctos.database.museum/agents.cfm?agent_id=21332262
Entity_ID: https://arctos.database.museum/entity/Kianga
Entity_Type: organism
|Assertion_Type|Assertion_Value|Assertion_URL|
|----|----|---|
|organism identification|Pan troglodytes|https://arctos.database.museum/name/Pan%20troglodytes|
|remark|chimpanzee in the Chimpanzee Species Survival Program |
|alternate identifier|Albuquerque Biopark Zoo Local ID: M07003 |
|alternate identifier|GAN: 22019550 |
|birth date|2007-02-04 |
|alive on date|2020-03-03 |https://www.arkansasonline.com/news/2020/apr/02/lr-zoo-welcomes-13-year-old-chimpanzee/|
|associate of|Rio Grande Zoo |https://arctos.database.museum/agent/21318999|
|associate of|Little Rock Zoo |https://arctos.database.museum/agent/21332263|
|more information|wikidata |https://www.wikidata.org/wiki/Q106716237|
Feedback on the model is greatly appreciated - what do you want to do that this can't, what could be better, have I somehow entirely missed the point, ?????
Feedback on the terms (not including table and column names) is not useful at this time; there will be code tables, ya'll can work that out while I'm building, after the model is solidified.
Documentation
"Good" IDs (eg urls) are preferred. "Not so great" IDs (bird band numbers, eartag numbers, whatever) are acceptable but should be expected to occasionally contane tipoez or be duplicates.
What to catalog guidance
Workflow:
- Locate or create an Entity ID, possibilities include but are not limited to
- USGS BBL number
- wikidata URLs
- "Entity" URLs (of existing entities or things you've just created)
- GAN numbers
- Mexican Wolf Studbook numbers
- Use that ID as Organism ID (or whatever we call it) with the involved catalog records
TODO
- allow no more than one Organism ID for a catalog record
- allow no more than one 'preferred id' (need firm name first) per entity
- HTML-wrap specimendetail when necessary
- For DWC export: IF has Organism ID THEN use it ELSE do whatever we're doing now
- consider (somehow) forcing this into downloads; it's a powerful way to detect relationships (eg bits in two collections) without having to include relationships or unwind complicated data
- consider (somehow, again) making Organism ID more prominent on catalog records
@campmlc @Jegelewicz does this work, need more discussion, ????
I'd like to try it with some examples. I just marked to load over 3000
Elephant records that are multiple repeat samples from the same
individuals. In terms of workflow to implement OrganismID, I assume I would
need to load an OrganismID use batch tools once these are loaded?
On Mon, May 10, 2021 at 8:51 AM dustymc @.*> wrote:
- [EXTERNAL]*
@campmlc https://github.com/campmlc @Jegelewicz
https://github.com/Jegelewicz does this work, need more discussion, ????—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-836795400,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBFOLPIUYJR4FBTIIV3TM7XHBANCNFSM4G5YNGMA
.
There's really nothing to try. You can load WHATEVER to Organism ID now, it'll work like any other identifier. If you use a URL for the identifier, it will (eventually, at least) be clickable. There's little to nothing new being proposed on that front.
Above is a proposal of one way to get a URL-identifier. The page that identifier opens can look like about anything, I need to know if the structure I've proposed is suitable before it gets built.
If "elephant number 7" is what you have and want, you can just load that as Organism ID using any of the existing tools. Do note that we'll need a 'zero or one' rule on Organism ID; if you eventually want something more than "elephant number 7" you should enter that as something other than Organism ID.
If you want an "entity" (and what I've laid out above is approved and gets built), you'd need to somehow "translate" 'elephant number 7' into https..../entity/{something}. You could create that "something" before you load and use it in the initial load, or create and bulk-add it afterward, or whatever. (I'd probably add them both, no matter when or how you create the entity, on the assumption that 'elephant number 7' might mean something to someone, but that's 100% curatorial call.)
OK, in this case I could use the ZIMS GAN number, which is the source of
the data we are using to catalog. I've already entered this number as
"self" and as "same individual as" when there is more than one event.
However, the "same individual as" will be a dead link - it just lets me
know that that record needs an OrganismID and eventual relationships to
cataloged items.
So if I eventually want to use the modified agent = entity model, I
shouldn't use the GAN number as "OrganismID", but instead create an agent
url and use that?
On Mon, May 10, 2021 at 10:16 AM dustymc @.*> wrote:
- [EXTERNAL]*
There's really nothing to try. You can load WHATEVER to Organism ID now,
it'll work like any other identifier. If you use a URL for the identifier,
it will (eventually, at least) be clickable. There's little to nothing new
being proposed on that front.Above is a proposal of one way to get a URL-identifier. The page that
identifier opens can look like about anything, I need to know if the
structure I've proposed is suitable before it gets built.If "elephant number 7" is what you have and want, you can just load that
as Organism ID using any of the existing tools. Do note that we'll need a
'zero or one' rule on Organism ID; if you eventually want something more
than "elephant number 7" you should enter that as something other than
Organism ID.If you want an "entity" (and what I've laid out above is approved and gets
built), you'd need to somehow "translate" 'elephant number 7' into
https..../entity/{something}. You could create that "something" before you
load and use it in the initial load, or create and bulk-add it afterward,
or whatever. (I'd probably add them both, no matter when or how you create
the entity, on the assumption that 'elephant number 7' might mean something
to someone, but that's 100% curatorial call.)—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-836907174,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBDUSI4C4O6DYXEABNLTNABHNANCNFSM4G5YNGMA
.
ZIMS GAN number,
You can
- Just use that, or
- Create an Entity, to which you could add some more information about that identifier
needs an OrganismID and eventual relationships
See above - I can almost assuredly suggest/create relationships (using resolvable identifiers) from OrganismID.
create an agent url
As far as I know that's completely off the table; that was a demonstration of the kinds of information that the "entity" would carry, and my understanding was that we'd all agreed to limit Agents to humans.
If you want something more than a string (and GAN doesn't provide one - you can just use their URL if they do), I'd recommend for now:
- Carefully review the proposal above so we can get started building something
- Add the GAN number as whatever identifier type is most appropriate
Then when/if Entities are built,
- Use the GAN number to create Entities
- Bulkload those IDs to the records
After Entities are built, you could do the same thing, or you could first Use the GAN number to create Entities and just include those in your initial catalog record bulkloader.
Immediate prioritization was approved by the AWG, but I don't want to build something that you're not going to use. If https://github.com/ArctosDB/arctos/issues/1966#issuecomment-834419533 works, I can build it. If it doesn't, we should find something that does and I can build that.
I guess I just need clarification as to how to proceed. Currently, the only ID I have to use in this case to identify individuals is the GAN number. There is no currently available url. I would prefer to use the entity model under development. Given that, how do I proceed?
1) Do I load the GAN now as an OrganismID identifier?
2) Or do I wait until an entity url is available?
prefer to use the entity model
Then I need confirmation that what I've proposed is what you want (or a better idea if it's not).
Do I load the GAN now as an OrganismID identifier?
I would NOT recommend that; my intention is to allow a maximum of one OrganismID (so we can feed it to GBIF and etc.), if you load this you'd have to unload it (there's a bulkunloader, but no reason to add work) before switching to something better.
I WOULD recommend loading that as something you can find later - GAN (if that's a type) or "original identifier" or WHATEVER, as long as you can use it to find these. (I can help with SQL or whatever, of course.)
Or do I wait until an entity url is available?
You could do that as well. I think the most basic version of what I've proposed would be pretty quick to build, but maybe that's not the final form or we'll run into some unanticipated complication or .....
In either case, if my proposal is approved I would like to use a copy of your data for development if you don't mind passing that along.
What you've proposed looks good to me, based on our conversations. I'd like
to see how it actually works, but I think the core components are there, eg
similar to agents.
On Mon, May 10, 2021, 10:59 AM dustymc @.*> wrote:
- [EXTERNAL]*
prefer to use the entity model
Then I need confirmation that what I've proposed is what you want (or a
better idea if it's not).Do I load the GAN now as an OrganismID identifier?
I would NOT recommend that; my intention is to allow a maximum of one
OrganismID (so we can feed it to GBIF and etc.), if you load this you'd
have to unload it (there's a bulkunloader, but no reason to add work)
before switching to something better.I WOULD recommend loading that as something you can find later - GAN (if
that's a type) or "original identifier" or WHATEVER, as long as you can use
it to find these. (I can help with SQL or whatever, of course.)Or do I wait until an entity url is available?
You could do that as well. I think the most basic version of what I've
proposed would be pretty quick to build, but maybe that's not the final
form or we'll run into some unanticipated complication or .....In either case, if my proposal is approved I would like to use a copy of
your data for development if you don't mind passing that along.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-836967228,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBAM2A4LPHYJBBJQPJTTNAGFJANCNFSM4G5YNGMA
.
see how it actually works,
http://test.arctos.database.museum/entity/boogity
Still lots of wrinkles to iron out, but probably to the point where some feedback could be useful.
Question on new entity editing page:
The "alternate entity name" -> this is a second website reference for the same animal, that was created as a mistake, but since it is linked to this entity page will still lead back to the correct entity page/reference?
Yes, that idea seems within scope.
Any assertion leads to whatever's in the URL, which can be nothing. You could also use this mechanism to "migrate" from some nonresolvable ID (eg bird band number) into this system.
preferred entity ID works the same way in the opposite direction - you can move to something outside of Arctos, or back to text identifiers, and leave a trail. http://test.arctos.database.museum/entity/baggity is a lateral move to another Arctos Entity, but that URL could be NULL (eg everything we know is in the value) or https://organismbank.org/.... or anything else.
You can also just ignore all of that - the tools to build a robust history are built in, but most of the time for most records I doubt they'll be necessary. I wouldn't mind being wrong about that....
My definitions (which probably need work) are in the usual place, http://test.arctos.database.museum/info/ctDocumentation.cfm?table=ctentity_assertion_type (and the bottom of the edit form, at least for now).
Can we change some of the controlled vocab, e.g. replace "component" with
"related catalog item" or "associated guid"? Related url? Something
human-recognizable?
Also, can we change the display order to be:
component_1 = related catalog item_1 = MSB:Mamm:326433 link
http://test.arctos.database.museum/guid/MSB:Mamm:326433
component_ID_1= related_catalog_item__ID_1 =Pan troglodytes
component_identifier_1 = related_catalog_item_identifier_1= Kianga
component_2=related_catalog_item_2 = Some mouse or something link
http://test.arctos.database.museum/guid/MSB:Mamm:999
component_ID_2 = related_url_ID_2 = Peromyscus leucopus
component_identifier_2= related_identifier_2= M07003
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Albuquerque%20Biopark%20Zoo%20Local%20ID&oidnum=M07003
component_3= organism_ID_3=
http://test.arctos.database.museum/entity/boogity
On Wed, May 12, 2021 at 7:22 PM dustymc @.*> wrote:
- [EXTERNAL]*
Yes, that idea seems within scope.
Any assertion leads to whatever's in the URL, which can be nothing. You
could also use this mechanism to "migrate" from some nonresolvable ID (eg
bird band number) into this system.preferred entity ID works the same way in the opposite direction - you
can move to something outside of Arctos, or back to text identifiers, and
leave a trail. http://test.arctos.database.museum/entity/baggity is a
lateral move to another Arctos Entity, but that URL could be NULL (eg
everything we know is in the value) or https://organismbank.org/.... or
anything else.You can also just ignore all of that - the tools to build a robust history
are built in, but most of the time for most records I doubt they'll be
necessary. I wouldn't mind being wrong about that....My definitions (which probably need work) are in the usual place,
http://test.arctos.database.museum/info/ctDocumentation.cfm?table=ctentity_assertion_type
(and the bottom of the edit form, at least for now).—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-840215764,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBBAEOBBB4UFLXTPQCLTNMSWLANCNFSM4G5YNGMA
.
"related catalog item" or "associated guid"? Related url?
I'm not sure what we should call this thing, but it's more _intimate_ than any of that - the "components" are parts of some sort of whole, not just "related" in some way.
organism_ID_3
- That sets us up to need organism_ID_3987, which isn't sustainable or usable.
- This isn't intended to be any sort of replacement for catalog records - "M07003" might be somehow handy for finding this thing (my thoughts are on avoiding duplicates), but that identifier belongs to the catalog record, at least more than it does to the Entity. (If it does belong to the entity then it's an
alternate entity namerather than acomponent identifier- I don't have any way to recognize those, they'd need to be added by humans.) - We may find some other use case at some point, but for now Entities are aimed at homogenous things - an outlying ID is cause to go checking data, not something that needs to be associated with the Entity in some more-defined way.
Shall we set up another call? There's something to look at now, would be good to be sure we're on the same page before I write much more code.
Yes, happy to do a call. Today before or after AWG?
On Thu, May 13, 2021 at 1:20 AM dustymc @.*> wrote:
- [EXTERNAL]*
"related catalog item" or "associated guid"? Related url?
I'm not sure what we should call this thing, but it's more intimate
than any of that - the "components" are parts of some sort of whole, not
just "related" in some way.organism_ID_3
- That sets us up to need organism_ID_3987, which isn't sustainable
or usable.- This isn't intended to be any sort of replacement for catalog
records - "M07003" might be somehow handy for finding this thing (my
thoughts are on avoiding duplicates), but that identifier belongs to the
catalog record, at least more than it does to the Entity. (If it does
belong to the entity then it's an alternate entity name rather than a component
identifier - I don't have any way to recognize those, they'd need to
be added by humans.)- We may find some other use case at some point, but for now Entities
are aimed at homogenous things - an outlying ID is cause to go checking
data, not something that needs to be associated with the Entity in some
more-defined way.Shall we set up another call? There's something to look at now, would be
good to be sure we're on the same page before I write much more code.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-840372304,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBD7HQLJHRIJVHUNKYLTNN4TJANCNFSM4G5YNGMA
.
Sure, either should work for me, say when.
For the terminology, it should cover things that are not Occurrence-like (which was my first thought) - think tea sets and whatever the rock-people might bring (chunks of a core maybe).
For scale, this should work to stitch together a lifetime of GPS tag pings (maybe for all the wolves of a pack).
For scope, it should accept "UAM, maybe the one in Madrid, we don't know, and not sure which department, number 1234, which probably isn't the primary identifier" (in the hope that someone will figure it out by association and add better IDs) and stuff from Arctos/GBIF/etc., which might have extensive data (from which we can just pull in what we want).
I don't much care HOW we do that, but I'd very much prefer to not do something that precludes those kinds of usage.
I can do 1:30-2pm MDT - gives us a little break after AWG? Or I can log in
early at 11am MDT. Or both.
On Thu, May 13, 2021 at 8:58 AM dustymc @.*> wrote:
- [EXTERNAL]*
Sure, either should work for me, say when.
For the terminology, it should cover things that are not Occurrence-like
(which was my first thought) - think tea sets and whatever the rock-people
might bring (chunks of a core maybe).For scale, this should work to stitch together a lifetime of GPS tag pings
(maybe for all the wolves of a pack).For scope, it should accept "UAM, maybe the one in Madrid, we don't know,
and not sure which department, number 1234, which probably isn't the
primary identifier" (in the hope that someone will figure it out by
association and add better IDs) and stuff from Arctos/GBIF/etc., which
might have extensive data (from which we can just pull in what we want).I don't much care HOW we do that, but I'd very much prefer to not do
something that precludes those kinds of usage.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-840618548,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBE2ZEIO6OEDXJJ6NKDTNPSIRANCNFSM4G5YNGMA
.
Mabe both, just in case?
sure, sounds good!
On Thu, May 13, 2021 at 9:14 AM dustymc @.*> wrote:
- [EXTERNAL]*
Mabe both, just in case?
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-840629479,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBCBBNMXLC5VSV7PIRDTNPUDNANCNFSM4G5YNGMA
.
Store what we've got, use it for error detection and search. (Clarify somewhere.)
For display, live pull data associated with components.
Component terminology: todo: fake it in the UI, keep 'component' in the model.
Start of documentation: https://handbook.arctosdb.org/documentation/entity.html
This is in production - will add live pull to the viewer when there's some use.
I can't wait to have a little time to mess with this!
Yes, wahoo! I have data in Arctos to immediately apply this to. Everything
in this project with a GAN number as the organism ID. Can you magic it?
https://arctos.database.museum/project/10002948
On Tue, May 18, 2021 at 9:57 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
I can't wait to have a little time to mess with this!
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-843292017,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBF2TOM3QG5CLTGGCK3TOKE7FANCNFSM4G5YNGMA
.
Can you magic it?
Probably, but I'd rather do that after there's some "normal" usage to make sure nothing's too broken/wrong/whatever.
I wouldn't mind magicking things like http://test.arctos.database.museum/entity/NK217004 as well, if everybody's down with that.
Sure, linking NKs is a very good test case. I imagine we will find all
sorts of interesting things. . . .
On Tue, May 18, 2021 at 10:13 AM dustymc @.*> wrote:
- [EXTERNAL]*
Can you magic it?
Probably, but I'd rather do that after there's some "normal" usage to make
sure nothing's too broken/wrong/whatever.I wouldn't mind magicking things like
http://test.arctos.database.museum/entity/NK217004 as well, if
everybody's down with that.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-843313754,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBB26DWXLIANJ3VKVNTTOKGYZANCNFSM4G5YNGMA
.
Comments on new Entity Node.
- Let's put "Entities" in the "Manage Data" drop down menu after "Agents" and not hide the new entity creation/edit in the search function.
- How about we display text instead of the link on the catalog record page? This looks nuts
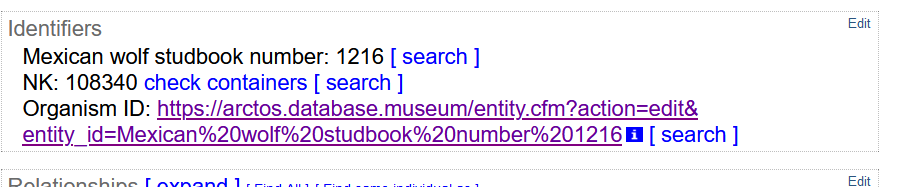
The text could be standard so that anything starting with "http" instead displays "Webpage" (or whatever - I'm not thinking too creatively right now) Also, as you can see from my example, we risk putting the wrong url in if you are creating the entity, then trying to copy pasta into the org id. - Add sex (use the ctsex_cde table)
- Add death date (same format as birth date)
- I think we need to allow for the use of other Agents in the "asserted by". For instance, in the case of the Mexican Wolves, I did not assert their sex, U. S. Fish and Wildlife Service Mexican Wolf Recovery Program did.
- "It is not possible to modify an Entity after creation, but a preferred alternative can be asserted." - Can we use the same wording we use with Agents for simplicity "bad duplicate of"? Also, entities CAN be updated, just not the entity name. I am on the fence about the assigned name becoming the GUID. Wouldn't it be better to use an index for entities and have the number be the GUID? This is how Agents work and as we have already seen - there is more than one Kianga.
- The magic Pull doesn't seem to work? Because it is looking in test? I was using this url https://arctos.database.museum/guid/MSB:Mamm:233261
- The link to the preferred Organism ID doesn't do anything?
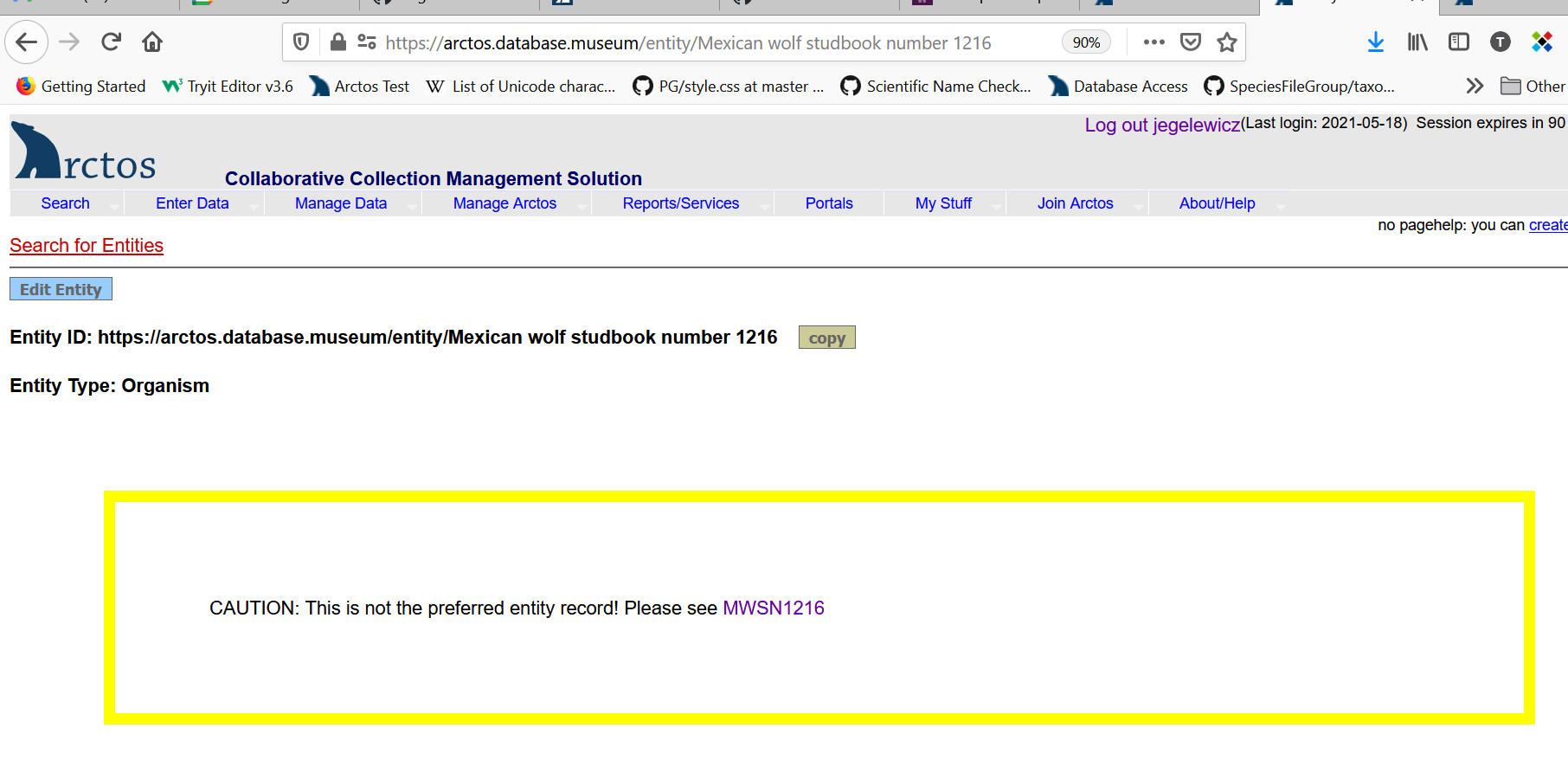
when I click MWSN1216, nothing happens - I clearly do not understand the "component" idea - I think I need a tutorial.
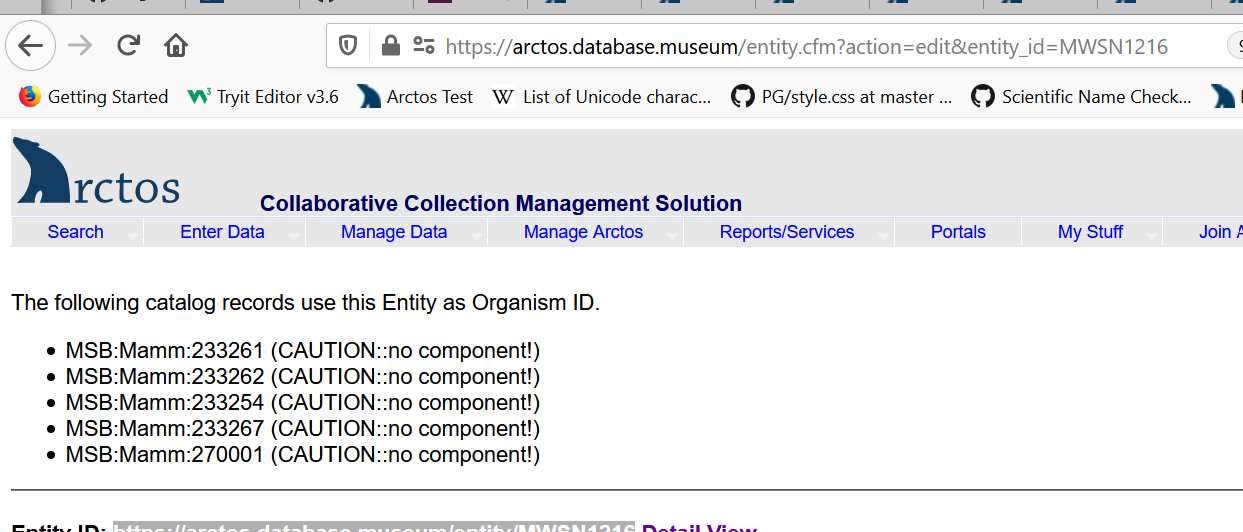
My adventure with Mexican Wolf Studbook Number 1216 was revealing. When I added the Organism ID to all of the "same individual as" from https://arctos.database.museum/guid/MSB:Mamm:233261 by clicking the linked records, I ended up with 6 records associated with this wolf. But when I clicked on "find all same individual as" from the same record, I get 5 (the record I was searching from was excluded) which seems like a bug.
In addition to the above, it seems like being able to add other identifiers to the entity (Mexican Wolf Studbook Number) would be useful. It would be great to add Other ID type = Mexican Wolf studbook number, Other ID value = 1216 to the entity MWSN1216 and have it propagate to any catalog record that has Organism ID = MWSN1216.
This looks nuts
You're using the wrong thing; along with looking nuts, that'll only work for "us." Use the copy button to get the ID

display text instead of the link
My major epiphany in this process is that we can't - OrganismID can be "3", or "http://someRandomWebsite.org/whatever" or something that Arctos issues, and the only way to deal with that is to treat them all the same (as strings, some of which DO STUFF in browsers, some of which might even be useful).
Add {stuff}
Whatever, it's just data, there's a CT process (which could also be used to get rid of the stuff I've added, if necessary). BUT: the stuff I added I added for discoverability, I'll pull "local" (Arctos, maybe GBIF, IDK) records into the view at some point, if you just want to SEE that then perhaps it doesn't need explicitly added.
not hide the new entity creation/edit in the search function.
It's there on purpose; creating 17 entities for one critter and then merging them is possible, but it's a lot of work and a mess. Searching those things I've pulled in first is a tiny bit of work that can avoid that mess. I think the step will prove very worthwhile, but whatever - there are ways around that path, maybe duplicates won't happen, IDK, seems worth trying out at least but still whatever....
I did not assert their sex, U. S. Fish and Wildlife Service Mexican Wolf Recovery Program did.
If we're pulling it, that can be in the display. If we're asserting it for the entity, then its not being asserted by USFWS (who was probably dealing with something closer to an Occurrence - if this is a fish that assertion could be less static). I'm not necessarily saying one or the other is some form of "correct" and we should do that, just that there are two very different kinds of data objects involved.
"bad duplicate of"
No terribly strong feelings on that, but it's not QUITE the same - maybe I just don't want to use your assertion for some reason, mine's not "bad" it's just not preferred by you.
entities CAN be updated, just not the entity name.
Or type, which is the whole of the "entity" - metadata can be updated. Splitting a lot of hairs, I know....
The magic Pull doesn't seem to work?
You have no components - it can't work without those (and then only those with public data in Arctos).
The link to the preferred Organism ID doesn't do anything?
You've got that one linked to itself - its doing stuff, but nothing very exciting.
I clearly do not understand the "component" idea - I think I need a tutorial.
Use eg https://arctos.database.museum/guid/MSB:Mamm:233261 instead of MSB:Mamm:233261 and it'll all make sense - again, this works outside of Arctos, identifiers can be anything, GUIDs do stuff, "DWC Triplets" can't.
. But when I clicked on "find all same individual as" from the same record, I get 5 (the record I was searching from was excluded) which seems like a bug.
Feature request maybe - assuming that everything's the same as itself doesn't seem completely unreasonable....
Question - why are we creating "MWSN" for Mexican Wolf Studbook Number? We
should spell it out as Mexican Wolf Studbook Number or use "SB 789" which
is the actual format in the studbook. MWSN is not used in the studbook or
in any context I have seen in the recovery program, so let's avoid
introducing something entirely new. Maybe we need a code table for entity
ID formats? Or at least for ones associated with major external projects?
I agree with using "bad duplicate of" to merge.
But when I clicked on "find all same individual as" from the same record, I
get 5 (the record I was searching from was excluded) which seems like a bug.
I just requested this recently, but can't find the issue.
On Wed, May 19, 2021 at 1:17 PM dustymc @.*> wrote:
- [EXTERNAL]*
This looks nuts
You're using the wrong thing; along with looking nuts, that'll only work
for "us." Use the copy button to get the ID[image: Screen Shot 2021-05-19 at 11 56 36 AM]
https://user-images.githubusercontent.com/5720791/118868664-4b6eed00-b899-11eb-965f-f6568c278246.pngdisplay text instead of the link
My major epiphany in this process is that we can't - OrganismID can be
"3", or "http://someRandomWebsite.org/whatever" or something that Arctos
issues, and the only way to deal with that is to treat them all the same
(as strings, some of which DO STUFF in browsers, some of which might even
be useful).Add {stuff}
Whatever, it's just data, there's a CT process (which could also be used
to get rid of the stuff I've added, if necessary). BUT: the stuff I added I
added for discoverability, I'll pull "local" (Arctos, maybe GBIF, IDK)
records into the view at some point, if you just want to SEE that then
perhaps it doesn't need explicitly added.not hide the new entity creation/edit in the search function.
It's there on purpose; creating 17 entities for one critter and then
merging them is possible, but it's a lot of work and a mess. Searching
those things I've pulled in first is a tiny bit of work that can avoid that
mess. I think the step will prove very worthwhile, but whatever - there are
ways around that path, maybe duplicates won't happen, IDK, seems worth
trying out at least but still whatever....I did not assert their sex, U. S. Fish and Wildlife Service Mexican Wolf
Recovery Program did.If we're pulling it, that can be in the display. If we're asserting it for
the entity, then its not being asserted by USFWS (who was probably dealing
with something closer to an Occurrence - if this is a fish that assertion
could be less static). I'm not necessarily saying one or the other is some
form of "correct" and we should do that, just that there are two very
different kinds of data objects involved."bad duplicate of"
No terribly strong feelings on that, but it's not QUITE the same - maybe I
just don't want to use your assertion for some reason, mine's not "bad"
it's just not preferred by you.entities CAN be updated, just not the entity name.
Or type, which is the whole of the "entity" - metadata can be updated.
Splitting a lot of hairs, I know....The magic Pull doesn't seem to work?
You have no components - it can't work without those (and then only those
with public data in Arctos).The link to the preferred Organism ID doesn't do anything?
You've got that one linked to itself - its doing stuff, but nothing very
exciting.I clearly do not understand the "component" idea - I think I need a
tutorial.Use eg https://arctos.database.museum/guid/MSB:Mamm:233261 instead of
MSB:Mamm:233261 and it'll all make sense - again, this works outside of
Arctos, identifiers can be anything, GUIDs do stuff, "DWC Triplets" can't.. But when I clicked on "find all same individual as" from the same
record, I get 5 (the record I was searching from was excluded) which seems
like a bug.Feature request maybe - assuming that everything's the same as itself
doesn't seem completely unreasonable....—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-844395884,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBC3HF5RXXIQWVDKGQ3TOQFCZANCNFSM4G5YNGMA
.
Another suggestion: Add the "find all same individual as" pull function to
the entity page, and make the following in Edit Entity into live links:
The following catalog records use this Entity as Organism ID.
- MSB:Mamm:233261 (CAUTION::no component!)
- MSB:Mamm:268079 (CAUTION::no component!)
- MSB:Mamm:233262 (CAUTION::no component!)
- MSB:Mamm:233254 (CAUTION::no component!)
- MSB:Mamm:233267 (CAUTION::no component!)
- MSB:Mamm:270001 (CAUTION::no component!)
On Wed, May 19, 2021 at 4:50 PM Mariel Campbell @.*>
wrote:
Question - why are we creating "MWSN" for Mexican Wolf Studbook Number?
We should spell it out as Mexican Wolf Studbook Number or use "SB 789"
which is the actual format in the studbook. MWSN is not used in the
studbook or in any context I have seen in the recovery program, so let's
avoid introducing something entirely new. Maybe we need a code table for
entity ID formats? Or at least for ones associated with major external
projects?I agree with using "bad duplicate of" to merge.
But when I clicked on "find all same individual as" from the same record,
I get 5 (the record I was searching from was excluded) which seems like a
bug.
I just requested this recently, but can't find the issue.On Wed, May 19, 2021 at 1:17 PM dustymc @.*> wrote:
- [EXTERNAL]*
This looks nuts
You're using the wrong thing; along with looking nuts, that'll only work
for "us." Use the copy button to get the ID[image: Screen Shot 2021-05-19 at 11 56 36 AM]
https://user-images.githubusercontent.com/5720791/118868664-4b6eed00-b899-11eb-965f-f6568c278246.pngdisplay text instead of the link
My major epiphany in this process is that we can't - OrganismID can be
"3", or "http://someRandomWebsite.org/whatever" or something that Arctos
issues, and the only way to deal with that is to treat them all the same
(as strings, some of which DO STUFF in browsers, some of which might even
be useful).Add {stuff}
Whatever, it's just data, there's a CT process (which could also be used
to get rid of the stuff I've added, if necessary). BUT: the stuff I added I
added for discoverability, I'll pull "local" (Arctos, maybe GBIF, IDK)
records into the view at some point, if you just want to SEE that then
perhaps it doesn't need explicitly added.not hide the new entity creation/edit in the search function.
It's there on purpose; creating 17 entities for one critter and then
merging them is possible, but it's a lot of work and a mess. Searching
those things I've pulled in first is a tiny bit of work that can avoid that
mess. I think the step will prove very worthwhile, but whatever - there are
ways around that path, maybe duplicates won't happen, IDK, seems worth
trying out at least but still whatever....I did not assert their sex, U. S. Fish and Wildlife Service Mexican Wolf
Recovery Program did.If we're pulling it, that can be in the display. If we're asserting it
for the entity, then its not being asserted by USFWS (who was probably
dealing with something closer to an Occurrence - if this is a fish that
assertion could be less static). I'm not necessarily saying one or the
other is some form of "correct" and we should do that, just that there are
two very different kinds of data objects involved."bad duplicate of"
No terribly strong feelings on that, but it's not QUITE the same - maybe
I just don't want to use your assertion for some reason, mine's not "bad"
it's just not preferred by you.entities CAN be updated, just not the entity name.
Or type, which is the whole of the "entity" - metadata can be updated.
Splitting a lot of hairs, I know....The magic Pull doesn't seem to work?
You have no components - it can't work without those (and then only those
with public data in Arctos).The link to the preferred Organism ID doesn't do anything?
You've got that one linked to itself - its doing stuff, but nothing very
exciting.I clearly do not understand the "component" idea - I think I need a
tutorial.Use eg https://arctos.database.museum/guid/MSB:Mamm:233261 instead of
MSB:Mamm:233261 and it'll all make sense - again, this works outside of
Arctos, identifiers can be anything, GUIDs do stuff, "DWC Triplets" can't.. But when I clicked on "find all same individual as" from the same
record, I get 5 (the record I was searching from was excluded) which seems
like a bug.Feature request maybe - assuming that everything's the same as itself
doesn't seem completely unreasonable....—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-844395884,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBC3HF5RXXIQWVDKGQ3TOQFCZANCNFSM4G5YNGMA
.
spell it out as Mexican Wolf Studbook Number
That's no real problem, but it will be less "portable."
http://test.arctos.database.museum/entity/Mexican Wolf Studbook Number One Two Three Four
If you copy-paste that, you'll probably miss something, especially from text, like this: http://test.arctos.database.museum/entity/Mexican Wolf Studbook Number One Two Three Four
If you email it to yourself, your email client will probably do SOMETHING weird.
If you copy it out of a URL, you'll get
which works fine as a URL, but it's not correct for an identifier (some apps will make it work anyway).
Etc etc. Hence

make the following in Edit Entity into live links:
Those aren't good IDs - if you put the documented thing into the URL, you'll get links. (If you don't you'll get complaints, like in your copypassta.)
Add the "find all same individual as" pull function to the entity page,
That can be Phase Three, but I think I'd rather go the other way and create relationships from Entities.
This looks nutsYou're using the wrong thing; along with looking nuts, that'll only work for "us." Use the copy button to get the ID
Screen Shot 2021-05-19 at 11 56 36 AM
Even when it's the RIGHT thing, it looks nuts and if we stick with Mexican Wolf Studbook Number 1216 it will look like this:
http://test.arctos.database.museum/entity/Mexican%20Wolf%20Studbook%20Number%201216
Which is why I tried MWSN1216 and why I think we should use an index to assign the url extension - separate form the name! Start with 1 and then the urls are simple:
even if the searchable names are not. This is how GBIF does taxonomy and if we aren't going to allow deletion of entities, re-indexing shouldn't be necessary.
The magic Pull doesn't seem to work?You have no components - it can't work without those (and then only those with public data in Arctos).
I beg to differ - there are a bunch of components! https://arctos.database.museum/entity/MWSN1216
Are you saying that the WHOLE url is supposed to go in the component VALUE? I had assumed the value was whatever and the url was supposed to go in the AssertionURL field.
display text instead of the linkMy major epiphany in this process is that we can't - OrganismID can be "3", or "http://someRandomWebsite.org/whatever" or something that Arctos issues, and the only way to deal with that is to treat them all the same (as strings, some of which DO STUFF in browsers, some of which might even be useful).
But it is just UI? If we can do this:
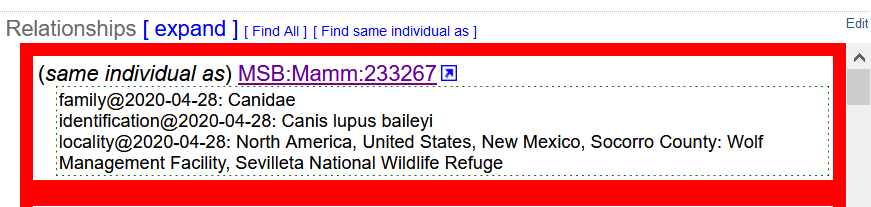
Why not this?
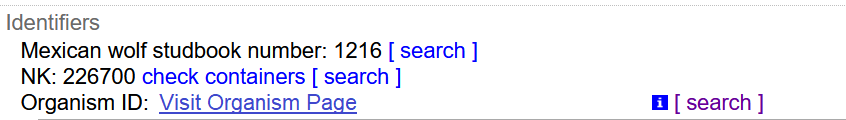
It is just to keep the catalog record from displaying three line long urls....
https://handbook.arctosdb.org/documentation/entity.html#the-process
Component:

No components:
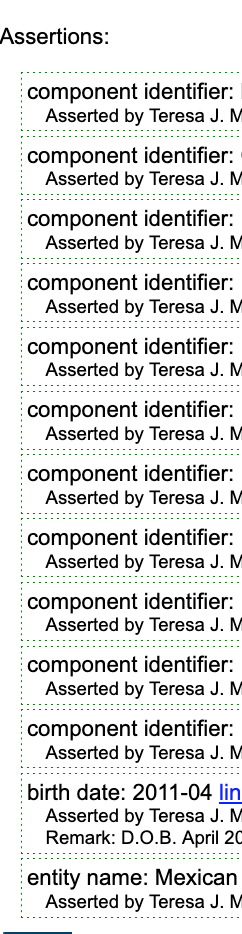

it is just UI?
Very much no.
we can do this:
All MSB:Mamm numbers have the same "prefix", OrganismID cannot.
displaying
- Find eerily familiar rat-bit on GBIF
- Click through to Arctos
- Splat-F
https://arctos.database.museum/entity/MWSN1216 - Nada
- Oh well, guess it wasn't the same rat.
No components
DOH!
All MSB:Mamm numbers have the same "prefix", OrganismID cannot.
So? I'm not saying that what you see on the record is what you get and what goes to GBIF should be https://arctos.database.museum/entity/MWSN1216.
I honestly don't understand Splat-F or what "find eerily familiar rat-bit on GBIF" means. I need a walk-through.
Anyway, maybe nobody cares if
is what you see for Organism ID OR that when we end up with Kianga the Giraffe, she will need to be
http://test.arctos.database.museum/entity/Kianga%20the%20Giraffe
I'm just saying, apparently not very clearly, that I think full identifiers are more useful to more people than partials. A record in GBIF (or anywhere outside of Arctos) must carry the full identifier, https://arctos.database.museum/entity/MWSN1216. Clicking through to that and then not finding it is potentially confusing, and seeing a partial identifier on other records might inspire users to _assign_ partials, which have limited functionality.
Kianga the Giraffe,
Yes! Any arbitrary string may or may not (probably not) be unique, and nonunique strings have limited functionality. "Kianga" likely refers to thousands of things, and different users will make different assumptions. http://test.arctos.database.museum/entity/Kianga is unambiguous - there can be only one, and even if it 404s there's some additional information that you might use to find your typo or get the issuer to clarify or something.
nobody cares if
See above and on the creation page - depending on their UI, they might very much care.
arctosprod@arctosutf>> select * from entity where entity_id='Mexican%20Wolf%20Studbook%20Number%20One%20Two%20Three%20Four';
entity_id | entity_type | created_by_agent_id | created_date
-----------+-------------+---------------------+--------------
(0 rows)
Maybe avoiding things that get encoded by browsers should be a rule rather than suggestion?
Edit: Or I could "issue" the encoded ID rather than the 'raw.'
there can be only one
But there isn't. I don't understand the opposition to assigning numbers as the entity identifier. Arctos can auto-assign, so there are no dupes. Anyone following the url will get all of the other goodies from there.
Edit: Or I could "issue" the encoded ID rather than the 'raw.'
THAT is what I have been saying for the last five comments! Also see above...
OK. Re-did https://arctos.database.museum/entity.cfm?action=edit&entity_id=MWSN1216 using pull. It wasn't clear to me at first that I could leave the component value blank and just put the Arctos guid in the AssertionURL, but I see that works as well as putting any old thing in the AssertionValue.
Not sure about putting the identifier type in remark. It seems like we need to be more specific about what the heck "NK" is or else we should just put the other_Identifier_type + otherid in the AssertionValue.
Also, I am not loving "component" but don't have a better term yet - I will think on it.
But there isn't.
There really is, the internet can't work without unique identifiers/URLs.
opposition to assigning numbers as the entity identifier
No real opposition from me, it's just an identifier. I sorta think https://arctos.database.museum/entity/MWSN1216 might be more informative than https://arctos.database.museum/entity/3 and I sorta think it might just be more confusing. The only really critical bit is whether Organism ID is issued and unique or not, everything else is icing from here.
Not sure about putting the identifier type in remark
Yea, I'm thinking the value should be "{otherIdType}{space}{display_value}" - I was thinking the raw number would be more searchable/less likely to produce duplicates, but now I'm not so sure.
not loving "component"
I'm not either, but I can't think of any other neutral term that covers Occurrences, pack members, family members, items in a set, and whatever else this might get used for.
putting any old thing in the AssertionValue
https://handbook.arctosdb.org/documentation/entity.html#the-process
Value needs sorted out and documented;
I don't think we can control that - someone at some point is going to be passionate about being able to use "Aunt Matilda's Favorite Fork" - but we could document, maybe even autofill/suggest for familiar things.
OK - I vote for assigned identifiers. They may not be informative, but they will create neater urls. Perhaps we should adopt something like the Wikidata Q numbers? "{Arctos}{integer}"?
component
"a part or element of a larger whole, especially a part of a machine or vehicle."
(why not just use part?)
part - a piece or segment of something such as an object, activity, or period of time, which combined with other pieces makes up the whole.
This seems most intuitive but possibly confused with catalog record parts?
unit - an individual thing or person regarded as single and complete but which can also form an individual component of a larger or more complex whole.
Also more intuitive than "component"? but not confused with catalog record parts.
constituent - a component part of something.
(UGH)
portion - a part of a whole.
(why not just use part?)
Given all of the above - I suggest we use part....
part
At least in my head, "parts" are simple things, "components" are somehow more complex - like the "parts" we already have, https://handbook.arctosdb.org/documentation/parts.html.
I dislike "element" less, but still seems arbitrarily different rather than better and would involve changing some code (but there will never be a better time than now to do that, if we must).
"element" also risks confusion with chemistry....
At least in my head, "parts" are simple things
IMO - not simple!
https://arctos.database.museum/info/ctDocumentation.cfm?table=ctspecimen_part_name#gastrointestinal_tract
https://arctos.database.museum/info/ctDocumentation.cfm?table=ctspecimen_part_name#head
https://arctos.database.museum/info/ctDocumentation.cfm?table=ctspecimen_part_name#heart__kidney__liver__spleen
"Part" of the Earth is North America - NOT simple....
Apologies for not following this critical discussion in more detail - I've
been getting repeatedly pulled off for other things. However, I do want and
need to understand what is going on here. I just got a loan request for
blood from three zoo wombats sampled multiple times, so there are 20
catalog records. I need to create the entities for these and all other zoo
material in the project I shared earlier. I still don't know how to create
an entity ID - it isn't in the list of Other IDs to add to from the Edit
Other Ids on the specimen page, which is the logical place I looked. It's
also not in the manage dropdown for the list of related items when I click
search next to the zoo or GAN number in the specimen page. I'm sure the how
tos are somewhere in this thread - can you reattach here?
As for component, I don't like the term either. When I look at the entity
page, all of those things seem like "associated identifier" etc, so going
back to my earlier comment:
Can we change some of the controlled vocab, e.g. replace "component" with
"related catalog item" or "associated guid"? Related url? Something
human-recognizable?
Also, can we change the display order to be:
component_1 = related catalog item_1 = MSB:Mamm:326433 link
http://test.arctos.database.museum/guid/MSB:Mamm:326433
component_ID_1= related_catalog_item__ID_1 =Pan troglodytes
component_identifier_1 = related_catalog_item_identifier_1= Kianga
component_2=related_catalog_item_2 = Some mouse or something link
http://test.arctos.database.museum/guid/MSB:Mamm:999
component_ID_2 = related_url_ID_2 = Peromyscus leucopus
component_identifier_2= related_identifier_2= M07003
http://arctos.database.museum/SpecimenResults.cfm?oidtype=Albuquerque%20Biopark%20Zoo%20Local%20ID&oidnum=M07003
component_3= organism_ID_3=
http://test.arctos.database.museum/entity/boogity
On Thu, May 20, 2021 at 1:09 PM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
At least in my head, "parts" are simple things
IMO - not simple!
https://arctos.database.museum/info/ctDocumentation.cfm?table=ctspecimen_part_name#head
"Part" of the Earth is North America - NOT simple....
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-845403554,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBAWTVWQAVKGVZ3LAZ3TOVM7HANCNFSM4G5YNGMA
.
on't know how to create
an entity ID
https://handbook.arctosdb.org/documentation/entity.html#the-process
replace "component" with
"related catalog item" or "associated guid"? Related url?
It's not necessarily any of those things. What is the objection to the term? That somehow seems to be a major hangup, I don't understand why. I'm not particularly attached to "component" but I think it's more accurate and less confusing than anything else that's been offered.
order
We agreed to just pull those data in a few days back??
Yes, but *where *do I go to do this? And can I get there from the specimen
record?
The simplest use case involves simply creating an Entity. Creation is
available to authorized users after search results; users are expected to
ensure that they are not creating functional duplicates. (True duplicates
are prevented.)
[image: Screen Shot 2021-05-12 at 8 46 58 AM]
This results in an Entity ID suitable for use as an Organism ID.
On Thu, May 20, 2021 at 5:12 PM dustymc @.*> wrote:
- [EXTERNAL]*
on't know how to create
an entity IDhttps://handbook.arctosdb.org/documentation/entity.html#the-process
replace "component" with
"related catalog item" or "associated guid"? Related url?It's not necessarily any of those things. What is the objection to the
term? That somehow seems to be a major hangup, I don't understand why. I'm
not particularly attached to "component" but I think it's more accurate and
less confusing than anything else that's been offered.order
We agreed to just pull those data in a few days back??
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-845541440,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBDVB5BXNPGABIAVEJDTOWJOTANCNFSM4G5YNGMA
.
Ah - I didn't put that in the docs because there's a request to move the link.
Oh - yes, please move it! And we do need to access it from the specimen
(catalog) record.
On Thu, May 20, 2021 at 5:42 PM dustymc @.*> wrote:
- [EXTERNAL]*
Ah - I didn't put that in the docs because there's a request to move the
link.[image: Screen Shot 2021-05-20 at 4 41 19 PM]
https://user-images.githubusercontent.com/5720791/119061884-450c6e00-b98a-11eb-925d-bca0c2f14957.png—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-845552425,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBCTPROPJ6ODPJ4M5Q3TOWM6HANCNFSM4G5YNGMA
.
we do need to access it from the specimen (catalog) record.
You're going to have to elaborate on that.
When I search on a particular ID, and get a list of all catalog records
that share it (e.g. try Albuquerque Biopark Zoo Local ID: M10031), and I
see that there are more than a single record, I want to be able to create
on the spot an entity ID to link them all. I can click "search" on the
Other ID M10031 and pull up all 6 records. Ideally, it makes most sense to
include entity create in the "manage" dropdown for the search results.
On Thu, May 20, 2021 at 5:46 PM dustymc @.*> wrote:
- [EXTERNAL]*
we do need to access it from the specimen (catalog) record.
You're going to have to elaborate on that.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-845553773,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBEHX253CK23OVBTDLLTOWNM7ANCNFSM4G5YNGMA
.
Worst case, that's a decent recipe for lots of duplicates. (And if the intent is to be that casual about creating Entities, why aren't we just using those existing IDs? I can't see much functional difference.)
Best case, it sounds like a fringe use case that won't ever get used after the initial obvious Entities are created, which I can magic after there's some real usage.
because there's a request to move the link.
Not a request to move a request to add a link. I'd like to treat Entities in the same way we treat Agents.
Search here

Manage here
@campmlc before you go creating a whole bunch of these I really think we need to sort out some stuff. Primarily whether the ids will be issued by Arctos or made up by people.
Ok, perhaps a call?
On Fri, May 21, 2021, 8:24 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
because there's a request to move the link.
Not a request to move a request to add a link. I'd like to treat Entities
in the same way we treat Agents.Search here
[image: Screen Shot 2021-05-20 at 4 41 19 PM]
https://user-images.githubusercontent.com/5720791/119061884-450c6e00-b98a-11eb-925d-bca0c2f14957.pngManage here
[image: image]
https://user-images.githubusercontent.com/5725767/119152642-bb05e900-ba0d-11eb-90ab-8a5908c8f583.png@campmlc https://github.com/campmlc before you go creating a whole
bunch of these I really think we need to sort out some stuff. Primarily
whether the ids will be issued by Arctos or made up by people.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-845986483,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBGWEH4TRRFWX3IXAT3TOZUKRANCNFSM4G5YNGMA
.
same way we treat Agents.
Agents is redundant (mostly because the technology has changed since the "private" stuff was built) - newer forms are all (when possible) more like Entity, in that there's one form with certain parts available only under certain permissions. Normalization ain't just for data....
ids will be issued by Arctos or made up by people.
Yes, I see three possibilities:
- accept any string (what we do now) - see above for potential problems and benefits, I think I'm with @Jegelewicz, this isn't great
- accept any constrained string (eg perhaps HTML encoding doesn't change anything - so no spaces, apostrophes, etc.) - avoids the encoding problems of above, still allows some limited "smartness" (NK123 could work) - not that that's always a good thing - might be frustrating when someone's already used all of the identifiers you'd like to use
- assign something (integers are easy) - just an arbitrary identifier, no "data" embedded in the ID, just click 'create' to get one
I don't have any real preferences.
I just patched in a new widget in response to some comment that's now buried in github's horrid little 'fold' - it uses 'component', we're getting a lot of code around that term, if there's something WRONG with it then we need a solution ASAP.
call
Yep, I was actually wondering if we shouldn't do that a lot more regularly - I just hang out on Zoom for some hour every week or something? Anyway, suggest a time.
Aforementioned new widget:
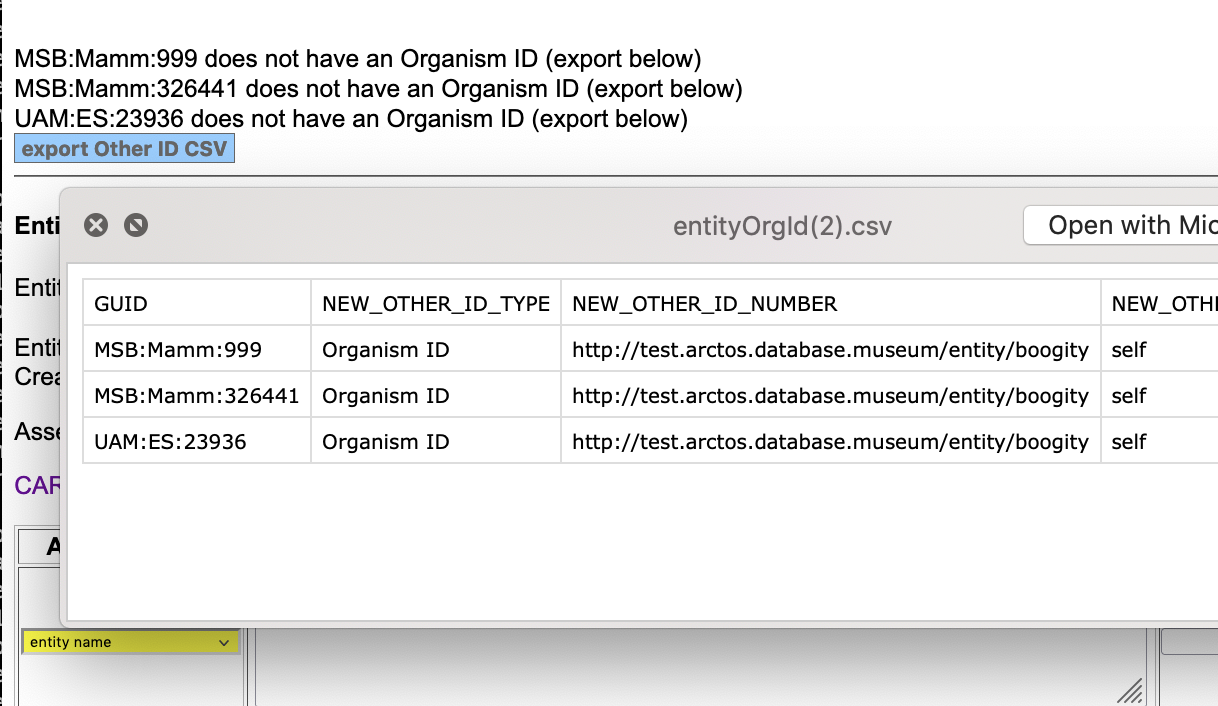
QnD demo of some potential non-Organism types of Entity:
http://test.arctos.database.museum/entity/someBonesFromARock ("set" - I couldn't find dinnerware in test, bones might be something different)
http://test.arctos.database.museum/entity/checkllistOfArbitraryTaxonomy ("checklist" - no particular reason the components all need to be the same type, could include agents or projects or publications or etc)
http://test.arctos.database.museum/entity/someGeneticLineOrSomething ("family" - things that share significant DNA, or whatever)
Right now I am free after 2 MDT today and all day Monday except from 2 to 4MDT.
I just hang out on Zoom for some hour every week or something?
I like it - Dusty's Office Hour
I think you should pick the day/time that works for you and put it on the Arctos Google Calendar.
Dusty office hour sounds good - Monday 1pm for this time around?
On Fri, May 21, 2021 at 12:03 PM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
Right now I am free after 2 MDT today and all day Monday except from 2 to
4MDT.I just hang out on Zoom for some hour every week or something?
I like it - Dusty's Office Hour
I think you should pick the day/time that works for you and put it on the
Arctos Google Calendar.—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-846140021,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBAFO3S7QELJMIUSD3DTO2N5JANCNFSM4G5YNGMA
.
Yeah office hours!
Do we keep it quiet for right now or would you like something to go out that is more formal?
Ok with me to announce as long as it is clear it is for Organism ID
discussion this time
On Sat, May 22, 2021, 7:52 PM Elizabeth Wommack @.*>
wrote:
- [EXTERNAL]*
Yeah office hours!
Do we keep it quiet for right now or would you like something to go out
that is more formal?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-846488420,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBGYFNUTSJUJ7NFBSWTTPBNVVANCNFSM4G5YNGMA
.
@ewommack if you'd facilitate scheduling that would be fabulous. I'm pretty flexible on the time, but noon (Pacific) isn't great. 10 is good but probably too early for the east coasters. Maybe 1 Pacific on Mondays as a starting point?
That works for me.
On Mon, May 24, 2021, 8:04 AM dustymc @.*> wrote:
- [EXTERNAL]*
@ewommack https://github.com/ewommack if you'd facilitate scheduling
that would be fabulous. I'm pretty flexible on the time, but noon (Pacific)
isn't great. 10 is good but probably too early for the east coasters. Maybe
1 Pacific on Mondays as a starting point?—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-847064352,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBGB4VD6BRZBPIIQ7I3TPJMFRANCNFSM4G5YNGMA
.
I currently have a standing meeting at 1 Pacific on Mondays....
And BTW 10 Pacific is 1PM eastern - I hope that isn't too early....
Coffee, THEN trying to figure out where the sun comes from, got it....
So noon Pacific today then 10 Pacific going forward?
I'm good with that!
For today's discussion -
1) how to change or edit an entity ID - or are they really necessary? Where
exactly are they visible? Internal to Arctos only? I see how to add a
preferred entity ID or an entity name, but not how to edit an existing
entity ID.
2) How do we link component data with their sources, e.g. component
identifiers, component identifications, with the source of the data - e.g.
the Arctos url/guid, some other online source, and differentiate these
clusters of data from other clusters?
[image: image.png]
On Mon, May 24, 2021 at 8:55 AM Teresa Mayfield-Meyer <
@.*> wrote:
- [EXTERNAL]*
I'm good with that!
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-847097225,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBBOGPZOCKBTJUYEYCLTPJSE3ANCNFSM4G5YNGMA
.
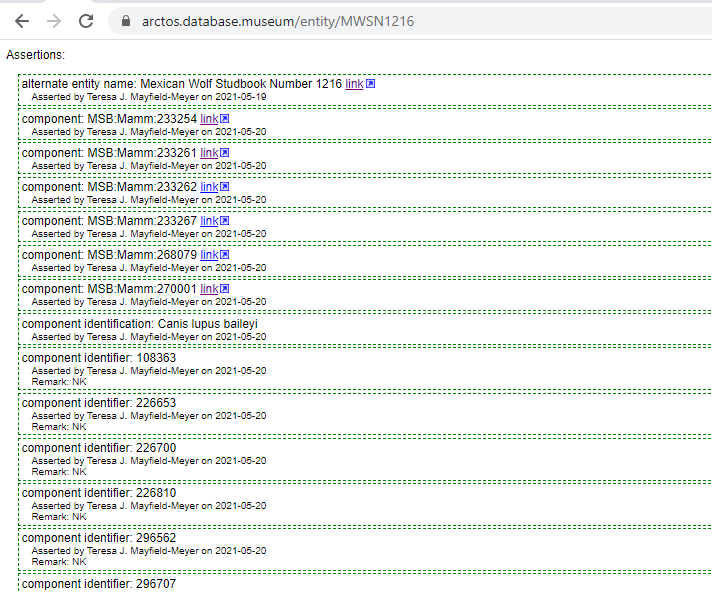
Also, where should "create entity" and "edit entity" be accessed from on the tool bars, catalog record page, etc.
Just back from the field. I gather there is a meeting today at 12 PM Pacific to discuss this?
entity ID - or are they really necessary?
Entity IDs are THE core component. (The format of the IDs - as long as they're resolvable - is not important, see https://github.com/ArctosDB/arctos/issues/1966#issuecomment-846006301)
not how to edit an existing entity ID.
You cannot, and if you could then this would be a very different (less stable, I believe) model.
2) How do we link component data with their sources,
I think this is a request to add a great deal of complexity and no additional functionality. (DISPLAY is another matter - we can pull data from Arctos, probably GBIF, at will.) That doesn't mean we shouldn't do it, but I think we should have an understanding of the big-picture goals, and the implications of adding this complexity, before changing the model. (My guess: it would be enough work that it won't get used, the entity will therefore bear less data, duplicates will be created, this will turn into a mess that nobody will use for any purpose.)
accessed from on the tool bars, catalog record page, etc.
That's generally trivial to adjust, but eg allowing create without first requiring search is almost certain to have functional implications (eg duplicates will be created and they will - rightly - make big enough messes that nobody will want to use this).
meeting today at 12 PM Pacific
Yes.
Very Q&D "fetch" in test:

OK, I'll be there at noon.
I'm not doing something right. Here is a record with two events:
https://arctos.database.museum/guid/MVZ:Bird:193195
I created an observational record for the second event:
https://arctos.database.museum/guid/MVZObs:Bird:4777
and selected for both a 'Organism ID' identifier
https://arctos.database.museum/entity/0709-02237
(manually entered the URL which I'm sure is not correct, but I didn't see a base URL in the code table)
When I click on the Organism ID link, I get "Entity not found! Please let us know what happened."
"Entity not found!
You didn't create one.
https://handbook.arctosdb.org/documentation/entity.html
I did this for you:


nope not there so


and now you have the bare minimum.
The next step would (ideally - this is now functional) be to add the components.
Then clicking "pull" and accepting whatever it says would add some discoverability.
@Jegelewicz was amazing and added the office hours to the calendar.
Do we want any note or explanation @dustymc?
"Dusty's Office Hours are discussions with Dusty on specific problems and production developments in Arctos. Come join the conversation and help us figure out how to make Arctos better"
Thanks!
I'm up for anything. I'll probably be more useful with some warning, I think we can/should prioritize if someone wants to schedule a topic, otherwise just see what happens?
I'll probably be more useful with some warning, I think we can/should prioritize if someone wants to schedule a topic
How about:
"Dusty's Office Hours are discussions with Dusty on specific problems and production developments in Arctos. Suggest a topic ahead of time in GitHub, or just come join the conversation and help us figure out how to make Arctos better"
From meeting:
- clarify search before create functionality
- Search is one field, hits everything possible, has usage hint
- show derived data (component IDs and such) in some less-central way
Changes
- entityID is assigned by Arctos; you get what you get and don't have a fit
- entity description (new field in table entity, required, editable, @campmlc will write documentation)
- pull is automagic
- manage_collection is required to create/edit
Unresolved:
- show more dynamic view in search result
- DO NOT show more dynamic view in search result
It's less-dynamic for now, not sure we have the CPU to pull everything in anyway. Looking forward, this needs to (theoretically) work for hundreds (zoo critters have a rough life) if not thousands (GPS collar, maybe) of components, which probably demands separate search results and 'details' views.
Needs further discussion:
Entities are but one option for Organism ID, and therefore the code is "Entity-centric." Organism ID can be exported from Entities to catalog records, but Entity ID cannot be exported/created from catalog records. I suggest that this is sufficient; Entities are "super objects" that only need exist when there's something additional to say. If the only goal it a common identifier for Organism ID, there are many options which do not involve Entities (bird banding lab numbers, for example). Entities are "better" identifiers, and making sure that they are in fact "better" requires a small amount of focus.
Yea But Anyway:
Consider something in SpecimenResults-->Manage-->Add All Records to {pick an entity}
- I think I'm comfortable with this to ADD, not so sure about CREATE
Needs Clarification
re: "bird banding lab numbers, for example" above: There is confusion around this point, it needs clarified somewhere. A number may/should be used in multiple types, because those types convey different information and have different functionality. For example, to use a BBL number as an Organism ID, the following should be entered (assuming BBL was an OtherID Type in Arctos):
- BBL: 12345
- Organism ID: BBL 12345
The BBL number supports "find records with a BBL number" (and perhaps value, but free-text fields aren't very good at that), and potentially (should BBL come online) can serve as a link to external resources or additional data.
The Organism ID serves as an Organism ID; it's an identifier that spans multiple Occurrences and links them together as one THING. In this case that link is dependent on users being consistent (eg, not using Organism ID: BBL{nospace}12345 in one of the involved records), and should be recognized as having limited scope (somewhere on the planet, there's probably an unrelated, perhaps even similar, "BBL 12345.") There's no realistic way for machines to determine if BBL{nospace}12345 and BBL 12345 should be the same thing; error detection requires (patient) humans.
Entities (of type Organism) serve the same purpose; they're linking identifiers. They differ in two significant ways:
- There's a verifiable "correct" format; identifiers issued by Arctos behave differently than those which were not (eg, typos).
- The Entity can carry data of its own, and this data can be used in things like error detection.
tl;dr: Any string can serve as Organism ID, but some can DO THINGS that others cannot.
Bulk Tools:
MSB's biopark data is recent and decent, but should have enough problems to be interesting. Try to make and "componentize" Entities from it, with a view towards developing bulk tools. (This may address any gaps left by the entity-centric approach described avove.)
Reports:
- See if the stuff from edit entity (components don't use entity ID, records using entity ID aren't components) can be made into reports and/or bulkloaders.
- All entities should have components or
preferred entity ID
Possibilities:
Rather than Export, we could write to the ID loader with status=autoload
- Yay: one click instead of ~4
- Not so yay: Fixing the giant messes that approach is capable of creating could be a tremendous amount of work (which usually means it'll never happen, and then nobody will use this because it's all a giant mess). Suggest the small amount of review required to manually use the loader is well invested.
"Reports" above has the same implications; we could save a few minutes by automating, which might then require much more than a few minutes to fix the giant mess which could result from a relatively minor error.
@campmlc @Jegelewicz @ccicero what'd I miss/mangle?
There's some new stuff in test, https://handbook.arctosdb.org/documentation/entity.html#the-process-v2 documents creating http://test.arctos.database.museum/entity/2
Questions:
- What should I auto-pull into Entity Assertions from catalog records; what data might lead someone to an existing Entity and prevent them from creating a duplicate?
- What should I dynamically pull on the detail page; what's useful there?
Not sure if this will be helpful, but here are several references for BBL bands: https://www.usgs.gov/centers/eesc/science/about-federal-bird-bands?qt-science_center_objects=0#qt-science_center_objects
BBL bands always have two sets of numbers XXXX-XXXX or XXXX-XXXXX. The first string relates to the size of the band, and the second string is in sequence numerically assigned to individual banders. I can't find a reference for the numeric codes for the different sizes, but I'm sure it exists somewhere. I could dig deeper if you need me to.
They keep strong track of which of us has which bands, because as you can guess mistakes get made all the time. That way they know who to poke/yell at if a warbler band comes back being reported on a Red-tailed Hawk.
Thanks. Nothing can really change how unresolvable strings work, but entities could serve as a place to gather identifiers - the Entity itself can hold all the variations that might be found in GBIF-n-such (BBL:XXXX-XXXX; BBL XXXX-XXXX; XXXX-XXXX, XXXXXXXX, etc., etc.) and that has some possibility of leading users to those records if they find the Arctos record.
What should I auto-pull into Entity Assertions from catalog records; what data might lead someone to an existing Entity and prevent them from creating a duplicate?
Identification (taxon)
All other identifiers
Attributes (of the catalog record item)
Latest is in production, I rebuilt the two Entities I could, old data is in arctos-assets.
Sorry I haven't worked on this - I've been busy cleaning ichnotaxa and part names.....
I think we've all had our distractions lately!
Yes, I can't wait to try these out. Any way to do a mass entity bulkload
for ABQ Biopark?
On Fri, May 28, 2021, 4:37 PM dustymc @.*> wrote:
- [EXTERNAL]*
I think we've all had our distractions lately!
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
https://github.com/ArctosDB/arctos/issues/1966#issuecomment-850706490,
or unsubscribe
https://github.com/notifications/unsubscribe-auth/ADQ7JBAW3LBY3PLFLIFOSYTTQALJZANCNFSM4G5YNGMA
.
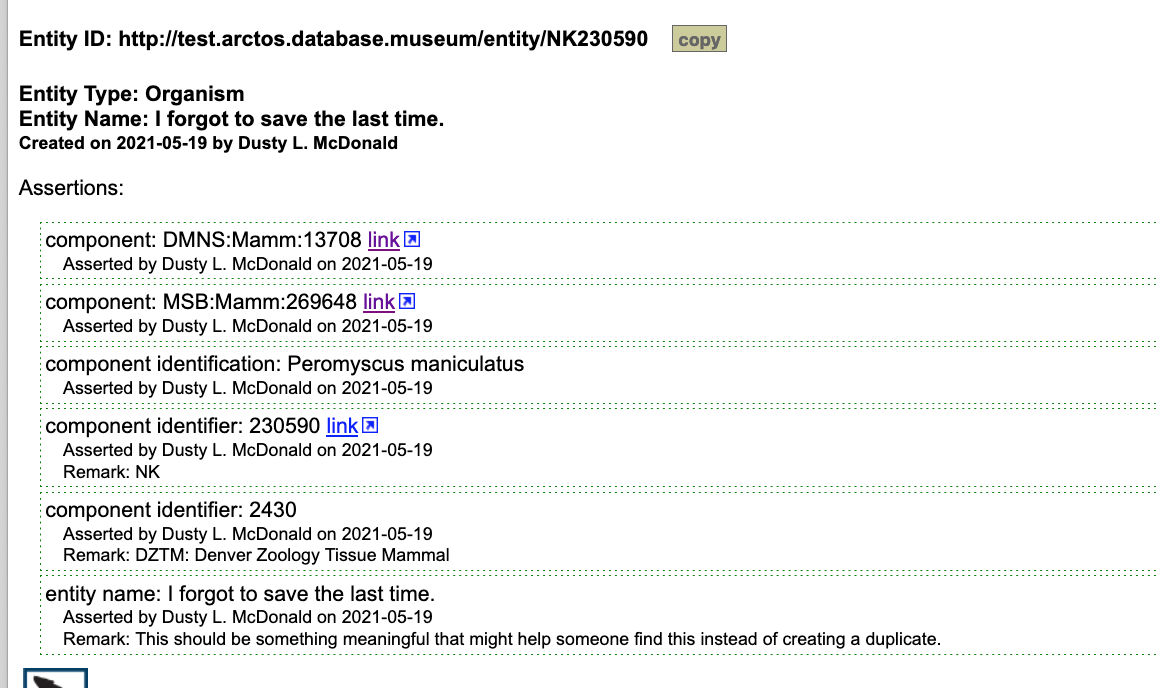

Most helpful comment
How about:
"Dusty's Office Hours are discussions with Dusty on specific problems and production developments in Arctos. Suggest a topic ahead of time in GitHub, or just come join the conversation and help us figure out how to make Arctos better"